Brie Noel Taylor — HTGAA Spring 2026


About me
Designer and artist based in Brooklyn, very excited to do some weird science!


Designer and artist based in Brooklyn, very excited to do some weird science!
Week 1 HW: Principles and Practices
Week One Homework Brie Dreams of Science Q1 First, describe a biological engineering application or tool you want to develop and why.
Week 1 HW: Principles and Practices
Benchling & In-Silico Gel Art I completed this bio lab last week at GenSpace! It was an exciting and easy-to-follow lecture. I’ll be attending Lab in-person this week, and will gather images of how the gel art turned out. DNA Design Challenge For my protein, I am choosing superfolder GFP (sfGFP) expressed in E. coli. This is primarily because I’m new to a science-lab environment, and my online research suggested this would be a simple, commonly-used protein that would help me learn and understand the tools and systems without the homework getting overly complicated (i.e. I don’t want to get stuck without a TA at home!).
First, describe a biological engineering application or tool you want to develop and why.
For the purposes of the How To Grow (Almost) Anything course, I am interested in exploring wetware. Meaning, I’m interested in exploring how plants and biomatter can be included in - or replace - traditional hardware elements. The scale and scope of this exploration will be ascertained through the class itself. For now, I am imagining someone small scale, such as replacing a conductive element in a circuit board with a plant or biomatter.
The intent is explorative and speculative. Could trees be electricity conductors? Unlikely, but I’d like to start with explorative speculation and work backwards from that, pushing the limits of my own knowledge and scientific understanding as far as I can.
The why here is that I intend to incorporate my learnings, eventually, into my art and design practice. I am an installation artist and designer, and I’d like to explore new ways the boundaries between “natural” and “technological” systems can break down.
Goal One: Non-pathogenic materials choices
Goal Two: Closed-growth system with no environmental release
Goal Three: Building towards an open system
Prompt citation
Brie. (2026, February 10). User prompt to ChatGPT: “I am taking a science class through MIT and Harvard called ‘How to Grow Almost Anything’…” [Large language model prompt]. OpenAI ChatGPT.
Governance action one: Safety tiering system for materials used
Purpose: A large part of my proposed project is using plant and biomaterials in place of traditional hardware elements in computing technology. This will likely evolve over time, but that is where I’m at currently. The purpose of this governance action is to create safety-focused guardrails and limitations around the biomaterials I will experiment with during this project. A tiering system - either relying on existing tiering systems, which I will research, or creating a new one - will do this. For example, I will not use toxic materials.
Design: A safety tiering system for ‘wetware’ materials that categorizes potential wetware materials into categories based on safety criteria such as being self-generating, difficult to safely contain, respiratory toxicity, toxicity to skin, difficulty of safe end of project disposal, and so on.
Assumptions: An assumption I am making is a that a tiering system - or open information needed to create a tiering system for the purposes of this project - exists and is readily available. A second assumption is that the project timeline allows space and capacity for creating a tiering system while also working on the project itself.
Risks of failure: Miscategorizing materials or misunderstanding safety indicators. If used at scale, could be a “box check” safety measure without active impact. Could slow down experimentation unecessarily if too strict.
Governance action two: End-of-project safety and sterilization plan
Purpose: A large part of this project centers on experimenting with various bio and plant materials, for wetware use. Even if materials are contained safely within the lab during project and experimentation, at the end of the project, I will need to be able to safely dispose of materials in a way that does not distribute them unsafely into the broader environment. Potentially, this end of life project plan for materials could become a mandatory governance structure for comparable wetware projects.
Design: A mandatory checklist of end-of-project disposal, documentation and sterilization processes for wetware experimentation and build projects. A checklsit and documentation structure.
Assumptions: Assumes that materials will need special disposal processes. Assumes there is a safe way to dispose of all wetware experimentation materials. Assumes normal sterilization and disposal processes for the lab space would not be sufficient.
Risk of failutre: Cost and complexity could prevent small-scale or artistic experimentation. Complexity and documentation adds time. May need to differentiate between ongoing disposal (for example, trying a material, it doesn’t work, and moving on) and end of project disposal (full shut-down) - if doing both, could add too much complexity and paperwork.
Governance action three: Open systems
Purpose: Keepoing learnings open and available. Avoiding information hoarding. Avoiding use for non-ethical means.
Design: An open-source platform where people can share and gather learnings about wetware experimentation and learning. A documentation plan and guidelines that promotes consistent documentation patterns, language and styles, so knowledge can be easily shared and learnings replicated.
Assumptions: That open source sharing would be collectively used and desired. That open source sharing would help avoid non-ethical use or private monetization of ideas.
Risk of failure: No one uses it! Documentation guidelines are insufficient, or learnings over time create new documentation style needs that don’t match the original documentation structures.
| Does the option: | Govn 1 | Govn 2 | Govn 3 |
|---|---|---|---|
| Enhance Biosafety | |||
| • By preventing incidents | 1 | 1 | 3 |
| • By helping respond | 3 | 2 | n/a |
| Foster Lab Safety | |||
| • By preventing incident | 1 | 1 | 3 |
| • By helping respond | 3 | 3 | n/a |
| Protect the environment | |||
| • By preventing incidents | 1 | 1 | 2 |
| • By helping respond | 1 | 3 | n/a |
| Other considerations | |||
| • Minimizing costs and burdens to stakeholders | 2 | 2 | 1 |
| • Feasibility? | 3 | 1 | 1 |
| • Not impede research | 2 | 3 | 2 |
| • Promote constructive applications | n/a | n/a | 1 |
Based on my thinking, writing and numbered options, I would prioritize governance idea 3 - working towards open source learnings with accessible, consistent documentation. This is because lab safety and materials safety measures exist that could be employed to cover governance concwerns one and two, i.e. ensuring only ‘safe’ materials are used, and ensuring safe and documented disposal practices at end-of-project.
Governance action three focuses, instead, on making sure _learnings are available to everyone*. As environmental conservation is an implicit intent of this project (as it focuses on using biomaterials for hardware), any positive learnings should be made reproducable and repeatable for scientists wanting to build on the learnings, and should be shared openly so that learnings can’t become privately monetized.
Relavent audiences - Scientists. Hardware companies. Designers. Engineers. Architects.
Reflecting on what you learned and did in class this week, outline any ethical concerns that arose, especially any that were new to you. Then propose any governance actions you think might be appropriate to address those issues. This should be included on your class page for this week.
Honestly - no ethical concerns have come up thus far that I haven’t addressed in my answers above.
Homework Questions from Professor Jacobson:
Q: Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy?
A: Based on the lecture slides, the error rate of polymerase is 1x106. I’m going to be honest and say that I can’t tell just from reading the slides how it compares to the length of the human genome - I don’t have a science background, so need to attend the week’s lecture to understand how to parse that information from the slides. I really want to learn, so I’m going to be honest and say I don’t know the answer here.
Homework Questions from Dr. LeProust:
Q: What’s the most commonly used method for oligo synthesis currently?
A: A mix of high-throughput oligo synthesizer by Illuma and electrochemical-based micro-array.
Q: Why is it difficult to make oligos longer than 200nt via direct synthesis?
A: Problems with uniformity. Expensive.
Q: Why can’t you make a 2000bp gene via direct oligo synthesis?
A: You need a cloned oligo pool.
Homework Question from George Church:
Q: What are the 10 essential amino acids in all animals and how does this affect your view of the “Lysine Contingency”?
A: The ten essential amino acids in animals are histidine, isoleucine, leucine, lysine, methionine, phenylalanine, threonine, tryptophan, valine, and (in many animals and developmental contexts) arginine. These are called “essential” because animals cannot synthesize them in sufficient quantities and must obtain them from their diet.
