Week 2 HW: DNA Read, Write, Edit
3.1. Choose your protein
I chose miniSOG (mini Singlet Oxygen Generator) using the protein table from FPbase. It is described as a cyan fluorescent protein that can be controlled with blue light. When illuminated, the molecule absorbs energy and transfers it to nearby oxygen, briefly converting it into a reactive form called singlet oxygen. This state lasts only a few microseconds inside cells and travels about 10–20 nanometers, making it useful for nanoscale targeting. Because it can repeatedly trigger localized reactions without being consumed, it behaves more like a catalyst than a reagent.
miniSOG is an engineered 106-amino-acid protein derived from a plant protein in Arabidopsis thaliana (mouse-ear cress). Plants need to sense light to grow toward the sun and regulate circadian rhythms, so they evolved blue-light-detecting proteins called phototropins. These proteins contain a LOV domain (Light, Oxygen, Voltage sensing domain) that holds a vitamin B2-derived molecule called FMN (flavin mononucleotide). FMN acts as a photosensitizer by absorbing blue light and transferring energy to oxygen.
By modifying this domain, researchers repurposed the natural light sensor into miniSOG, which generates localized oxidative reactions when activated. This allows scientists to mark proteins, damage specific organelles such as mitochondria, selectively kill cells, and stain cellular structures for imaging.
The protein sequence is as follows:
MEKSFVITDP RLPDNPIIFA SDGFLELTEY SREEILGRNG RFLQGPETDQ ATVQKIRDAI RDQREITVQL INYTKSGKKF WNLLHLQPMR DQKGELQYFI GVQLDG
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence
I used the Bioinformatics reverse translation tool to obtain 318 base sequence of most likely codons:
atggaaaaaagctttgtgattaccgatccgcgcctgccggataacccgattatttttgcg agcgatggctttctggaactgaccgaatatagccgcgaagaaattctgggccgcaacggc cgctttctgcagggcccggaaaccgatcaggcgaccgtgcagaaaattcgcgatgcgatt cgcgatcagcgcgaaattaccgtgcagctgattaactataccaaaagcggcaaaaaattt tggaacctgctgcatctgcagccgatgcgcgatcagaaaggcgaactgcagtattttatt ggcgtgcagctggatggc
3.3. Codon optimization
E. coli is the easiest organism to work with in the lab so for this exercise the below is the codon optimization using GenSmart:
ATGGAAAAATCATTTGTAATAACAGATCCCCGCCTGCCAGATAACCCGATTATTTTCGCTAGCGACGGCTTTTTGGAGTTGACTGAGTACAGCCGTGAGGAAATTCTGGGTCGCAACGGCCGTTTTCTGCAAGGTCCGGAAACCGATCAGGCAACGGTGCAAAAGATCCGTGACGCGATTCGCGACCAACGTGAGATCACCGTCCAGCTTATCAACTATACCAAATCCGGTAAAAAGTTCTGGAATCTGCTGCACCTGCAGCCGATGCGTGACCAGAAGGGTGAATTACAGTACTTCATCGGCGTTCAACTGGATGGC
3.4. What technologies could be used to produce this protein from your DNA?
To produce the miniSOG protein, the DNA sequence encoding miniSOG is inserted into a plasmid expression vector containing a promoter, ribosome binding site, and terminator. The plasmid is introduced into E. coli through transformation. Inside the bacteria, RNA polymerase binds to the promoter and transcribes the miniSOG gene into messenger RNA (mRNA). Ribosomes then translate the mRNA codons into amino acids with the help of tRNAs, forming the 106-amino-acid miniSOG polypeptide. The protein folds into its functional structure and binds FMN, allowing it to function as a light-activated singlet oxygen generator.
