from opentrons import types
metadata = {
'author': 'Beyza Batır',
'protocolName': 'HTGAA Opentrons Lab',
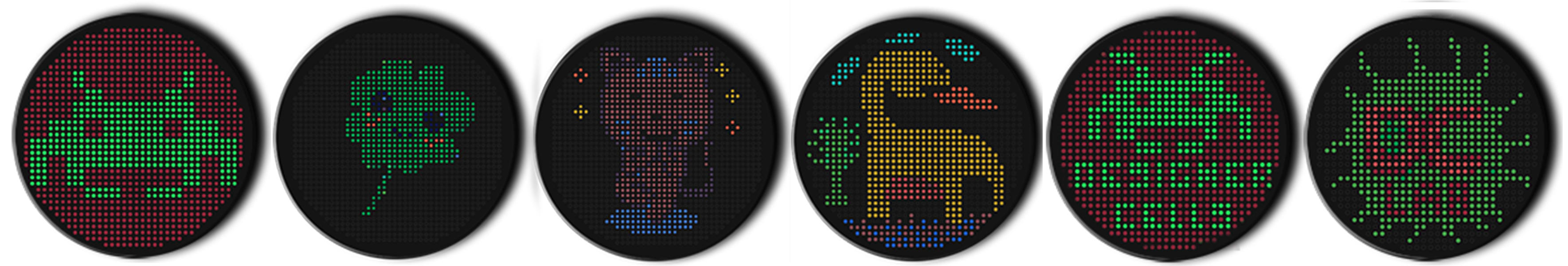
'description': 'SpaceInvaders',
'source': 'HTGAA 2026 Opentrons Lab',
'apiLevel': '2.20'
}
TIP_RACK_DECK_SLOT = 9
COLORS_DECK_SLOT = 6
AGAR_DECK_SLOT = 5
PIPETTE_STARTING_TIP_WELL = 'A1'
well_colors = {
'A1': 'Red',
'B1': 'Green',
}
def run(protocol):
# --- Load labware ---
tips_20ul = protocol.load_labware('opentrons_96_tiprack_20ul', TIP_RACK_DECK_SLOT, 'Opentrons 20uL Tips')
pipette_20ul = protocol.load_instrument("p20_single_gen2", "right", [tips_20ul])
temperature_module = protocol.load_module('temperature module gen2', COLORS_DECK_SLOT)
temperature_plate = temperature_module.load_labware('opentrons_96_aluminumblock_generic_pcr_strip_200ul', 'Cold Plate')
color_plate = temperature_plate
agar_plate = protocol.load_labware('htgaa_agar_plate', AGAR_DECK_SLOT, 'Agar Plate')
center_location = agar_plate['A1'].top()
pipette_20ul.starting_tip = tips_20ul.well(PIPETTE_STARTING_TIP_WELL)
# --- Helper functions ---
def location_of_color(color_string):
for well, color in well_colors.items():
if color.lower() == color_string.lower():
return color_plate[well]
raise ValueError(f"No well found with color {color_string}")
def dispense_and_detach(pipette, volume, location):
above_location = location.move(types.Point(z=5))
pipette.move_to(above_location)
pipette.dispense(volume, location)
pipette.move_to(above_location)
# --- Coordinates from GUI ---
mrfp1_points = [(-9.9, 38.5),(-7.7, 38.5),(-5.5, 38.5),(-3.3, 38.5),(-1.1, 38.5),(1.1, 38.5),(3.3, 38.5),(5.5, 38.5),(7.7, 38.5),(9.9, 38.5),(-16.5, 36.3),(-14.3, 36.3),(-12.1, 36.3),(-9.9, 36.3),(-7.7, 36.3),(-5.5, 36.3),(-3.3, 36.3),(-1.1, 36.3),(1.1, 36.3),(3.3, 36.3),(5.5, 36.3),(7.7, 36.3),(9.9, 36.3),(12.1, 36.3),(14.3, 36.3),(16.5, 36.3),(-20.9, 34.1),(-18.7, 34.1),(-16.5, 34.1),(-14.3, 34.1),(-12.1, 34.1),(-9.9, 34.1),(-7.7, 34.1),(-5.5, 34.1),(-3.3, 34.1),(-1.1, 34.1),(1.1, 34.1),(3.3, 34.1),(5.5, 34.1),(7.7, 34.1),(9.9, 34.1),(12.1, 34.1),(14.3, 34.1),(16.5, 34.1),(18.7, 34.1),(20.9, 34.1),(-23.1, 31.9),(-20.9, 31.9),(-18.7, 31.9),(-16.5, 31.9),(-14.3, 31.9),(-12.1, 31.9),(-9.9, 31.9),(-7.7, 31.9),(-5.5, 31.9),(-3.3, 31.9),(-1.1, 31.9),(1.1, 31.9),(3.3, 31.9),(5.5, 31.9),(7.7, 31.9),(9.9, 31.9),(12.1, 31.9),(14.3, 