Week 2 HW: DNA read-write-edit
Part 1: Gel Electrophoresis
Due to no access to equipment and space for gel electrophoresis I simulated the same to understand the process on https://www.labxchange.org/library/items/lb:LabXchange:9548bee3:lx_simulation:1?fullscreen=true
 Workflow
Design plasmid DNA with protein of interest →Transform bacteria with plasmid DNA→Get many copies of plasmid DNA→introduction of plasmid DNA to cells
Workflow
Design plasmid DNA with protein of interest →Transform bacteria with plasmid DNA→Get many copies of plasmid DNA→introduction of plasmid DNA to cells
Working in Benchling
After signing in I imported it into Benching and ran digests for EcoRI HindIII BamHI KpnI EcoRV SacI SalI
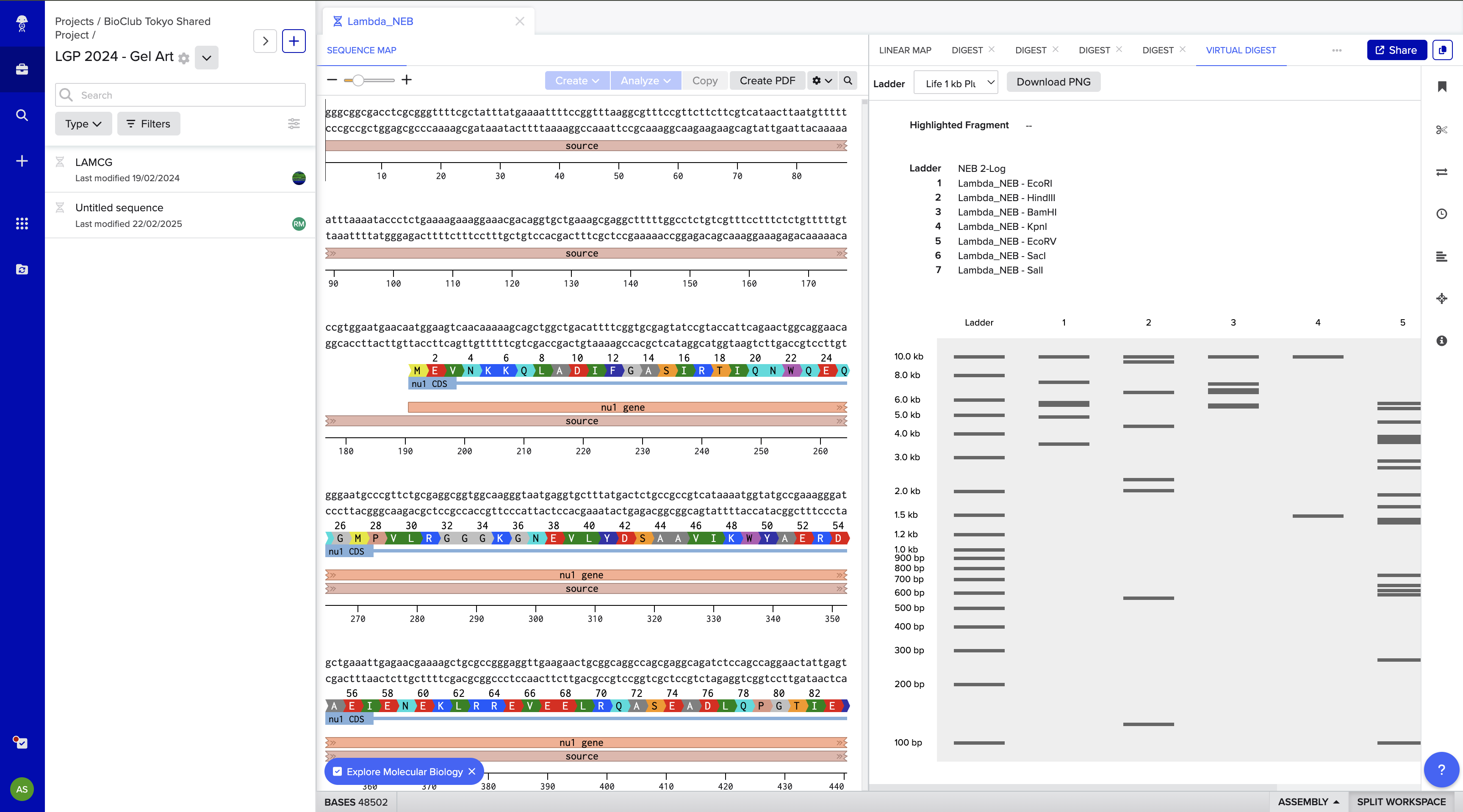

And then ran digests on SalI SacI BamHI KpnI EcoRV BamHI KpnI SacI SalI
to create an Elephant! 🐘
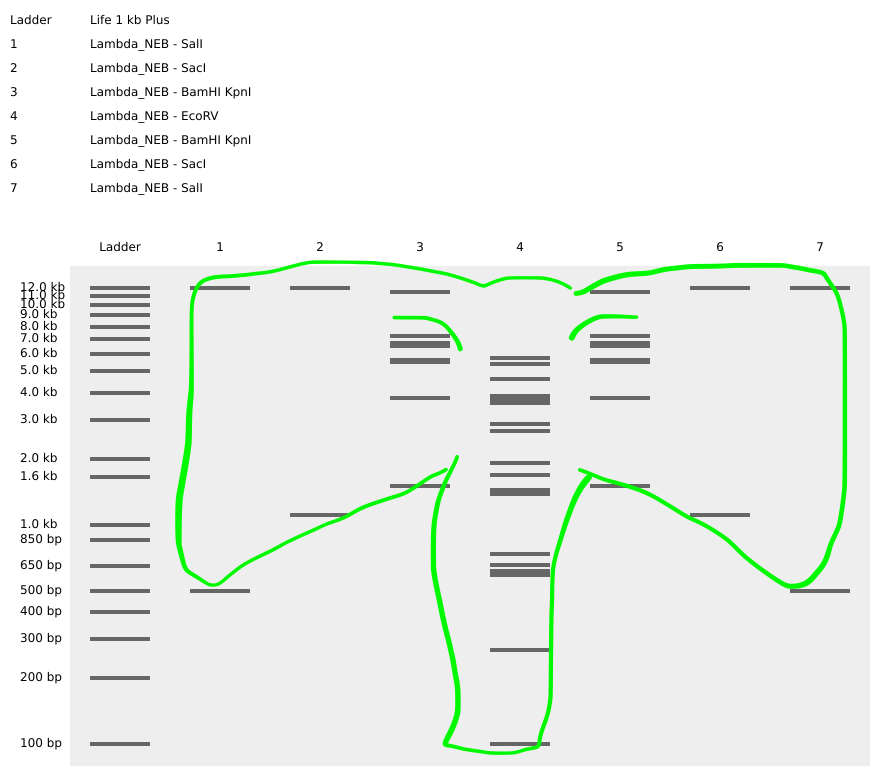

For this, I referred to an iGem video to understand how enzyme digesting works as well https://www.youtube.com/watch?v=7cGev-SKLao
DNA design challenge
Chosen protein : Actin
tr|D3BD07|D3BD07_HETP5 Actin OS=Heterostelium pallidum (strain ATCC 26659 / Pp 5 / PN500) OX=670386 GN=act10 PE=3 SV=1 MEGEDVQALVIDNGSGMCKAGFAGDDAPRAVFPSIVGRPRHTGVMVGMGQKDSYVGDEAQ SKRGILTLKYPIEHGIVTNWDDMEKIWHHTFYNELRVAPEEHPVLLTEAPLNPKANREKM TQIMFETFNTPAMYVAIQAVLSLYASGRTTGIVMDSGDGVSHTVPIYEGYALPHAILRLD LAGRDLTDYMMKILTERGYSFTTTAEREIVRDIKEKLAYVALDFENEMQTAASSSALEKS YELPDGQVITIGNERFRCPEALFQPSFLGMESAGIHETTYNSIMKCDVDIRKDLYGNVVL SGGTTMFPGIADRMQKELTALAPSTMKIKIIAPPERKYSVWIGGSILASLSTFQQMWISK EEYDESGPSIVHRKCF
Reverse translated