This list reframes the idea of the “Lysine Contingency.” Lysine is not uniquely special in being essential—rather, it is one of a broader set of amino acids that animals fundamentally depend on external sources to obtain. What makes lysine interesting is not that life hinges on it alone, but that multiple core building blocks of proteins are metabolically unavailable to animals and must come from ecological or industrial supply chains (plants, microbes, or synthetic production). From this perspective, the “Lysine Contingency” is better understood as part of a more general nutritional and biochemical contingency: animal life is structurally dependent on a network of externally produced amino acids, with lysine being a prominent but not singular example.
Prompt citation: Brie. (2026, February 10). Question to ChatGPT about essential amino acids and the “Lysine Contingency” based on course slides [Large language model query]. OpenAI ChatGPT.
I completed this bio lab last week at GenSpace! It was an exciting and easy-to-follow lecture. I’ll be attending Lab in-person this week, and will gather images of how the gel art turned out.
For my protein, I am choosing superfolder GFP (sfGFP) expressed in E. coli. This is primarily because I’m new to a science-lab environment, and my online research suggested this would be a simple, commonly-used protein that would help me learn and understand the tools and systems without the homework getting overly complicated (i.e. I don’t want to get stuck without a TA at home!).
Additionally, it sounds like superfolder GFP (sfGFP) expressed in E. coli can be used as a bio-indicator (emitting light), which may end up being helpful for my final project.
I have used Uniprot to obtain the following protein sequence:
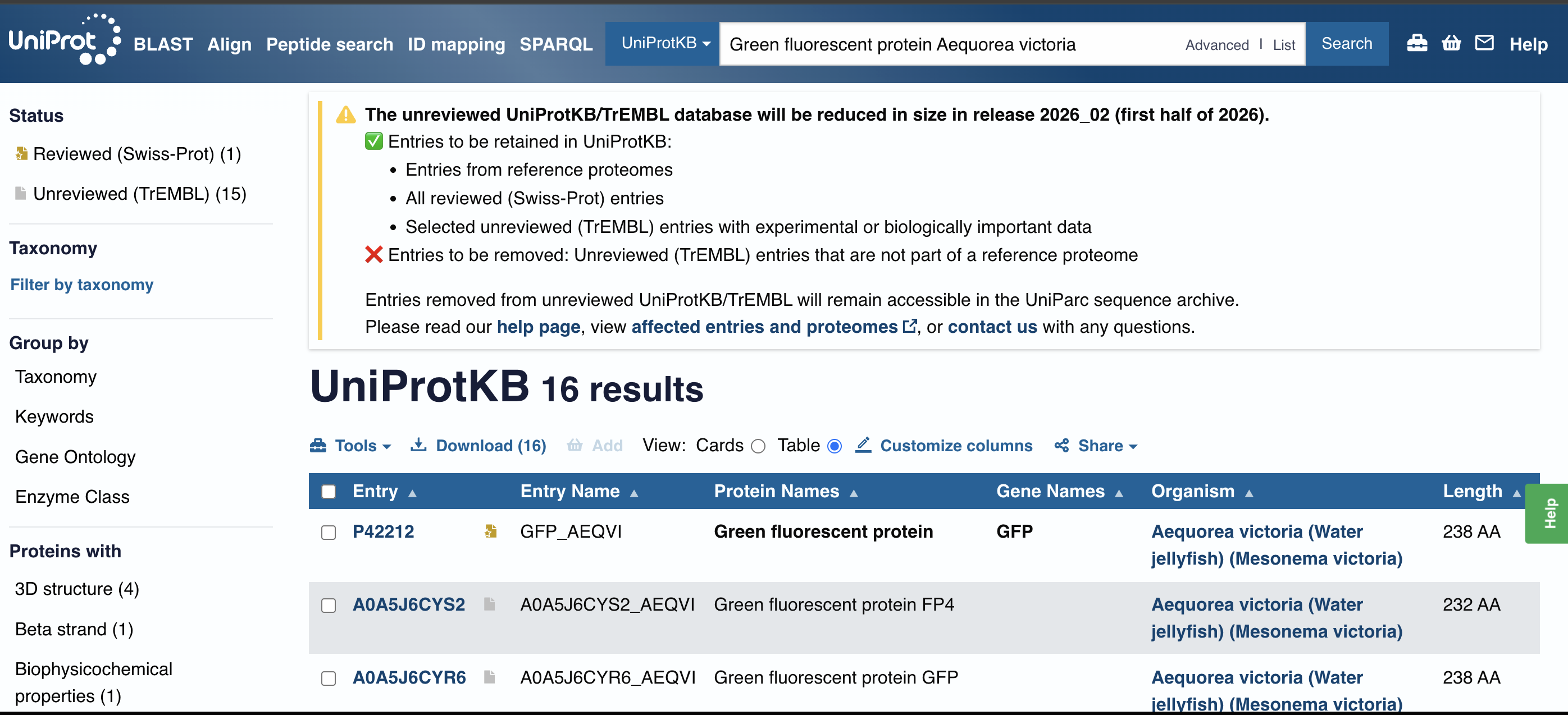

I asked ChatGPT to help me understand exactly which name to search in Uniprot to find my protein:
Taylor, B. N. (2026). Consultation with ChatGPT (OpenAI, GPT-5) to determine appropriate UniProt search term for selected protein (“Green fluorescent protein Aequorea victoria”). Personal communication, February 16, 2026.