31.9),(16.5, 31.9),(18.7, 31.9),(20.9, 31.9),(23.1, 31.9),(-25.3, 29.7),(-23.1, 29.7),(-20.9, 29.7),(-18.7, 29.7),(-16.5, 29.7),(-14.3, 29.7),(-12.1, 29.7),(-9.9, 29.7),(-7.7, 29.7),(-5.5, 29.7),(-3.3, 29.7),(-1.1, 29.7),(1.1, 29.7),(3.3, 29.7),(5.5, 29.7),(7.7, 29.7),(9.9, 29.7),(12.1, 29.7),(14.3, 29.7),(16.5, 29.7),(18.7, 29.7),(20.9, 29.7),(23.1, 29.7),(25.3, 29.7),(-27.5, 27.5),(-25.3, 27.5),(-23.1, 27.5),(-20.9, 27.5),(-18.7, 27.5),(-16.5, 27.5),(-14.3, 27.5),(-12.1, 27.5),(-9.9, 27.5),(-7.7, 27.5),(-5.5, 27.5),(-3.3, 27.5),(-1.1, 27.5),(1.1, 27.5),(3.3, 27.5),(5.5, 27.5),(7.7, 27.5),(9.9, 27.5),(12.1, 27.5),(14.3, 27.5),(16.5, 27.5),(18.7, 27.5),(20.9, 27.5),(23.1, 27.5),(25.3, 27.5),(27.5, 27.5),(-29.7, 25.3),(-27.5, 25.3),(-25.3, 25.3),(-23.1, 25.3),(-20.9, 25.3),(-18.7, 25.3),(-16.5, 25.3),(-14.3, 25.3),(-12.1, 25.3),(-9.9, 25.3),(-7.7, 25.3),(-5.5, 25.3),(-3.3, 25.3),(-1.1, 25.3),(1.1, 25.3),(3.3, 25.3),(5.5, 25.3),(7.7, 25.3),(9.9, 25.3),(12.1, 25.3),(14.3, 25.3),(16.5, 25.3),(18.7, 25.3),(20.9, 25.3),(23.1, 25.3),(25.3, 25.3),(27.5, 25.3),(29.7, 25.3),(-31.9, 23.1),(-29.7, 23.1),(-27.5, 23.1),(-25.3, 23.1),(-23.1, 23.1),(-16.5, 23.1),(-14.3, 23.1),(-12.1, 23.1),(-9.9, 23.1),(-7.7, 23.1),(-5.5, 23.1),(-3.3, 23.1),(-1.1, 23.1),(1.1, 23.1),(3.3, 23.1),(5.5, 23.1),(7.7, 23.1),(9.9, 23.1),(12.1, 23.1),(14.3, 23.1),(16.5, 23.1),(23.1, 23.1),(25.3, 23.1),(27.5, 23.1),(29.7, 23.1),(31.9, 23.1),(-34.1, 20.9),(-31.9, 20.9),(-29.7, 20.9),(-27.5, 20.9),(-25.3, 20.9),(-23.1, 20.9),(-16.5, 20.9),(-14.3, 20.9),(-12.1, 20.9),(-9.9, 20.9),(-7.7, 20.9),(-5.5, 20.9),(-3.3, 20.9),(-1.1, 20.9),(1.1, 20.9),(3.3, 20.9),(5.5, 20.9),(7.7, 20.9),(9.9, 20.9),(12.1, 20.9),(14.3, 20.9),(16.5, 20.9),(23.1, 20.9),(25.3, 20.9),(27.5, 20.9),(29.7, 20.9),(31.9, 20.9),(34.1, 20.9),(-34.1, 18.7),(-31.9, 18.7),(-29.7, 18.7),(-27.5, 18.7),(-25.3, 18.7),(-23.1, 18.7),(-16.5, 18.7),(-14.3, 18.7),(-12.1, 18.7),(-9.9, 18.7),(-7.7, 18.7),(-5.5, 18.7),(-3.3, 18.7),(-1.1, 18.7),(1.1, 18.7),(3.3, 18.7),(5.5, 18.7),(7.7, 18.7),(9.9, 18.7),(12.1, 18.7),(14.3, 18.7),(16.5, 18.7),(23.1, 18.7),(25.3, 18.7),(27.5, 18.7),(29.7, 18.7),(31.9, 18.7),(34.1, 18.7),(-36.3, 16.5),(-34.1, 16.5),(-31.9, 16.5),(-29.7, 16.5),(-27.5, 16.5),(-25.3, 16.5),(-23.1, 16.5),(-20.9, 16.5),(-18.7, 16.5),(-9.9, 16.5),(-7.7, 16.5),(-5.5, 16.5),(-3.3, 