Week 10 HW: Imaging and Measurement
Waters Part I Molecular Weight
Question 1: Based on the predicted amino acid sequence of eGFP (see below) and any known modifications, what is the calculated molecular weight?
Using the ExPASy Compute pI/Mw tool with the provided eGFP sequence
Theoretical MW = 28,006.60 Da
Question 2: Calculate the molecular weight of the eGFP using the adjacent charge state approach described in the recitation.
Using the two adjacent charge state peaks
- $m/z_n = 875.4421$
- $m/z_{n+1} = 848.9758$
Using the first peak ($z = 32$):
MW = 32 \times (875.4421 - 1.0073) = 32 \times 874.4348 \approx 27{,}981.9 \text{ Da}Confirming with the second peak ($z = 33$):
MW = 33 \times (848.9758 - 1.0073) = 33 \times 847.9685 \approx 27{,}982.0 \text{ Da}Deconvoluted MW ≈ 27,982 Da
Step 2c: Measurement Accuracy
$$\text{Accuracy} = \frac{|MW_{\text{experiment}} - MW_{\text{theory}}|}{MW_{\text{theory}}} = \frac{|27{,}982 - 28{,}006|}{28{,}006} = \frac{24}{28{,}006} \approx 8.5 \times 10^{-4} \approx 0.085%$$
Question 3: Can you observe the charge state for the zoomed-in peak in the mass spectrum for the intact eGFP? If yes, what is it? If no, why not?
No, the charge state cannot be determined from the zoomed-in peak. Determining the charge state requires at least two adjacent charge-state peaks so their spacing can be used to calculate $z$. In the zoomed region, only a single isolated peak is shown with no neighboring charge-state peak visible, so there is insufficient information to assign a charge state.
Waters Part III — Peptide Mapping (Primary Structure)
Q1.How many Lysines (K) and Arginines (R) are in eGFP? Please circle or highlight them in the eGFP sequence given in Waters Part I question 1 above.