My sequence (yay!):
sp|P42212|GFP_AEQVI Green fluorescent protein OS=Aequorea victoria OX=6100 GN=GFP PE=1 SV=1 MSKGEELFTGVVPILVELDGDVNGHKFSVSGEGEGDATYGKLTLKFICTTGKLPVPWPTL VTTFSYGVQCFSRYPDHMKQHDFFKSAMPEGYVQERTIFFKDDGNYKTRAEVKFEGDTLV NRIELKGIDFKEDGNILGHKLEYNYNSHNVYIMADKQKNGIKVNFKIRHNIEDGSVQLAD HYQQNTPIGDGPVLLPDNHYLSTQSALSKDPNEKRDHMVLLEFVTAAGITHGMDELYK
Next, I found an online reserve translation tool to reverse my protein sequence. I chose Bioinformatics. I copied and pasted my protein sequence, and hit ‘submit’ to get the following codons:
atgagcaaaggcgaagaactgtttaccggcgtggtgccgattctggtggaactggatggc gatgtgaacggccataaatttagcgtgagcggcgaaggcgaaggcgatgcgacctatggc aaactgaccctgaaatttatttgcaccaccggcaaactgccggtgccgtggccgaccctg gtgaccacctttagctatggcgtgcagtgctttagccgctatccggatcatatgaaacag catgatttttttaaaagcgcgatgccggaaggctatgtgcaggaacgcaccatttttttt aaagatgatggcaactataaaacccgcgcggaagtgaaatttgaaggcgataccctggtg aaccgcattgaactgaaaggcattgattttaaagaagatggcaacattctgggccataaa ctggaatataactataacagccataacgtgtatattatggcggataaacagaaaaacggc attaaagtgaactttaaaattcgccataacattgaagatggcagcgtgcagctggcggat cattatcagcagaacaccccgattggcgatggcccggtgctgctgccggataaccattat ctgagcacccagagcgcgctgagcaaagatccgaacgaaaaacgcgatcatatggtgctg ctggaatttgtgaccgcggcgggcattacccatggcatggatgaactgtataaa
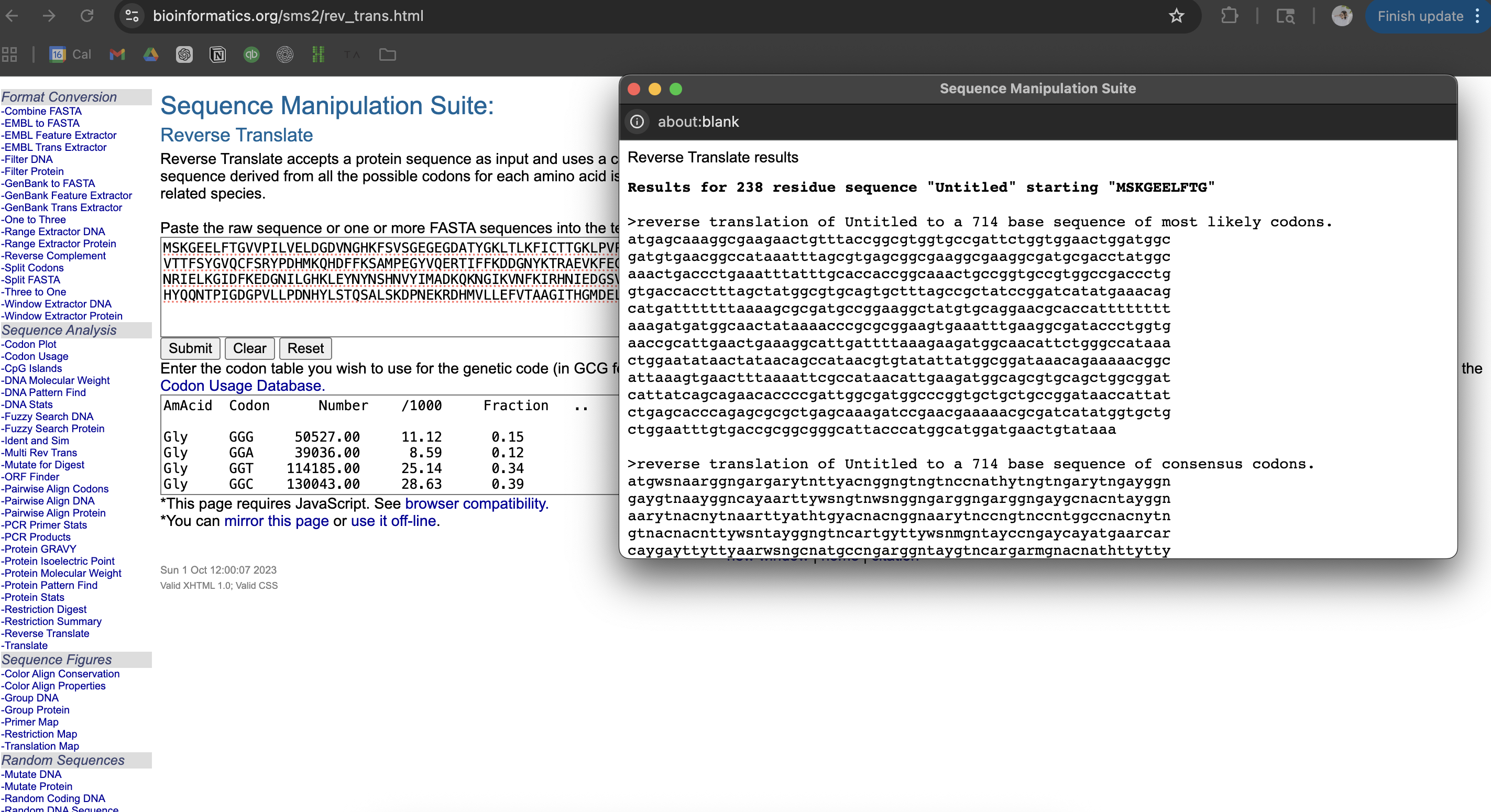

Next, I did some online reading about codon optimization, as I wasn’t sure about the ‘why’ and didn’t recall the why from the relevant lecture. This included Google and ChatGPT research:
Taylor, B. N. (2026). Consultation with ChatGPT (OpenAI, GPT-5) regarding codon optimization rationale and host organism selection (E. coli) for GFP expression. Personal communication, February 16, 2026.
I will be using e.coli as, again, my research suggests it is simple and commonly-used, and I am optimizing for minimizing complexity while I am learning new skills.