16.5),(-1.1, 16.5),(1.1, 16.5),(3.3, 16.5),(5.5, 16.5),(7.7, 16.5),(9.9, 16.5),(18.7, 16.5),(20.9, 16.5),(23.1, 16.5),(25.3, 16.5),(27.5, 16.5),(29.7, 16.5),(31.9, 16.5),(34.1, 16.5),(36.3, 16.5),(-36.3, 14.3),(-34.1, 14.3),(-31.9, 14.3),(-29.7, 14.3),(-27.5, 14.3),(-25.3, 14.3),(-23.1, 14.3),(-20.9, 14.3),(-18.7, 14.3),(-9.9, 14.3),(-7.7, 14.3),(-5.5, 14.3),(-3.3, 14.3),(-1.1, 14.3),(1.1, 14.3),(3.3, 14.3),(5.5, 14.3),(7.7, 14.3),(9.9, 14.3),(18.7, 14.3),(20.9, 14.3),(23.1, 14.3),(25.3, 14.3),(27.5, 14.3),(29.7, 14.3),(31.9, 14.3),(34.1, 14.3),(36.3, 14.3),(-36.3, 12.1),(-34.1, 12.1),(-31.9, 12.1),(-29.7, 12.1),(-27.5, 12.1),(-25.3, 12.1),(-23.1, 12.1),(23.1, 12.1),(25.3, 12.1),(27.5, 12.1),(29.7, 12.1),(31.9, 12.1),(34.1, 12.1),(36.3, 12.1),(-38.5, 9.9),(-36.3, 9.9),(-34.1, 9.9),(-31.9, 9.9),(-29.7, 9.9),(-27.5, 9.9),(-25.3, 9.9),(-23.1, 9.9),(23.1, 9.9),(25.3, 9.9),(27.5, 9.9),(29.7, 9.9),(31.9, 9.9),(34.1, 9.9),(36.3, 9.9),(38.5, 9.9),(-38.5, 7.7),(-36.3, 7.7),(-34.1, 7.7),(-31.9, 7.7),(-29.7, 7.7),(-27.5, 7.7),(-25.3, 7.7),(-23.1, 7.7),(23.1, 7.7),(25.3, 7.7),(27.5, 7.7),(29.7, 7.7),(31.9, 7.7),(34.1, 7.7),(36.3, 7.7),(38.5, 7.7),(-38.5, 5.5),(-36.3, 5.5),(-34.1, 5.5),(-31.9, 5.5),(-16.5, 5.5),(-14.3, 5.5),(-12.1, 5.5),(12.1, 5.5),(14.3, 5.5),(16.5, 5.5),(31.9, 5.5),(34.1, 5.5),(36.3, 5.5),(38.5, 5.5),(-38.5, 3.3),(-36.3, 3.3),(-34.1, 3.3),(-31.9, 3.3),(-16.5, 3.3),(-14.3, 3.3),(-12.1, 3.3),(12.1, 3.3),(14.3, 3.3),(16.5, 3.3),(31.9, 3.3),(34.1, 3.3),(36.3, 3.3),(38.5, 3.3),(-38.5, 1.1),(-36.3, 1.1),(-34.1, 1.1),(-31.9, 1.1),(-16.5, 1.1),(-14.3, 1.1),(-12.1, 1.1),(12.1, 1.1),(14.3, 1.1),(16.5, 1.1),(31.9, 1.1),(34.1, 1.1),(36.3, 1.1),(38.5, 1.1),(-38.5, -1.1),(38.5, -1.1),(-38.5, -3.3),(38.5, -3.3),(-38.5, -5.5),(38.5, -5.5),(-38.5, -7.7),(-29.7, -7.7),(-27.5, -7.7),(-25.3, -7.7),(25.3, -7.7),(27.5, -7.7),(29.7, -7.7),(38.5, -7.7),(-38.5, -9.9),(-29.7, -9.9),(-27.5, -9.9),(-25.3, -9.9),(25.3, -9.9),(27.5, -9.9),(29.7, -9.9),(38.5, -9.9),(-29.7, -12.1),(-27.5, -12.1),(-25.3, -12.1),(-16.5, -12.1),(-14.3, -12.1),(-12.1, -12.1),(-9.9, -12.1),(-7.7, -12.1),(-5.5, -12.1),(-3.3, -12.1),(-1.1, -12.1),(1.1, -12.1),(3.3, -12.1),(5.5, -12.1),(7.7, -12.1),(9.9, -12.1),(12.1, -12.1),(14.3, -12.1),(16.5, -12.1),(25.3, -12.1),(27.5, -12.1),(29.7, -12.1),(-29.7, -14.3),(-27.5, -14.3),(-25.3, -14.3),(-16.5, -14.