There are 20 Lysines (K) and 6 Arginines (R) in the eGFP sequence, for a total of 26 cleavage sites.
Highlighted sequence (K and R in bold):
MVSKGEELFTGVVPILVELDGDVNGHKFSVSGEGEGDATYGKLTLKFICTTGKLPVPWPTLVTTLTYGVQCFSRYPDHMKQHDFFKSAMPEGYVQERTIFFKDDGNYKTRAEVKFEGDTLVNRIELKGIDFKEDGNILGHKLEYNYNSHNVYIMADKQKNGIKVNFKIRHNIEDGSVQLADHYQQNTPIGDGPVLLPDNHYLSTQSALSKDPNEK****RDHMVLLEFVTAAGITLGMDELYKLEHHHHHH
Q2. How many peptides will be generated from tryptic digestion of eGFP?
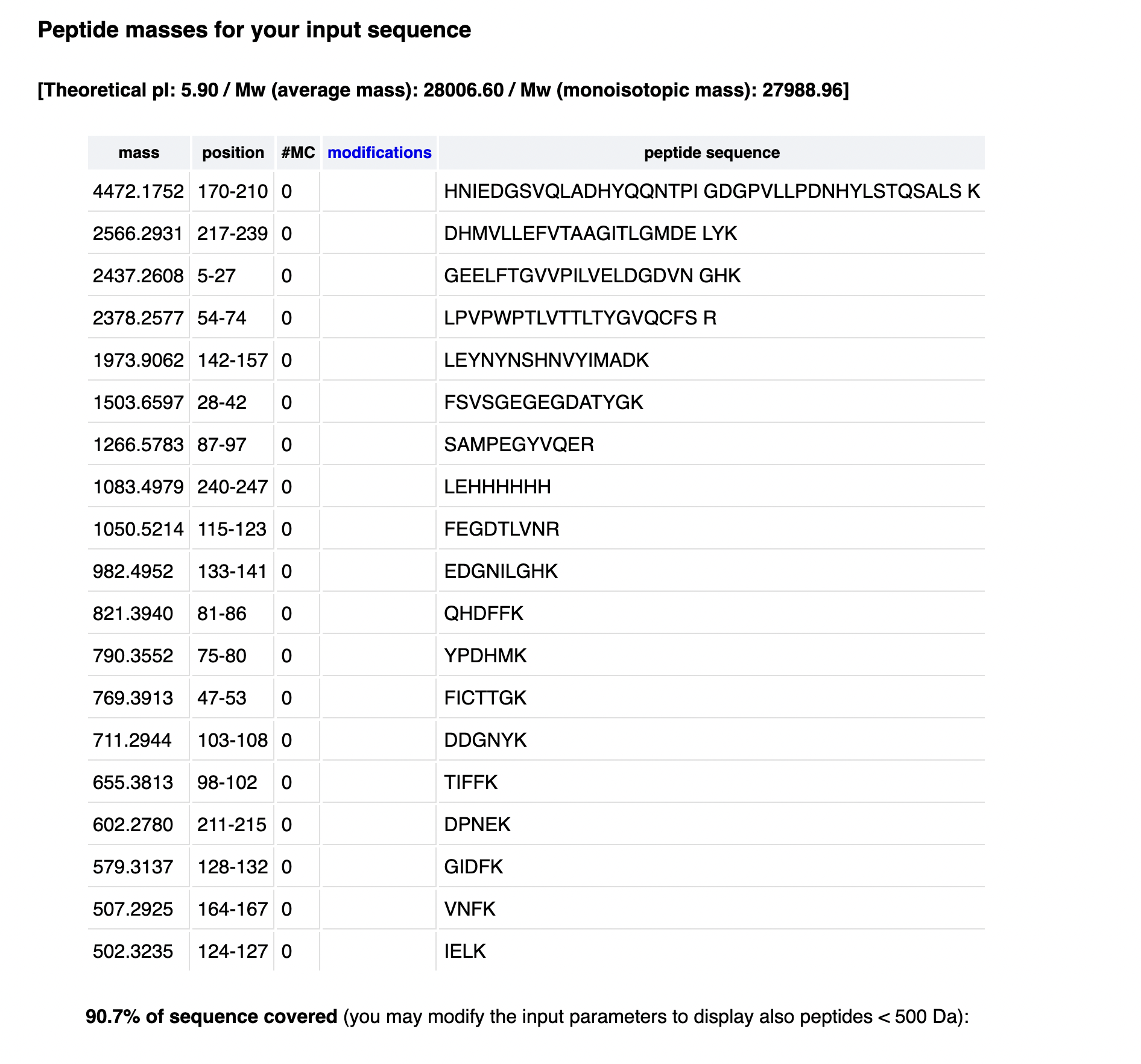
Using the PeptideMass tool at https://web.expasy.org/peptide_mass/ with the eGFP sequence, trypsin as enzyme, no missed cleavages, and the parameters shown in Figure 4, the tool generates 19 tryptic peptides.

Q3. Based on the LC-MS data for the Peptide Map data generated in lab (please use Figure 5a as a reference) how many chromatographic peaks do you see in the eGFP peptide map between 0.5 and 6 minutes? You may count all peaks that are >10% relative abundance.
There are around 19 peaks between .5 and 6 minutes
Q4. Assuming all the peaks are peptides, does the number of peaks match the number of peptides predicted from question 2 above? Are there more peaks in the chromatogram or fewer?
Accounting for all peaks the total would be 22, more than the predicted 19 peaks
Q5. Identify the mass-to-charge of the peptide shown in Figure 5b. What is the charge of the most abundant charge state of the peptide (use the separation of the isotopes to determine the charge state). Calculate the mass of the singly charged form of the peptide ([M+H])+ based on its m/z and z.
The highlighted peak shows the most abundant isotope at m/z = 2.78 (the apex of the green-circled envelope).