I tried using the Twist Codon Optimization Tool, as that’s what’s referenced in the homework examples, but hit a roadblock figuring out how to input my codon sequence into the tool. I therefore pivoted to IDTDNA. I input my reverse codon ouput (and removed line breaks!), and then hit optimize:
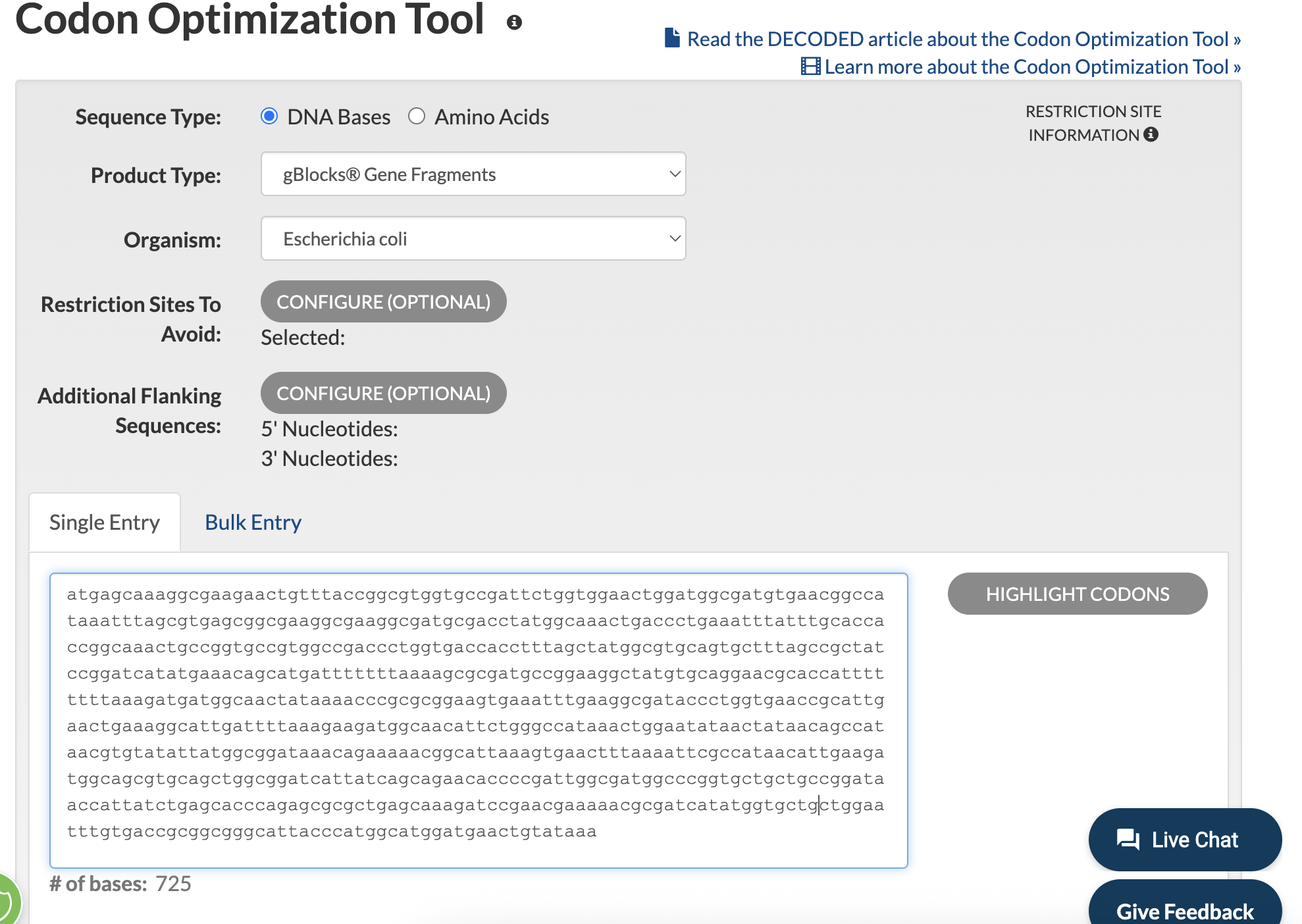

I then obtained the follow results:
ATG AGC AAG GGC GAA GAA TTG TTC ACT GGC GTT GTT CCT ATT CTG GTG GAA CTG GAT GGT GAT GTA AAC GGT CAT AAA TTT AGT GTC TCC GGA GAG GGG GAA GGC GAC GCG ACT TAC GGT AAG CTG ACC CTG AAA TTC ATT TGC ACA ACT GGC AAA TTG CCG GTT CCT TGG CCA ACC CTG GTA ACA ACA TTT TCA TAT GGT GTT CAA TGC TTT AGC CGT TAT CCT GAC CAT ATG AAA CAA CAT GAT TTT TTC AAA AGT GCG ATG CCC GAA GGT TAC GTC CAA GAA AGA ACG ATT TTC TTC AAA GAT GAC GGT AAT TAT AAA ACA CGG GCC GAA GTT AAG TTT GAA GGA GAT ACA TTA GTA AAC CGT ATT GAG CTG AAA GGC ATT GAT TTC AAG GAA GAT GGT AAT ATT CTG GGT CAC AAG TTA GAA TAT AAC TAC AAC AGC CAT AAT GTT TAT ATT ATG GCT GAT AAA CAG AAA AAC GGC ATT AAG GTA AAC TTT AAG ATC CGG CAT AAT ATT GAG GAT GGA TCA GTG CAG TTA GCT GAT CAT TAT CAA CAG AAT ACA CCT ATA GGT GAC GGG CCG GTC CTG TTG CCT GAT AAC CAT TAT CTG AGT ACC CAA TCC GCG TTG TCA AAA GAC CCG AAC GAA AAA CGT GAC CAT ATG GTT TTG TTA GAA TTT GTT ACC GCC GCT GGG ATA ACT CAT GGA ATG GAC GAA CTG TAT AAA
Plus, restriction enzyme sites found:
BclI (TGATCA) BspEI (TCCGGA) MscI (TGGCCA) NdeI (CATATG) PsiI (TTATAA) SspI (AATATT)
Once I have the optimized GFP DNA sequence, the protein can be made either inside living cells or using a cell-free system.