3),(-14.3, -14.3),(-12.1, -14.3),(-9.9, -14.3),(-7.7, -14.3),(-5.5, -14.3),(-3.3, -14.3),(-1.1, -14.3),(1.1, -14.3),(3.3, -14.3),(5.5, -14.3),(7.7, -14.3),(9.9, -14.3),(12.1, -14.3),(14.3, -14.3),(16.5, -14.3),(25.3, -14.3),(27.5, -14.3),(29.7, -14.3),(-29.7, -16.5),(-27.5, -16.5),(-25.3, -16.5),(-16.5, -16.5),(-14.3, -16.5),(-12.1, -16.5),(-9.9, -16.5),(-7.7, -16.5),(-5.5, -16.5),(-3.3, -16.5),(-1.1, -16.5),(1.1, -16.5),(3.3, -16.5),(5.5, -16.5),(7.7, -16.5),(9.9, -16.5),(12.1, -16.5),(14.3, -16.5),(16.5, -16.5),(25.3, -16.5),(27.5, -16.5),(29.7, -16.5),(-34.1, -18.7),(-31.9, -18.7),(-29.7, -18.7),(-27.5, -18.7),(-25.3, -18.7),(-23.1, -18.7),(-20.9, -18.7),(-18.7, -18.7),(-3.3, -18.7),(-1.1, -18.7),(1.1, -18.7),(3.3, -18.7),(18.7, -18.7),(20.9, -18.7),(23.1, -18.7),(25.3, -18.7),(27.5, -18.7),(29.7, -18.7),(31.9, -18.7),(34.1, -18.7),(-34.1, -20.9),(-31.9, -20.9),(-29.7, -20.9),(-27.5, -20.9),(-25.3, -20.9),(-23.1, -20.9),(-20.9, -20.9),(-18.7, -20.9),(-3.3, -20.9),(-1.1, -20.9),(1.1, -20.9),(3.3, -20.9),(18.7, -20.9),(20.9, -20.9),(23.1, -20.9),(25.3, -20.9),(27.5, -20.9),(29.7, -20.9),(31.9, -20.9),(34.1, -20.9),(-31.9, -23.1),(-29.7, -23.1),(-27.5, -23.1),(-25.3, -23.1),(-23.1, -23.1),(-20.9, -23.1),(-18.7, -23.1),(-16.5, -23.1),(-14.3, -23.1),(-12.1, -23.1),(-9.9, -23.1),(-7.7, -23.1),(-5.5, -23.1),(-3.3, -23.1),(-1.1, -23.1),(1.1, -23.1),(3.3, -23.1),(5.5, -23.1),(7.7, -23.1),(9.9, -23.1),(12.1, -23.1),(14.3, -23.1),(16.5, -23.1),(18.7, -23.1),(20.9, -23.1),(23.1, -23.1),(25.3, -23.1),(27.5, -23.1),(29.7, -23.1),(31.9, -23.1),(-29.7, -25.3),(-27.5, -25.3),(-25.3, -25.3),(-23.1, -25.3),(-20.9, -25.3),(-18.7, -25.3),(-16.5, -25.3),(-14.3, -25.3),(-12.1, -25.3),(-9.9, -25.3),(-7.7, -25.3),(-5.5, -25.3),(-3.3, -25.3),(-1.1, -25.3),(1.1, -25.3),(3.3, -25.3),(5.5, -25.3),(7.7, -25.3),(9.9, -25.3),(12.1, -25.3),(14.3, -25.3),(16.5, -25.3),(18.7, -25.3),(20.9, -25.3),(23.1, -25.3),(25.3, -25.3),(27.5, -25.3),(29.7, -25.3),(-27.5, -27.5),(-25.3, -27.5),(-23.1, -27.5),(-20.9, -27.5),(-18.7, -27.5),(-16.5, -27.5),(-14.3, -27.5),(-12.1, -27.5),(-9.9, -27.5),(-7.7, -27.5),(-5.5, -27.5),(-3.3, -27.5),(-1.1, -27.5),(1.1, -27.5),(3.3, -27.5),(5.5, -27.5),(7.7, -27.5),(9.9, -27.5),(12.1, -27.5),(14.3, -27.5),(16.5, -27.5),(18.7, -27.5),(20.9, -27.5),(23.1, -27.5),(25.3, -27.5),(27.5, -27.5),(-25.3, -29.7),(-