Week 7: Genetic Circuits Part II
Part 1: Intracellular Artificial Neural Networks (IANNs)
What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions?
IANNs have an advantage over traditional genetic circuits because they can handle inputs in a more continuous way instead of just on/off logic. Regular genetic circuits are basically Boolean, so they struggle with noisy or overlapping signals. IANNs can take in multiple inputs, weigh them differently, and produce a more gradual response, which is closer to how biological systems actually behave.
Describe a useful application for an IANN; include a detailed description of input/output behavior, as well as any limitations an IANN might face to achieve your goal
One application could be a living environmental sensor that detects pollution levels. For example, it could take in inputs like pH, temperature, and the presence of certain toxins, and then output a level of fluorescence based on how severe the conditions are, instead of just turning on or off. This would make it more useful for real-world monitoring where conditions aren’t binary. That said, there are still limitations. Cells are noisy systems, so the outputs might not always be consistent. There’s also the issue of metabolic burden and slower response times, and it’s still difficult to precisely tune or “train” these networks inside living cells.
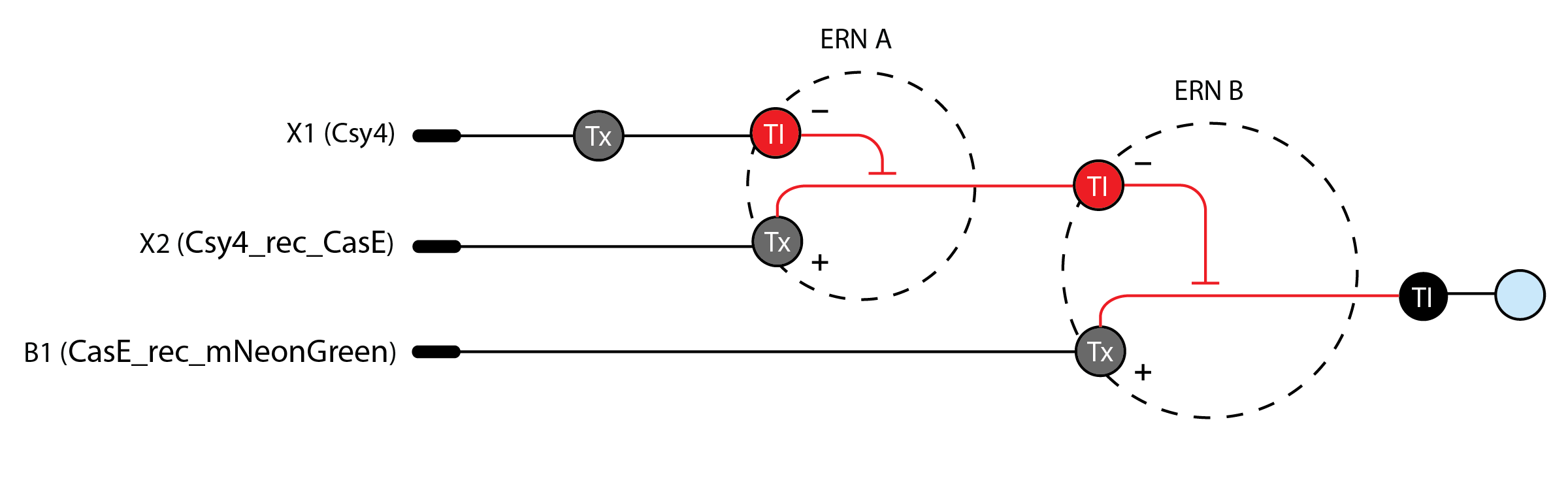
Below is a diagram depicting an intracellular single-layer perceptron where the X1 input is DNA encoding for the Csy4 endoribonuclease and the X2 input is DNA encoding for a fluorescent protein output whose mRNA is regulated by Csy4. Tx: transcription; Tl: translation
Draw a diagram for an intracellular multilayer perceptron where layer 1 outputs an endoribonuclease that regulates a fluorescent protein output in layer 2

Part 2: Fungal Materials
What are some examples of existing fungal materials and what are they used for? What are their advantages and disadvantages over traditional counterparts?
Fungal materials, especially those made from filamentous fungi, are starting to be used as sustainable alternatives in areas like construction, textiles, packaging, and insulation. These materials come from mycelium, which is a network of hyphae that grows through organic matter and binds it together into a solid structure. The properties of these materials are largely influenced by what makes up the fungal cell wall… mainly chitin, which adds rigidity, and β-glucans, which provide flexibility.