In a cell-dependent method, the GFP DNA would be inserted into a plasmid with a promoter and ribosome binding site. The plasmid would then be put into E. coli bacteria. Inside the bacteria, the cell’s machinery reads the DNA. First, RNA polymerase transcribes the DNA into messenger RNA (mRNA). Then, the ribosome reads the mRNA and translates it into the GFP protein using the genetic code. As the protein folds, it becomes fluorescent.
In a cell-free system, the same DNA can be added to a mixture that contains the necessary transcription and translation machinery (enzymes, ribosomes, nucleotides, amino acids). The system produces mRNA and then translates it into protein outside of a living cell. This method is useful when working conditions need to be tightly controlled.
Taylor, B. N. (2026). Consultation with ChatGPT (OpenAI, GPT-5) to clarify transcription and translation steps for GFP expression from optimized DNA. Personal communication, February 17, 2026.
I opened Benchling, and entered my DNA string:
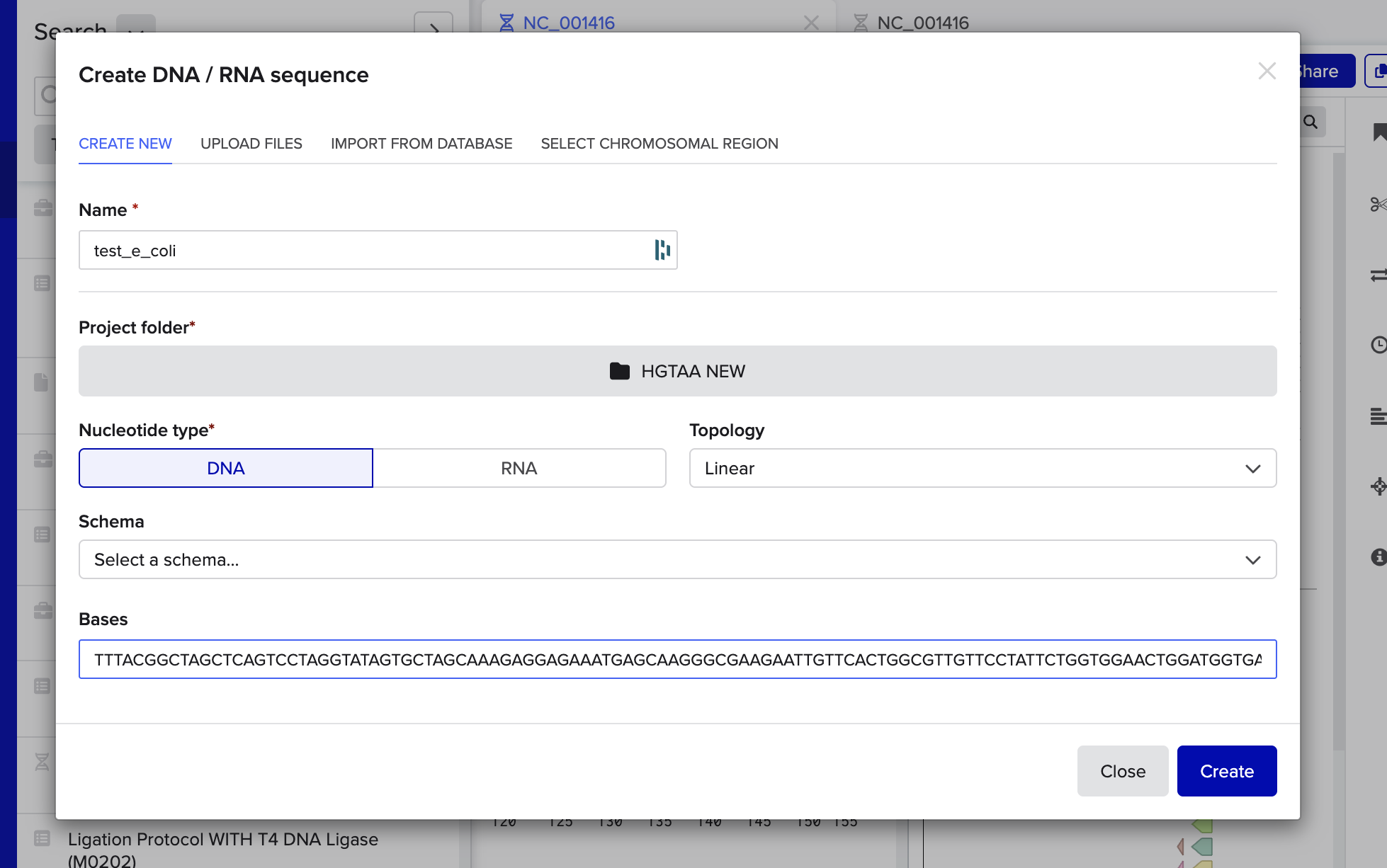

Then I began annotating:
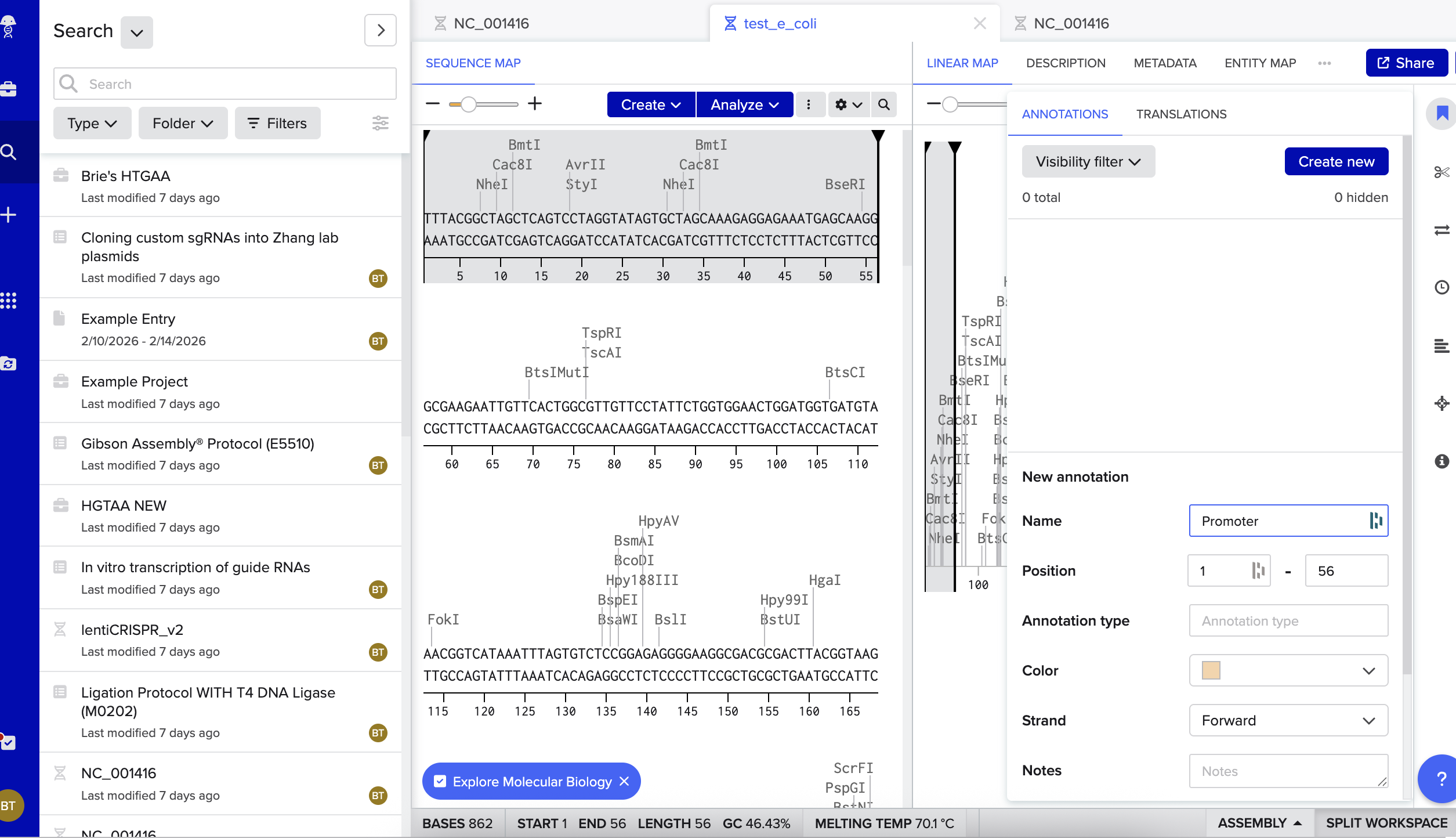

Next up, I added annotations. Oh boy. This was really hard. I may have made mistakes here. Though I tried very very hard! I mostly tried to mimic the sequence and annotations listed in the homework file, though did also ask ChatGPT for help talking me through the steps and understanding what is going on:
Taylor, B. N. (2026). Consultation with ChatGPT (OpenAI, GPT-5) for guidance on assembling and annotating a GFP expression cassette in Benchling (promoter, RBS, CDS, His tag, stop codon, terminator). Personal communication, February 17, 2026.
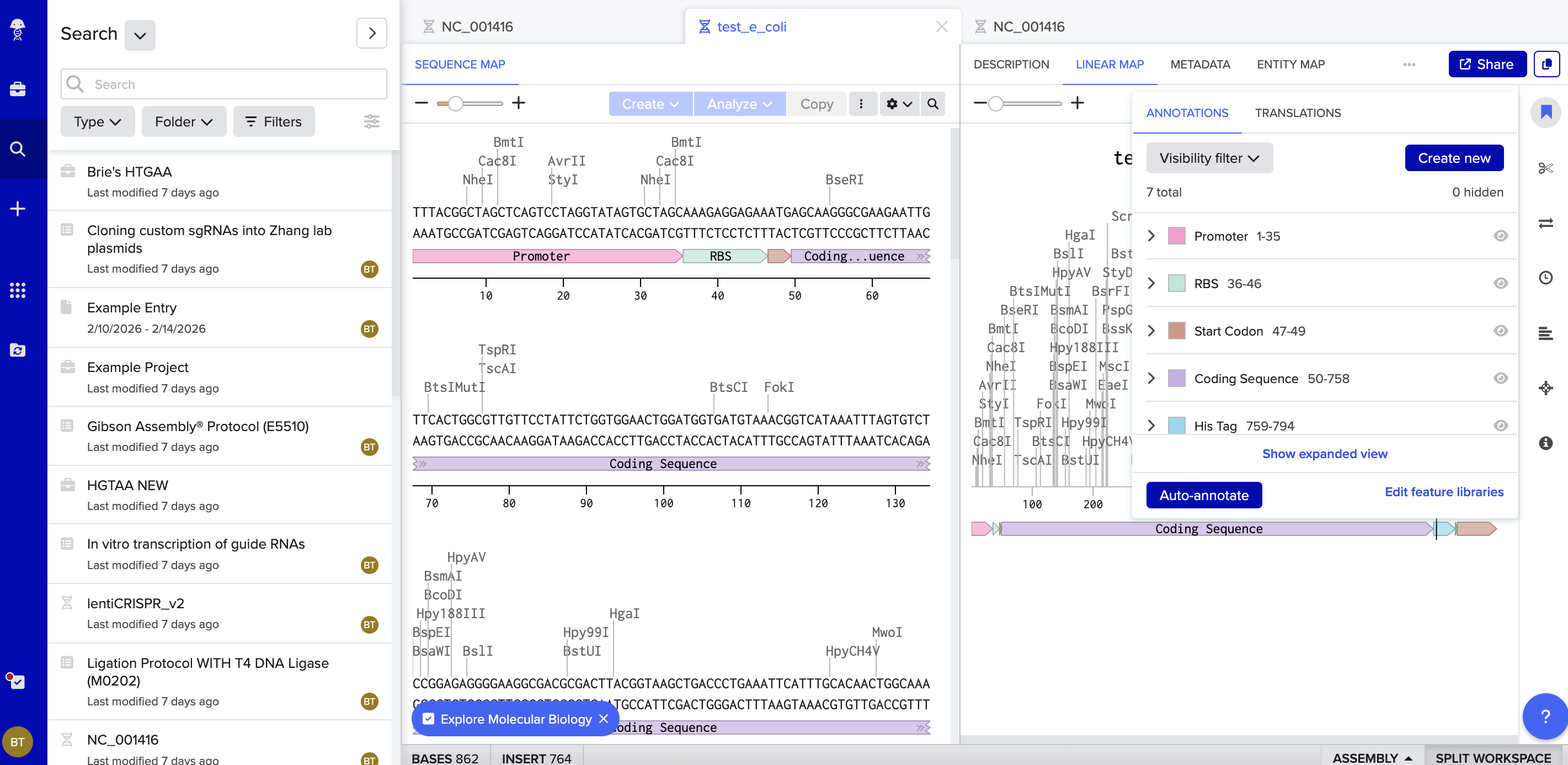

Here is the link to my linear map: https://benchling.com/s/seq-DQElyxqQGlBDYM96Iy9P?m=slm-LevVABkxM5ZzWgkQx9rv
I then followed the homework instructions to download my sequence, and upload it into Benchling. Here’s the receipts!