23.1, -29.7),(-20.9, -29.7),(-18.7, -29.7),(-16.5, -29.7),(-14.3, -29.7),(-12.1, -29.7),(-9.9, -29.7),(-7.7, -29.7),(-5.5, -29.7),(-3.3, -29.7),(-1.1, -29.7),(1.1, -29.7),(3.3, -29.7),(5.5, -29.7),(7.7, -29.7),(9.9, -29.7),(12.1, -29.7),(14.3, -29.7),(16.5, -29.7),(18.7, -29.7),(20.9, -29.7),(23.1, -29.7),(25.3, -29.7),(-23.1, -31.9),(-20.9, -31.9),(-18.7, -31.9),(-16.5, -31.9),(-14.3, -31.9),(-12.1, -31.9),(-9.9, -31.9),(-7.7, -31.9),(-5.5, -31.9),(-3.3, -31.9),(-1.1, -31.9),(1.1, -31.9),(3.3, -31.9),(5.5, -31.9),(7.7, -31.9),(9.9, -31.9),(12.1, -31.9),(14.3, -31.9),(16.5, -31.9),(18.7, -31.9),(20.9, -31.9),(23.1, -31.9),(-20.9, -34.1),(-18.7, -34.1),(-16.5, -34.1),(-14.3, -34.1),(-12.1, -34.1),(-9.9, -34.1),(-7.7, -34.1),(-5.5, -34.1),(-3.3, -34.1),(-1.1, -34.1),(1.1, -34.1),(3.3, -34.1),(5.5, -34.1),(7.7, -34.1),(9.9, -34.1),(12.1, -34.1),(14.3, -34.1),(16.5, -34.1),(18.7, -34.1),(20.9, -34.1),(-16.5, -36.3),(-14.3, -36.3),(-12.1, -36.3),(-9.9, -36.3),(-7.7, -36.3),(-5.5, -36.3),(-3.3, -36.3),(-1.1, -36.3),(1.1, -36.3),(3.3, -36.3),(5.5, -36.3),(7.7, -36.3),(9.9, -36.3),(12.1, -36.3),(14.3, -36.3),(16.5, -36.3),(-9.9, -38.5),(-7.7, -38.5),(-5.5, -38.5),(-3.3, -38.5),(-1.1, -38.5),(1.1, -38.5),(3.3, -38.5),(5.5, -38.5),(7.7, -38.5),(9.9, -38.5)]
sfgfp_points = [(-20.9, 23.1),(-18.7, 23.1),(18.7, 23.1),(20.9, 23.1),(-20.9, 20.9),(-18.7, 20.9),(18.7, 20.9),(20.9, 20.9),(-20.9, 18.7),(-18.7, 18.7),(18.7, 18.7),(20.9, 18.7),(-16.5, 16.5),(-14.3, 16.5),(-12.1, 16.5),(12.1, 16.5),(14.3, 16.5),(16.5, 16.5),(-16.5, 14.3),(-14.3, 14.3),(-12.1, 14.3),(12.1, 14.3),(14.3, 14.3),(16.5, 14.3),(-20.9, 12.1),(-18.7, 12.1),(-16.5, 12.1),(-14.3, 12.1),(-12.1, 12.1),(-9.9, 12.1),(-7.7, 12.1),(-5.5, 12.1),(-3.3, 12.1),(-1.1, 12.1),(1.1, 12.1),(3.3, 12.1),(5.5, 12.1),(7.7, 12.1),(9.9, 12.1),(12.1, 12.1),(14.3, 12.1),(16.5, 12.1),(18.7, 12.1),(20.9, 12.1),(-20.9, 9.9),(-18.7, 9.9),(-16.5, 9.9),(-14.3, 9.9),(-12.1, 9.9),(-9.9, 9.9),(-7.7, 9.9),(-5.5, 9.9),(-3.3, 9.9),(-1.1, 9.9),(1.1, 9.9),(3.3, 9.9),(5.5, 9.9),(7.7, 9.9),(9.9, 9.9),(12.1, 9.9),(14.3, 9.9),(16.5, 9.9),(18.7, 9.9),(20.9, 9.9),(-20.9, 7.7),(-18.7, 7.7),(-16.5, 7.7),(-14.3, 7.7),(-12.1, 7.7),(-9.9, 7.7),(-7.7, 7.7),(-5.5, 7.7),(-3.3, 7.7),(-1.1, 7.7),(1.1, 7.7),(3.3, 7.7),(5.5, 7.7),(7.7, 7.7),(9.9, 7.7),(12.1, 7.7),(14.3, 7.7),(16.5, 7.7),(18.7, 7.7),(20.9, 7.7),(-29.7, 5.5),(-27.5, 5.5),(-25.3, 5.5),(-23.1, 5.5),(-20.9, 5.5),(-18.7, 5.5),(-9.9, 5.5),(-7.7, 5.5),(-5.5, 5.5),(-3.3, 5.5),(-1.1, 5.5),(1.1, 5.5),(3.3, 5.5),(5.5, 5.5),(7.7, 5.5),(9.9, 5.5),(18.7, 5.5),(20.9, 5.5),(23.1, 5.5),(25.3, 5.5),(27.5, 5.5),(29.7, 5.5),(-29.7, 3.3),(-27.5, 3.3),(-25.3, 3.3),(-23.1, 3.3),(-20.9, 3.3),(-18.7, 3.3),(-9.9, 3.3),(-7.7, 3.3),(-5.5, 3.3),(-3.3, 3.3),(-1.1, 3.3),(1.1, 3.3),(3.3, 3.3),(5.5, 3.3),(7.7, 3.3),(9.9, 3.3),(18.7, 3.3),(20.9, 3.3),(23.1, 3.3),(25.3, 3.3),(27.5, 3.3),(29.7, 3.3),(-29.7, 1.1),(-27.5, 1.1),(-25.3, 1.1),(-23.1, 1.1),(-20.9, 1.1),(-18.7, 1.1),(-9.9, 1.1),(-7.7, 1.1),(-5.5, 1.1),(-3.3, 1.1),(-1.1, 1.1),(1.1, 1.1),(3.3, 1.1),(5.5, 1.1),(7.7, 1.1),(9.9, 1.1),(18.7, 1.1),(20.9, 1.1),(23.1, 1.1),(25.3, 1.1),(27.5, 1.1),(29.7, 1.1),(-36.3, -1.1),(-34.1, -1.1),(-31.9, -1.1),(-29.7, -1.1),(-27.5, -1.1),(-25.3, -1.1),(-23.1, -1.1),(-20.9, -1.1),(-18.7, -1.1),(-16.5, -1.1),(-14.3, -1.1),(-12.1, -1.1),(-9.9, -1.1),(-7.7, -1.1),(-5.5, -1.1),(-3.3, -1.1),(-1.1, -1.1),(1.1, -1.1),(3.3, -1.1),(5.5, -1.1),(7.7, -1.1),(9.9, -1.1),(12.1, -1.1),(14.3, -1.1),(16.5, -1.1),(18.7, -1.1),(20.9, -1.1),(23.1, -1.1),(25.3, -1.1),(27.5, -1.1),(29.7, -1.1),(31.9, -1.1),(34.1, -1.1),(36.3, -1.1),(-36.3, -3.3),(-34.1, -3.3),(-31.9, -3.3),(-29.7, -3.3),(-27.5, -3.3),(-25.3, -3.3),(-23.1, -3.3),(-20.9, -3.3),(-18.7, -3.3),(-16.5, -3.3),(-14.3, -3.3),(-12.1, -3.3),(-9.9, -3.3),(-7.7, -3.3),(-5.5, -3.3),(-3.3, -3.3),(-1.1, -3.3),(1.1, -3.3),(3.3, -3.3),(5.5, -3.3),(7.7, -3.3),(9.9, -3.3),(12.1, -3.3),(14.3, -3.3),(16.5, -3.3),(18.7, -3.3),(20.9, -3.3),(23.1, -3.3),(25.3, -3.3),(27.5, -3.3),(29.7, -3.3),(31.9, -3.3),(34.1, -3.3),(36.3, -3.3),(-36.3, -5.5),(-34.1, -5.5),(-31.9, -5.5),(-29.7, -5.5),(-27.5, -5.5),(-25.3, -5.5),(-23.1, -5.5),(-20.9, -5.5),(-18.7, -5.5),(-16.5, -5.5),(-14.3, -5.5),(-12.1, -5.5),(-9.9, -5.5),(-7.7, -5.5),(-5.5, -5.5),(-3.3, -5.5),(-1.1, -5.5),(1.1, -5.5),(3.3, -5.5),(5.5, -5.5),(7.7, -5.5),(9.9, -5.5),(12.1, -5.5),(14.3, -5.5),(16.5, -5.5),(18.7, -5.5),(20.9, -5.5),(23.1, -5.5),(25.3, -5.5),(27.5, -5.5),(29.7, -5.5),(31.9, -5.5),(34.1, -5.5),(36.3, -5.5),(-36.3, -7.7),(-34.1, -7.7),(-31.9, -7.7),(-23.1, -7.7),(-20.9, -7.7),(-18.7, -7.7),(-16.5, -7.7),(-14.3, -7.7),(-12.1, -7.7),(-9.9, -7.7),(-7.7, -7.7),(-5.5, -7.7),(-3.3, -7.7),(-1.1, -7.7),(1.1, -7.7),(3.3, -7.7),(5.5, -7.7),(7.7, -7.7),(9.9, -7.7),(12.1, -7.7),(14.3, -7.7),(16.5, -7.7),(18.7, -7.7),(20.9, -7.7),(23.1, -7.7),(31.9, -7.7),(34.1, -7.7),(36.3, -7.7),(-36.3, -9.9),(-34.1, -9.9),(-31.9, -9.9),(-23.1, -9.9),(-20.9, -9.9),(-18.7, -9.9),(-16.5, -9.9),(-14.3, -9.9),(-12.1, -9.9),(-9.9, -9.9),(-7.7, -9.9),(-5.5, -9.9),(-3.3, -9.9),(-1.1, -9.9),(1.1, -9.9),(3.3, -9.9),(5.5, -9.9),(7.7, -9.9),(9.9, -9.9),(12.1, -9.9),(14.3, -9.9),(16.5, -9.9),(18.7, -9.9),(20.9, -9.9),(23.1, -9.9),(31.9, -9.9),(34.1, -9.9),(36.3, -9.9),(-36.3, -12.1),(-34.1, -12.1),(-31.9, -12.1),(-23.1, -12.1),(-20.9, -12.1),(-18.7, -12.1),(18.7, -12.1),(20.9, -12.1),(23.1, -12.1),(31.9, -12.1),(34.1, -12.1),(36.3, -12.1),(-36.3, -14.3),(-34.1, -14.3),(-31.9, -14.3),(-23.1, -14.3),(-20.9, -14.3),(-18.7, -14.3),(18.7, -14.3),(20.9, -14.3),(23.1, -14.3),(31.9, -14.3),(34.1, -14.3),(36.3, -14.3),(-36.3, -16.5),(-34.1, -16.5),(-31.9, -16.5),(-23.1, -16.5),(-20.9, -16.5),(-18.7, -16.5),(18.7, -16.5),(20.9, -16.5),(23.1, -16.5),(31.9, -16.5),(34.1, -16.5),(36.3, -16.5),(-16.5, -18.7),(-14.3, -18.7),(-12.1, -18.7),(-9.9, -18.7),(-7.7, -18.7),(-5.5, -18.7),(5.5, -18.7),(7.7, -18.7),(9.9, -18.7),(12.1, -18.7),(14.3, -18.7),(16.5, -18.7),(-16.5, -20.9),(-14.3, -20.9),(-12.1, -20.9),(-9.9, -20.9),(-7.7, -20.9),(-5.5, -20.9),(5.5, -20.9),(7.7, -20.9),(9.9, -20.9),(12.1, -20.9),(14.3, -20.9),(16.5, -20.9)]
# --- Patterning ---
VOLUME = 1 # µL per dot
# Red layer (mrfp1)
pipette_20ul.pick_up_tip()
for (x, y) in mrfp1_points:
pipette_20ul.aspirate(VOLUME, location_of_color('Red'))
target = center_location.move(types.Point(x=x, y=y, z=0))
dispense_and_detach(pipette_20ul, VOLUME, target)
pipette_20ul.drop_tip()
# Green layer (sfgfp)
pipette_20ul.pick_up_tip()
for (x, y) in sfgfp_points:
pipette_20ul.aspirate(VOLUME, location_of_color('Green'))
target = center_location.move(types.Point(x=x, y=y, z=0))
dispense_and_detach(pipette_20ul, VOLUME, target)
pipette_20ul.drop_tip()
###
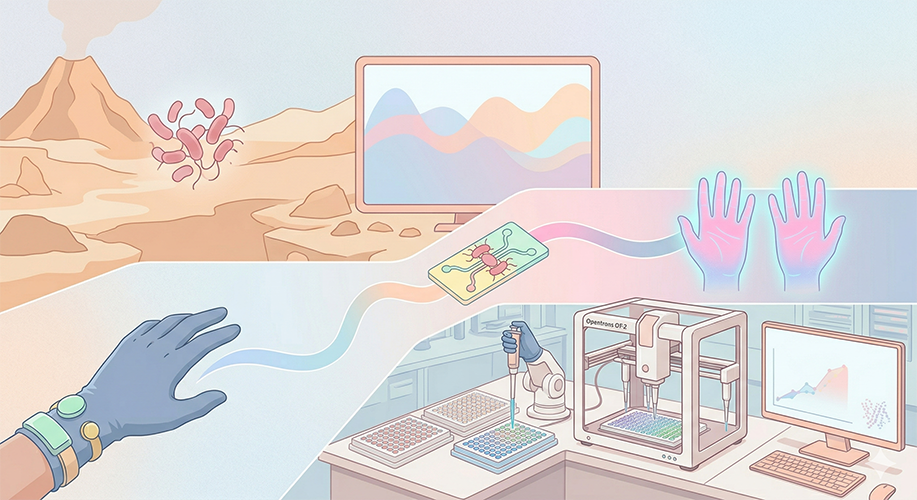
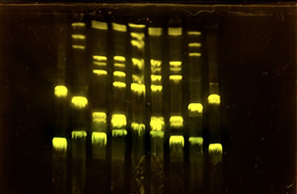
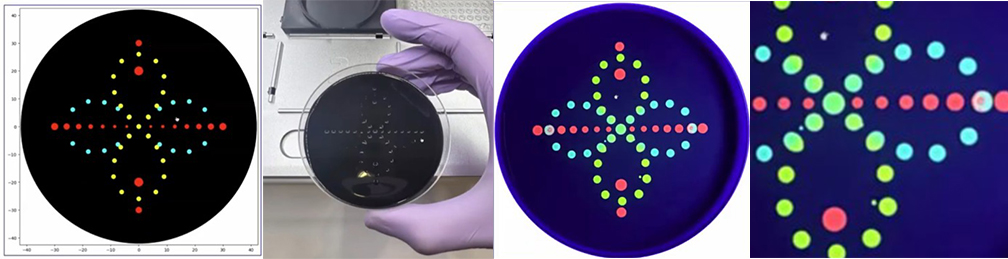
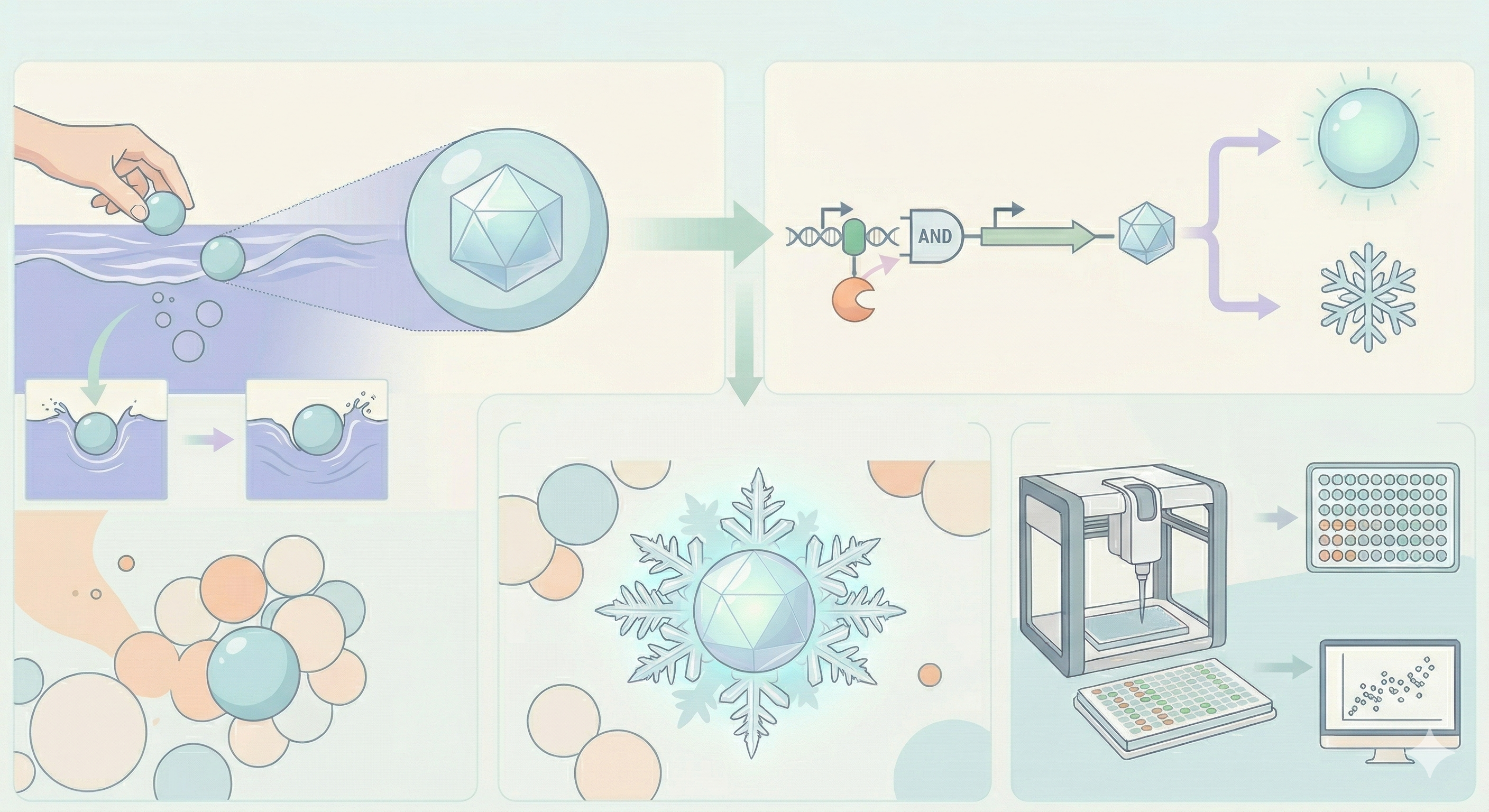
One of the great parts about having an automated robot is being able to precisely mix, deposit, and run reactions without much intervention, and design and deploy experiments remotely.