ATGGAAGGCGACGTTCAAGCGCTGGTGATCGACAATGGTTCTGGCATGTAAAGCGGGTTTCGCAGGCGACGACGCACCGCGCGCGCGCGTTCTTTCCTTCGATTGTGCGCCGTCGCCGTCATACCGGCGTGATGGTTGTGGGGATGCAGCAAGAGGACTCCTACGTGGGCGACGAGGCGCAGTCGAAAGGTGGGATCCTGACCCTGAAGTACCCGATCGAACACGGGATTGGTGACTAACAATGGGACGATATGAAGGAAATCTGGCACCACACGTTCTTATAACGAATTAAGAGTGGCGCCGGAAGAACCAGTTCCTGTGCTGCTGACCGAGGCGCCGCTGAACCCGAAAGCCAACCGTGAAGAAATGAAGACCAGGATTATGTTTGAACCTTTC AACACGCCGGCGATGTATGTGGCGATTCAAGCGGTGTTGTCGCTGTATGCCTCGGGTCGTACCACC GGTATTGTGATGGATTCTGGCGACGGCGTGTCCCATACGGTGCCCATCTATGAAGGTTATGCCTTACCGCACCGCATCCTCCGCCTGGATCTGGCGGGTCGCGATCTGACTGAC TATATGATGAAGATCCTGACTGAACGTGGTTATTCGTTTACGACCACCGCCGAAAGGGAaatcgtcgacatc aaagagaagctggcgtatgtggcacttgatttcgagaacgagatgcaaacggcggcgTCGTCGTCGTCGCGTTGAA AAGTCG TATGAACTGCCG GACGGCCAGGTCATCACTATCGGTAACGAACGTTTC CGCTGCCCTGCGCTTTCAACCGTCGTTCTTAGGCATGGAAAGCGCGGGCA TACACGAAACCACGTACAACAGCATTATGAAATGC GATGTCGACATT CGCAAGGATCTGTATGGTAACGTGGTCCTGGGCGGCACCACGATGTTCCCGGGCATCGCCGAACG CATGCAAGAAACTGACCACCGCGCTGGCGCCGTCGACCATGAAAATCAAGATCATTGCCGCGCCGGAACGTAAGTCTTGGGTCATCGGCGGC TCGTTGGCCTCGTCGACCTTC CAGCAGATG TGGATCAGCAAAGAAGAG TATGACGAAAGCGGTCCTTCGGTGATCCACCGTAAGTTCTTCGCGAAACCGCAAGATTAA
optimized codon sequence
Optimized DNA Sequence (1122 bp)
atggaaggcgatgtccaggcgctggtgatcgacaacggctccggcatgaaggccggcttcgccggcgatgatgcccccagggcggcggtgtcttcccctcgatcgtgggccgtccgcgtcacaccggtgtgatggtggtgggtatgcagcagaaagattcctatgtgggcgacgaagcgcaatcgaaagggcatcctgaccctgaagtatccgatcgagcatggcatcgtgaacaactgggacatggagaagatctggcaccacatgttctacaacgagctgcgtgtggcgccggaagaaccccacgtgctgctgaccgaggcgccgctgaacccgaaggccaaccgtgaacgcaagatgaagaccaggatcatgatgttcgaacagttcaacacgccggcgatgtatgtggcgattcaagcggtgctgtcgctgtatgcctcgggccgtaccaccggcatcgtgatggactccggcgatggcgtttcccacatcgtgcccatctatgaaggctatgcgctgccgcatgccatcctgcgcctggatctggcgggcagggatctgaccgactacatgatgaagatcctgaccgaacgcggttatagcttcaccaccaccgcggagaagatcgtccgggacatcaagaagaaactggcgtatgtggcgctcgatttcgaaaacgaaatgcaagcgaccgcgagctcgagcgccctggagaagtcgtatgagctgccggacggccaggtgatcaccatcggcaacgaacgcttccgttgccctgccgctgttccagccctcgttcggcatggagagcgccggcatccatgagaccacctacaacagcatcatgaagacctgcgatgtggacatccgcaaggacctgtatggcaacgtggtgctcggcggcaccaccatgttccccggcatcgccgacaggatgcaaaaggagctgaccgccgcgctgccgcccagcaccatgaagatcaagatcatcgcgccgccggagcgtaagtcgtgggtgatcggcggctcgctggcgagcctgagcacgttccagcagatgtggatcagcaaggaggaatacgacgagtcgggcccgagcatcgtgcaccgcaagtgcttcggcaagcgcaagatgaa
I would recommend a plasmid-based cloning approach for initial expression work. The optimized DNA can be inserted into a standard expression plasmid and introduced into E. coli via transformation. Once inside the bacterial cells, the plasmid replicates autonomously, allowing the host machinery to transcribe my DNA into mRNA and subsequently translate it into actin protein. However, since actin is a eukaryotic cytoskeletal protein and my sequence lacks the signal peptides and targeting sequences necessary for membrane localization or secretion, the expressed protein will likely accumulate intracellularly. This necessitates cell lysis and downstream purification via affinity chromatography or other protein separation techniques to isolate and characterize my recombinant actin. Alternatively, the PURE system (Protein synthesis Using Recombinant Elements) presents a compelling option due to its turnaround time. In this cell-free approach, my DNA template is incubated with a defined set of recombinant enzymes and cellular extracts that provide the necessary transcriptional and translational machinery. This in vitro reaction proceeds rapidly without the overhead of maintaining living cells, generating my actin protein directly in the reaction mixture. The resulting product must subsequently be purified via affinity chromatography to obtain homogeneous, functional protein suitable for my downstream biochemical investigations.