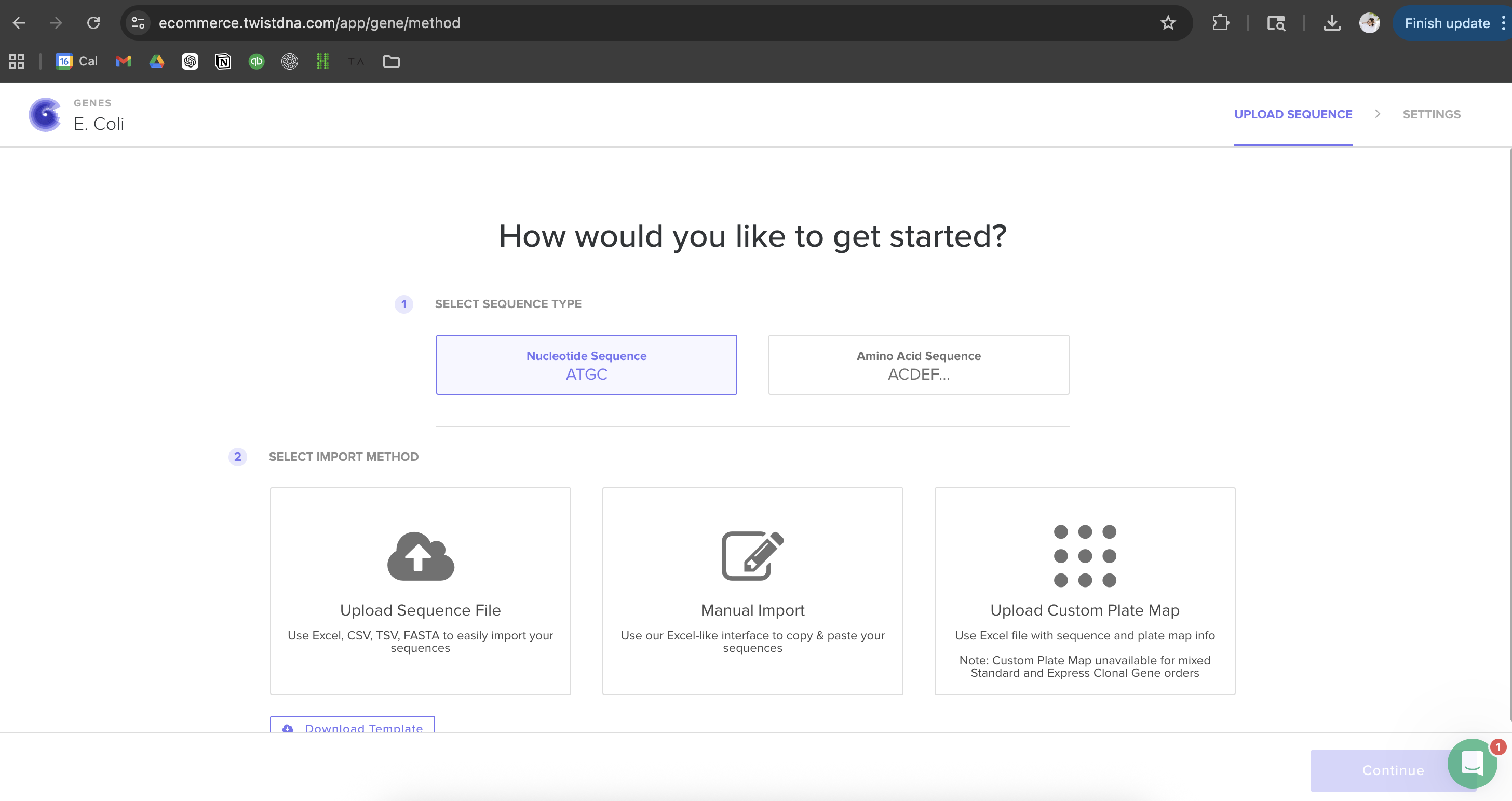

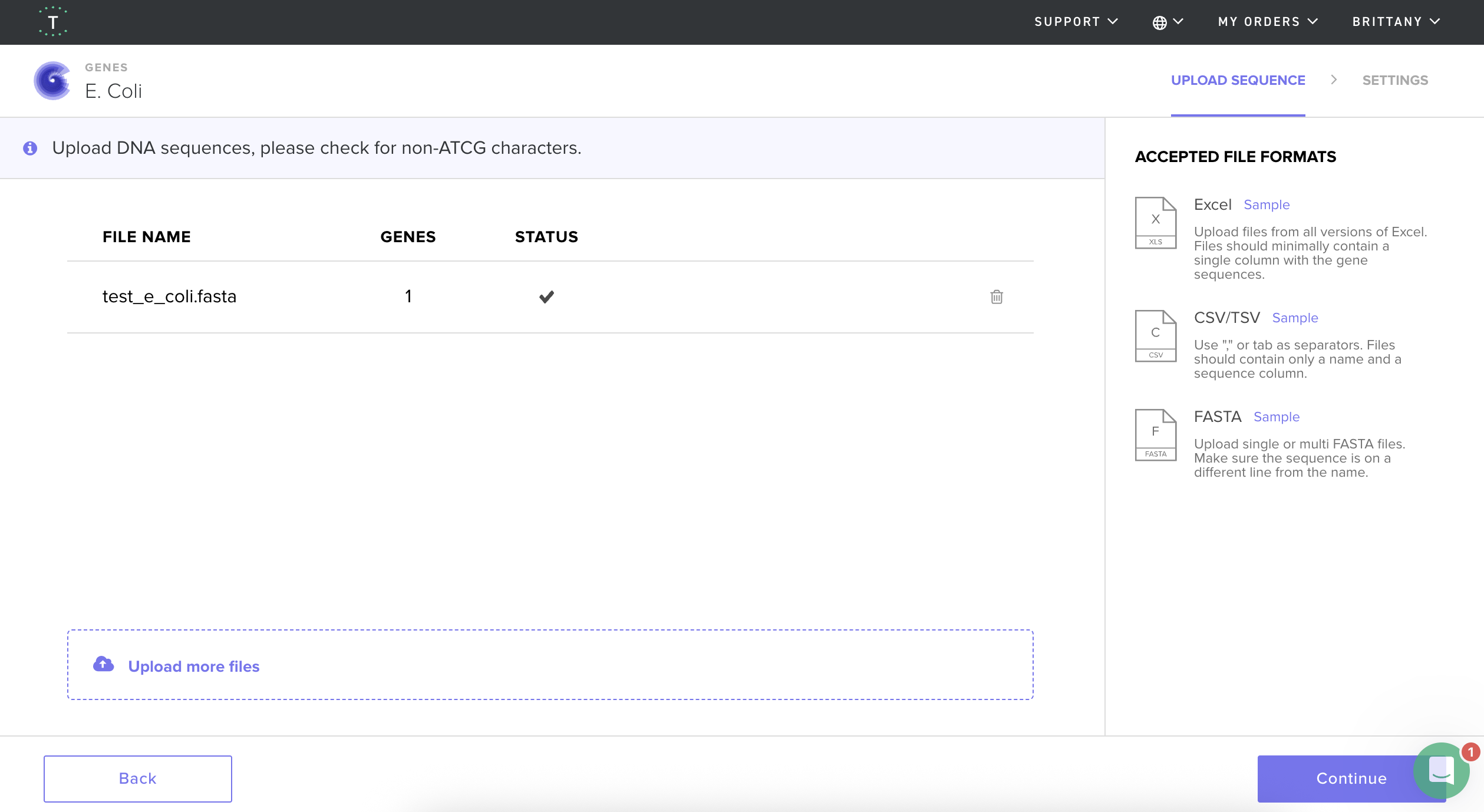

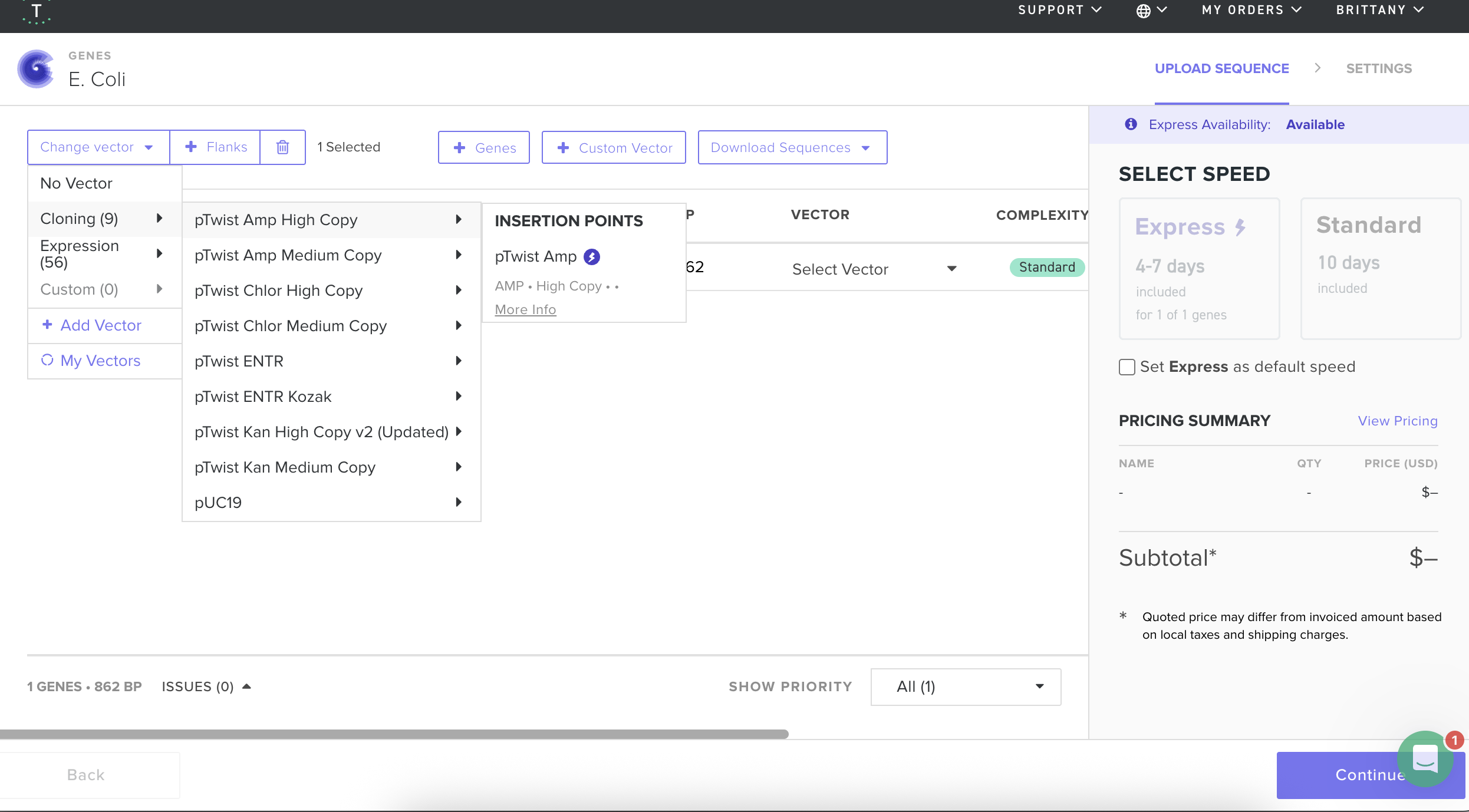

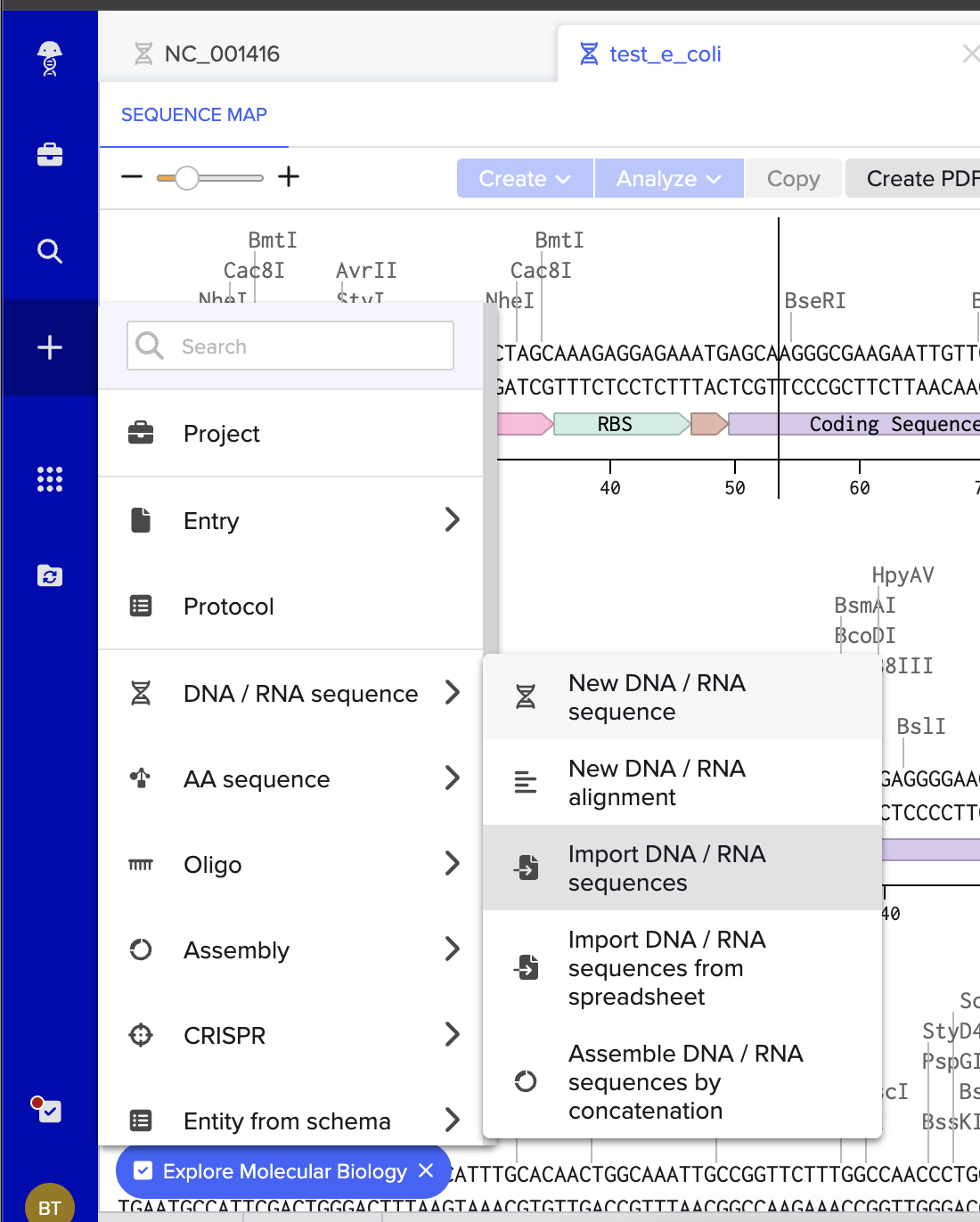

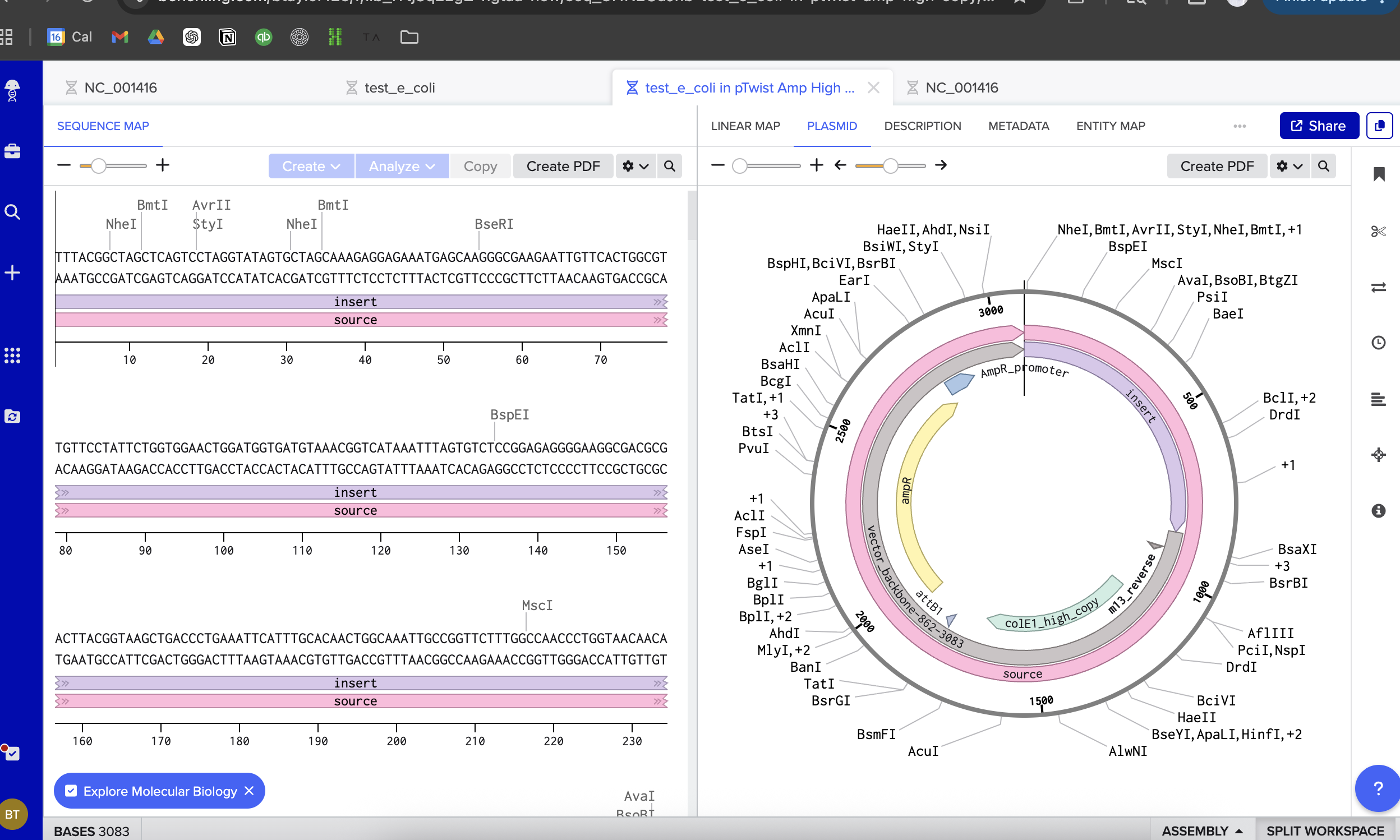

I would want to sequence DNA from environmental water samples, such as river or wastewater samples. This could help identify which bacteria or microorganisms are present. I’m interested in environmental systems, and sequencing environmental DNA (eDNA) could help detect pollution or biodiversity changes.
What technology would I use?
I would use Illumina sequencing. It is widely used, high accuracy, and good for sequencing many short DNA fragments at once.
This is a second-generation sequencing method.
Input: DNA extracted from a sample.
Preparation: DNA is fragmented, adapters are added, and PCR is used to amplify fragments.
During sequencing, fluorescent signals are detected as nucleotides are added one by one.