Compared to traditional materials like plastics or leather, fungal materials are biodegradable, require less energy to produce, and can be grown using agricultural waste. Another advantage is that their properties can be adjusted by changing the fungal species, the substrate, or how they are grown and processed. That said, they still have some limitations, like lower strength, sensitivity to moisture, and inconsistency due to natural biological variation. Different species also behave quite differently, meaning material performance can vary a lot depending on what fungus is used. Overall, fungal materials show a lot of promise as sustainable alternatives, but they still need better control and development before they can fully compete with conventional materials.
Examples of projects that incorporates mycelium:
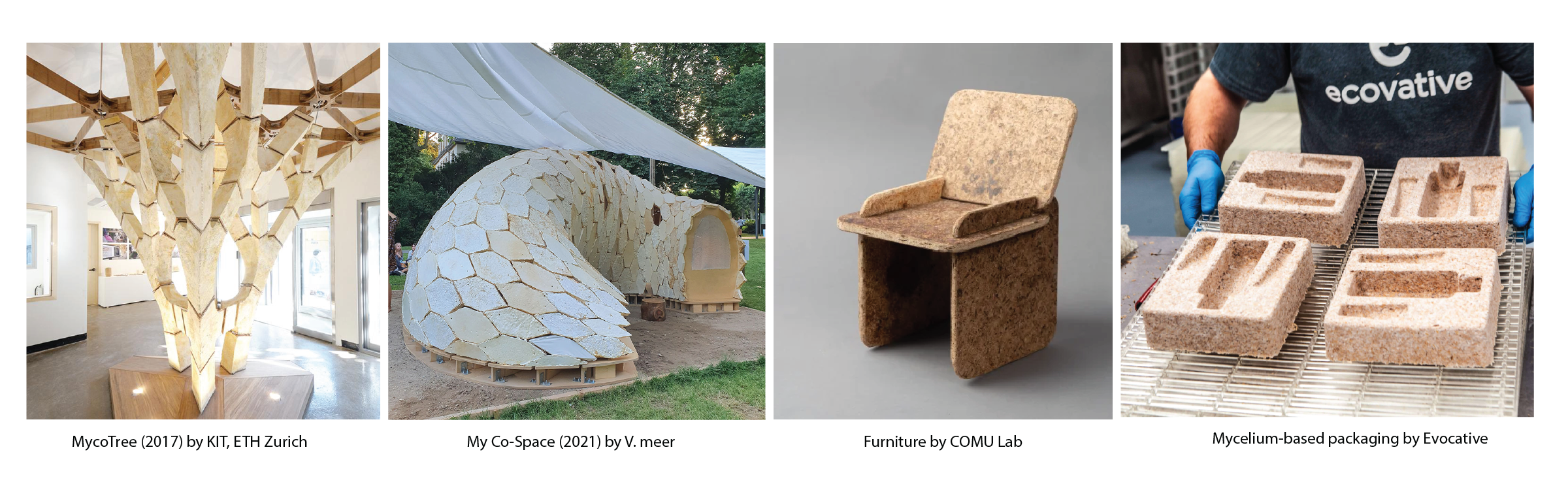


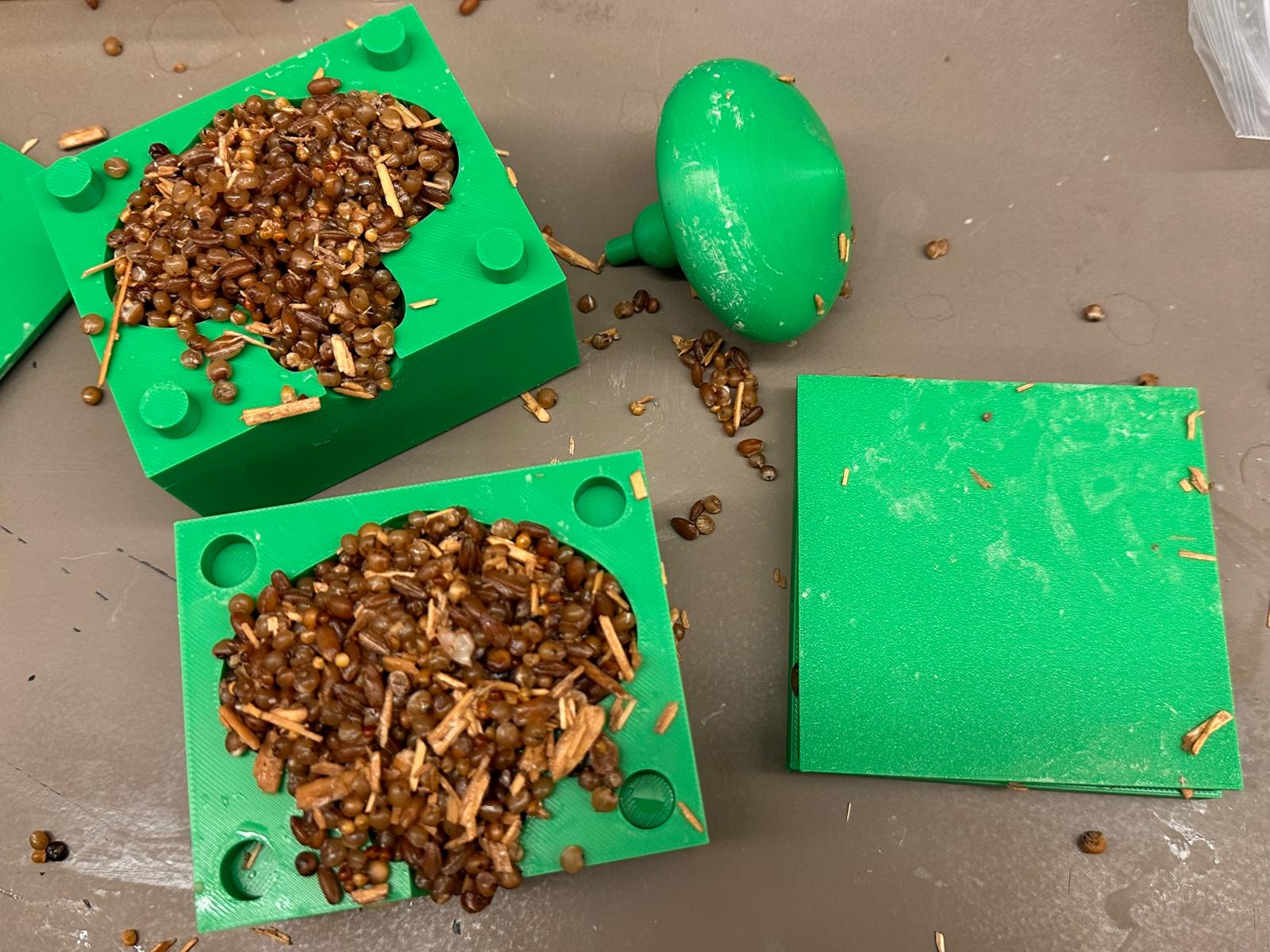
Lab last week: I 3D printed a negative mold of a small mushroom geometry to cast mycelium…


What might you want to genetically engineer fungi to do and why? What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
One reason to genetically engineer fungi is to gain more control over how they grow and behave as materials. For instance, they could be designed to respond to environmental conditions, such as changing stiffness, color, or triggering self-repair when damaged. It would also be useful to control their growth patterns, since the way mycelium branches and densifies directly affects the strength and structure of the material. There is also potential to introduce functions like conductivity or sensing, allowing fungal systems to act as responsive or interactive materials rather than purely passive ones. Overall, genetic engineering would make fungal materials more consistent and tunable instead of relying on natural variability.