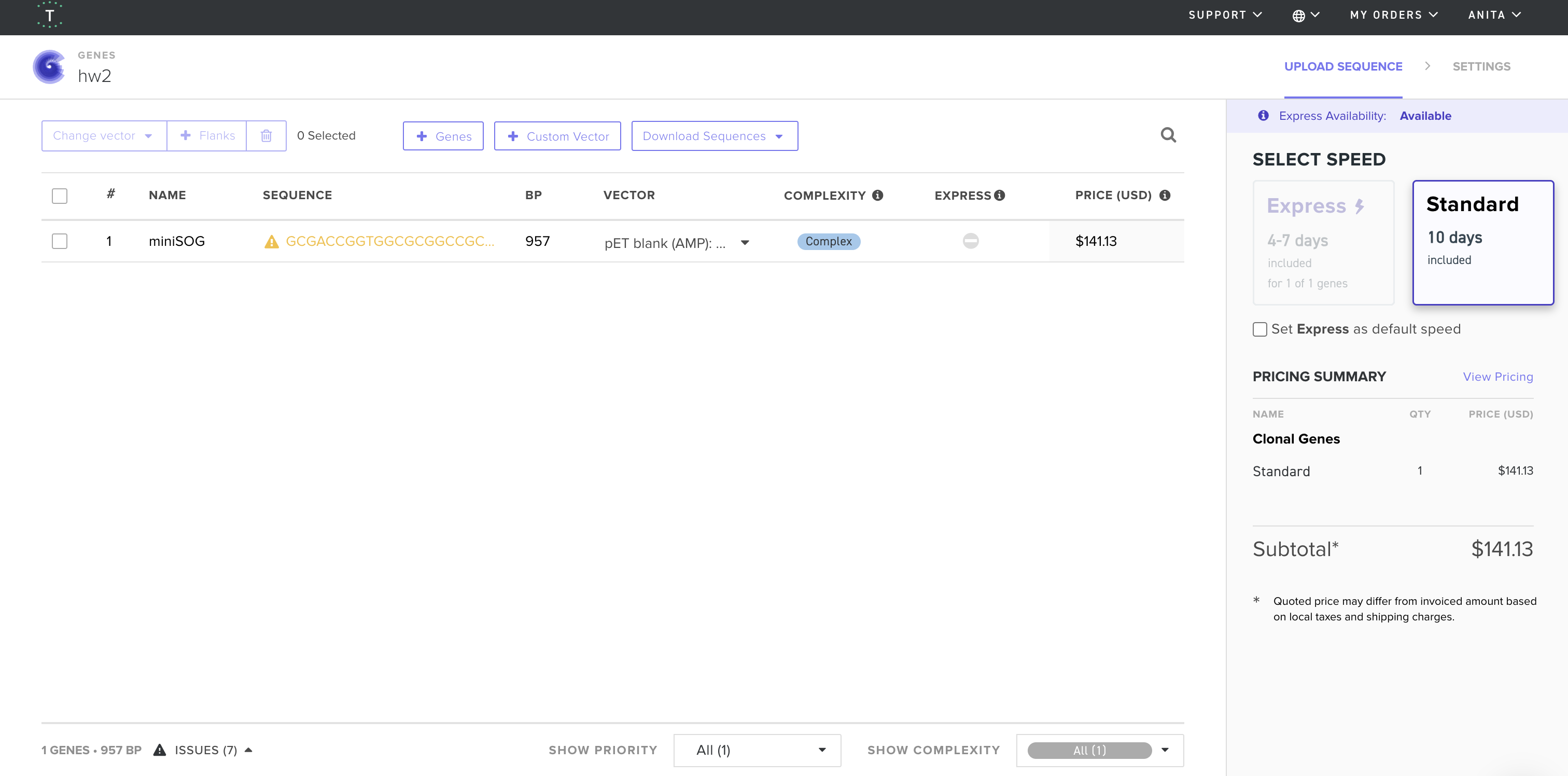
4. Twist and Benchling
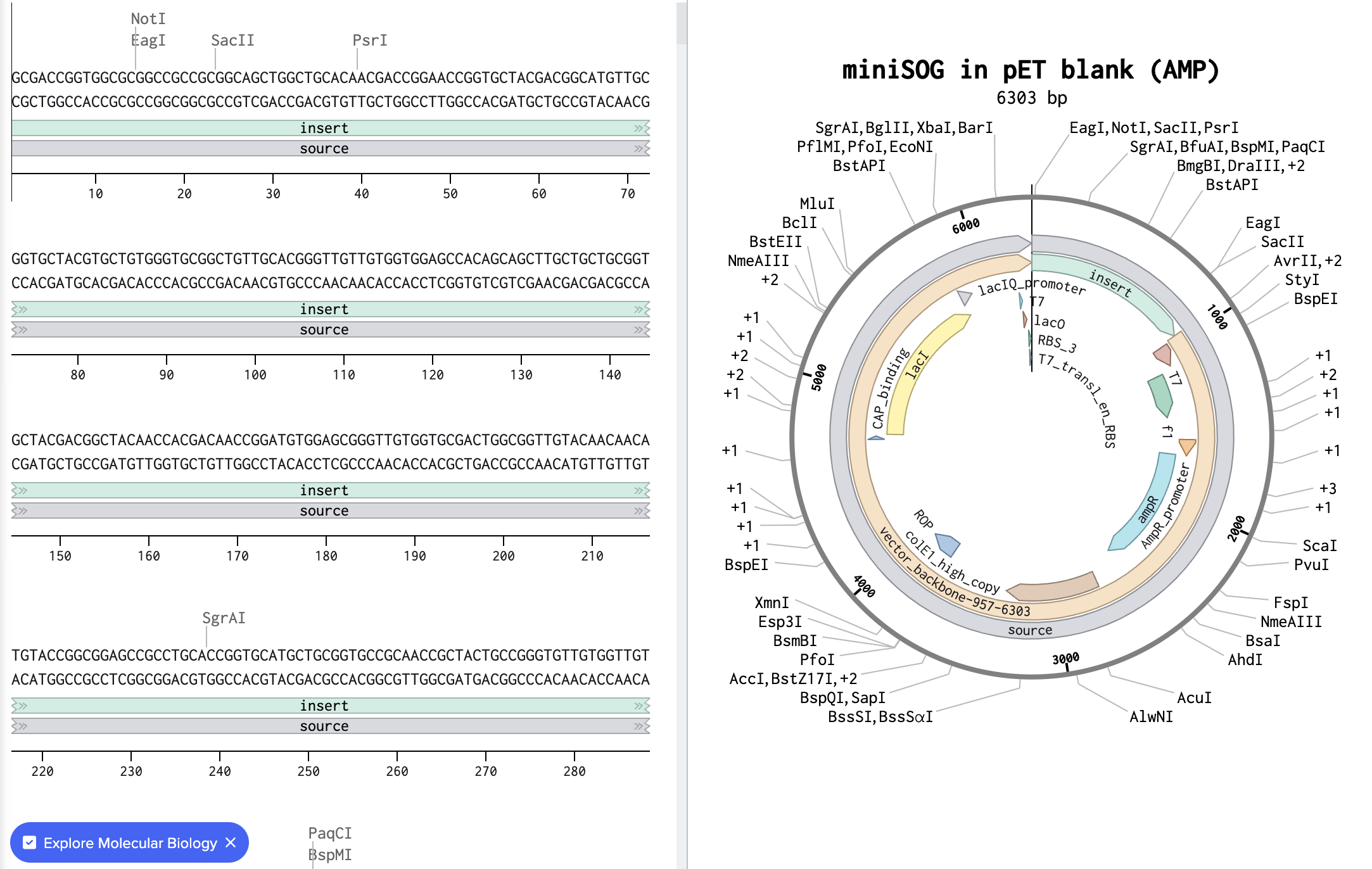
I imported the GenBank file from Twist Bioscience into Benchling and below is the resulted visualization:
5.1. DNA Read I would use nanopore sequencing (for example Oxford Nanopore sequencing) to sequence the DNA encoding miniSOG. This method can read very long DNA molecules, so it can sequence the entire plasmid containing the gene in a single read instead of reconstructing it from short fragments.
Nanopore sequencing is a third-generation sequencing technology because it reads single DNA molecules directly in real time without PCR amplification or synthesis reactions. Unlike second-generation methods, it does not rely on fluorescence or imaging.
Input and sample preparation: purified plasmid DNA isolated from E. coli.
Preparation steps: Extract plasmid DNA from the bacteria > Linearize or lightly fragment the plasmid > Ligate sequencing adapters and a motor protein to the DNA ends > Load the DNA library onto the nanopore flow cell
Essential sequencing steps and base calling. The flow cell contains a membrane with tiny protein nanopores. An electrical voltage is applied across the membrane, causing ions to flow through each pore and creating a measurable current.
A motor protein feeds a single DNA strand through the nanopore > As bases pass through the pore, they partially block ion flow > Each base (A, T, C, G) produces a characteristic change in electrical current > The instrument records this current trace > Software analyzes the signal and converts the current pattern into nucleotide identities.
The output is long sequencing reads that contains the nucleotide sequence and a Q score for each base