Output: Short DNA reads in digital format that can be analyzed on a computer.
Limitations: Reads are short, so assembly can be complex. It also requires lab preparation and equipment.
Taylor, B. N. (2026). Consultation with ChatGPT (OpenAI, GPT-5) to clarify sequencing, synthesis, and genome editing concepts for introductory-level coursework. Personal communication, February 17, 2026.
I would synthesize a GFP expression cassette for E. coli, like the one I designed. This could allow bacteria to produce a visible signal (fluorescence), which could later be adapted for sensing systems.
What technology would I use?
I would use chemical DNA synthesis (phosphoramidite synthesis) combined with gene assembly.
Short DNA fragments are chemically synthesized.
These fragments are assembled into a full gene.
The gene can then be cloned into a plasmid.
Limitations: There are size limits and cost constraints. Longer sequences require assembly steps.
Taylor, B. N. (2026). Consultation with ChatGPT (OpenAI, GPT-5) to clarify sequencing, synthesis, and genome editing concepts for introductory-level coursework. Personal communication, February 17, 2026.
I would edit bacterial DNA to insert a reporter gene like GFP under control of a promoter that responds to environmental signals (for example, stress or pollution). This could allow bacteria to act as biosensors.
What technology would I use?
I would use CRISPR-Cas9.
A guide RNA is designed to match a target DNA sequence.
Cas9 cuts the DNA at that location.
A repair template can be added to insert new DNA.
Limitations: Off-target effects can happen. Efficiency is not always 100%.
Taylor, B. N. (2026). Consultation with ChatGPT (OpenAI, GPT-5) to clarify sequencing, synthesis, and genome editing concepts for introductory-level coursework. Personal communication, February 17, 2026.