This paper is particularly relevant because it addresses a critical but often overlooked problem in laboratory automation: the gap between intended and actual liquid deposition. As demonstrated in the images captured by Chen (2026), this discrepancy becomes strikingly clear when comparing the physical petri dish under normal lighting conditions with its UV-illuminated counterpart. Under standard light, the dish appears largely as expected, with the deposited pattern barely distinguishable to the naked eye (Chen, 20256, Figure 2). However, when the same plate is examined under UV light, small, unwanted droplets become clearly visible in places where they were not present in the original design (Chen, 2026, Figures 3-4). The computer vision algorithm developed in this study successfully detects and maps these deviations, marking the spots with color codes according to their size to highlight the extent of the error.
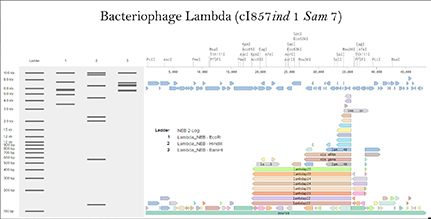
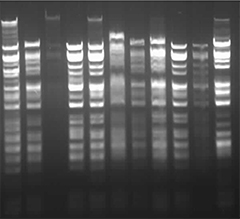
Figure 1-4: Images captured from the Opentrons OT-2 liquid handling experiments by Yanchen Chen. Credit: Yanchen Chen (24.02.2026).
These satellite droplets arise from well-known physical phenomena in liquid handling, such as surface tension-driven splashing or residual liquid remaining on pipette tips between transfers. What makes this finding biologically significant is that in high-precision applications such as drug screening, dose-response assays, or microbial growth experiments, even a small unintended deposit can introduce a compound or organism into a zone where it was never meant to be. This cross-contamination would silently corrupt experimental results, and without a real-time quality control system, the researcher would have no way of knowing the data was compromised.
The novel contribution of this work is therefore not purely engineering: by enabling the Opentrons OT-2 to detect and flag these errors autonomously using computer vision, the system directly protects the integrity of biological experiments. This transforms the robot from a simple liquid-dispensing tool into a self-monitoring platform capable of ensuring experimental validity; a meaningful advancement for any biological application that depends on precise, contamination-free liquid handling.
Echo transfer biosensor constructs and any required cofactors into specified wells.
Bravo stamp in CPFS reagent master mix into all wells of a 96-well / 384-well plate.
Multiflo dispense the CFPS lysate to all wells to start protein expression.
PlateLoc seal the plate.
Inheco incubate the plate at 37°C while the biosensor proteins are synthesized.
XPeel remove the seal.
PHERAstar measure fluorescence to compare biosensor responses.
Embedded slide deck of 1-3 slides with 3 ideas you have for an Individual Final Project. by naming (Beyza Batır, Izmir, Turkey)
I will upload my slides on CL powerpoint in DC Labs Student#7 section.
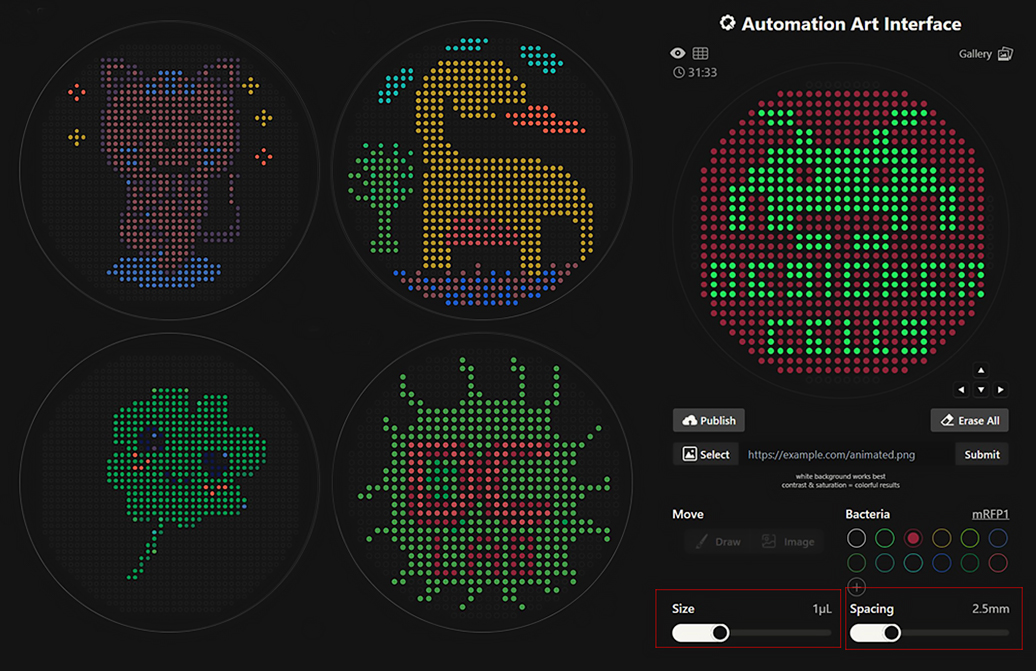
These extra designs were created within the “Automation Art Interface” to explore possibilities with different weights, colors, and area usage. I also created Designer Cells artworks for our node and this time corrected my mistakes (size, spacing, safe canvas margin) that did not comply with the requirements announced on HTGAA Google Colab. If needed, I can prepare Google Colab for all designs.