Compared to bacteria, fungi are better suited for material applications because they naturally form filamentous networks that grow into three-dimensional structures. They can bind substrates and create solid composites as they grow, which aligns more directly with fabrication. Their cell walls also provide more mechanical strength, making them more useful for structural uses. While bacteria are faster and easier to engineer, they are typically limited to producing molecules in liquid systems. Fungi, on the other hand, are more relevant when the goal is to grow materials with form and function at larger scales.
Final Project Proposal
Fabric-Based Cell-Free Biosensors for Heavy Metal Detection This project develops a distributed sensing system that integrates textiles, capillary-driven microfluidics, and cell-free synthetic biology to detect environmental toxins in wetland-adjacent conditions. Embroidered fabric pathways guide fluid transport through capillary action, while hydrogel inclusions regulate flow, timing, and exposure, enabling passive operation without external pumping or electronics.
Localized sensing regions contain cell-free gene expression systems with DNA constructs responsive to heavy metals. Lead (Pb²⁺) detection is mediated by the PbrR regulatory protein, which activates transcription in the presence of lead ions. Upon activation, the system expresses a chromoprotein reporter, producing a visible color change directly within the textile.
By combining fluid transport, material structure, and biochemical response, the fabric functions as a distributed sensing network that translates invisible contamination into persistent visual patterns. Designed for intermittent exposure such as stormwater runoff and wetland edges, this approach enables deployable, low-cost environmental monitoring through responsive textiles and landscape-integrated installations.
AIM 1: Design and implement a cell-free, lead-responsive genetic circuit embedded within a textile-based microfluidic system to produce a visible chromoprotein signal upon exposure to Pb²⁺.
This aim focuses on integrating a PbrR-regulated promoter system with a chromoprotein reporter in a cell-free format, enabling localized, on-site detection of heavy metal contamination without the use of living cells.