In TOF-MS, the isotope spacing reveals the charge state. The relationship is: \text{Isotope spacing} = \frac{1}{z}
Looking at the isotope pattern around the peak at retention time ~2.7 min, you can see fine structure. The isotope spacing (distance between consecutive 13C isotopes) is approximately 0.33-0.35 m/z units.
z = \frac{1}{\text{isotope spacing}} = \frac{1}{0.33} \approx 3 The charge state is z = 3 (triply charged ion, [M+3H]³⁺)
Using the relationship between observed m/z, charge state, and molecular mass: m/z = \frac{M + nH}{z} Rearranging to solve for M: M = (m/z \times z) - nH Where n = z (number of protons added). For z = 3: M = (2.78 \times 3) - (3 \times 1.0073) = 8.34 - 3.0219 = 5.3181 \text{ kDa} \approx 5.32 \text{ kDa} The singly charged form [M+H]⁺ would have m/z equal to the neutral mass plus one proton: [M+H]^+ = 5318.1 + 1.0073 = 5319.1 \text{ m/z} \approx 5319 \text{ Da}
Q6. Identify the peptide based on comparison to expected masses in the PeptideMass tool. What is mass accuracy of measurement? Please calculate the error in ppm.
- Observed peptide mass (MW_experiment): 1050.52438 Da
- Theoretical peptide mass (MW_theory): 1050.5214 Da
The mass accuracy is calculated using the absolute difference between experimental and theoretical mass values, normalized to the theoretical mass:
\text{Accuracy} = \frac{|MW_{\text{experiment}} - MW_{\text{theory}}|}{MW_{\text{theory}}}Substituting the measured values:
\text{Accuracy} = \frac{|1050.52438 - 1050.5214|}{1050.5214} \text{Accuracy} = \frac{0.00298}{1050.5214} = 2.84 \times 10^{-6}Error in ppm
To express the accuracy as a ppm error, multiply the accuracy by 10^6:
\text{ppm error} = \text{Accuracy} \times 10^6 \text{ppm error} = 2.84 \times 10^{-6} \times 10^6 = 2.84 \text{ ppm}Final Answer
- Observed peptide mass: 1050.52438 Da
- Closest predicted peptide mass: 1050.5214 Da
- Mass error: 2.84 ppm
- Assessment: This error is well below the <10 ppm threshold, indicating excellent measurement accuracy for high-resolution mass spectrometry
Q7. Number of peptides from tryptic digestion
88 percent
Homework: Waters Part IV — Oligomers
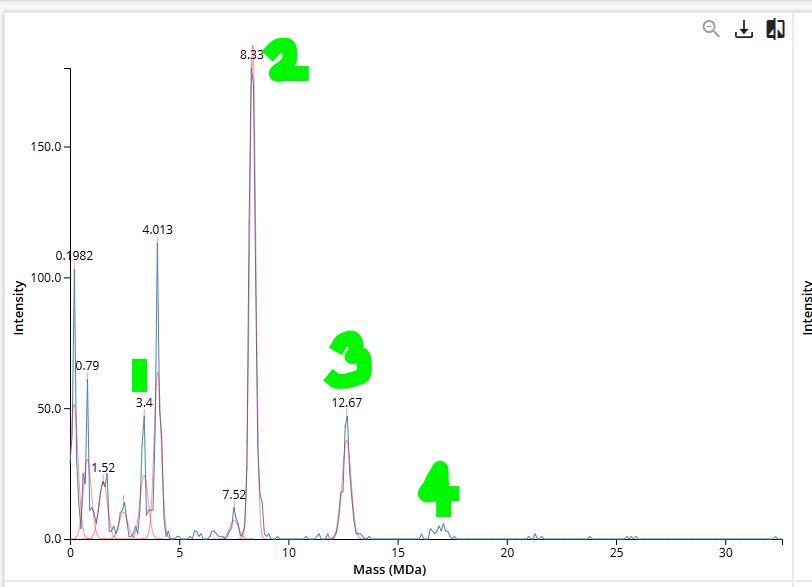
- 7FU Decamer: Mass = 10 x 340 kDa = 3400 kDa (3.4 (MDa))
- 8FU Didecamer: Mass = 20 x 400 kDa = 8000 kDa (8 (MDa))
- 8FU 3-Decamer: Mass = 30 x 400 kDa = 3400 kDa (12 (MDa))
- 8FU 4-Decamer: Mass = 40 x 400 kDa = 3400 kDa (16 (MDa))

Homework: Waters Part V — Did I make GFP?
| Parameter | Theoretical (Da) | Observed (Da) | PPM Mass Error |
|---|