DNA synthesis order
I want to use Green Fluorescent Protein because it is a good medium to understand and track other proteins. Physarum polycephalum has actin and myosis predominantly, and to understand the movements within the Physarum tubes, fluorescence can help.
Protein sequnece copied from UnitProt:
sp|P42212|GFP_AEQVI Green fluorescent protein OS=Aequorea victoria OX=6100 GN=GFP PE=1 SV=1 MSKGEELFTGVVPILVELDGDVNGHKFSVSGEGEGDATYGKLTLKFICTTGKLPVPWPTL VTTFSYGVQCFSRYPDHMKQHDFFKSAMPEGYVQERTIFFKDDGNYKTRAEVKFEGDTLV NRIELKGIDFKEDGNILGHKLEYNYNSHNVYIMADKQKNGIKVNFKIRHNIEDGSVQLAD HYQQNTPIGDGPVLLPDNHYLSTQSALSKDPNEKRDHMVLLEFVTAAGITHGMDELYK
Translating AA (amino acid) sequence to DNA sequence: (using https://www.bioinformatics.org/sms2/rev_trans.html)
reverse translation of sp|P42212|GFP_AEQVI Green fluorescent protein OS=Aequorea victoria OX=6100 GN=GFP PE=1 SV=1 to a 714 base sequence of most likely codons. atgagcaaaggcgaagaactgtttaccggcgtggtgccgattctggtggaactggatggc gatgtgaacggccataaatttagcgtgagcggcgaaggcgaaggcgatgcgacctatggc aaactgaccctgaaatttatttgcaccaccggcaaactgccggtgccgtggccgaccctg gtgaccacctttagctatggcgtgcagtgctttagccgctatccggatcatatgaaacag catgatttttttaaaagcgcgatgccggaaggctatgtgcaggaacgcaccatttttttt aaagatgatggcaactataaaacccgcgcggaagtgaaatttgaaggcgataccctggtg aaccgcattgaactgaaaggcattgattttaaagaagatggcaacattctgggccataaa ctggaatataactataacagccataacgtgtatattatggcggataaacagaaaaacggc attaaagtgaactttaaaattcgccataacattgaagatggcagcgtgcagctggcggat cattatcagcagaacaccccgattggcgatggcccggtgctgctgccggataaccattat ctgagcacccagagcgcgctgagcaaagatccgaacgaaaaacgcgatcatatggtgctg ctggaatttgtgaccgcggcgggcattacccatggcatggatgaactgtataaa