Cheap and Open-Source DIY DNA Synthesis


BLP-2000D by Georg Tremmel and Shiho Fukuhara, who produced a DNA printer as a bioart work which asks the question about governance, biopolitics and biosecurity in relation to DIY DNA synthesis.
Having had the privilege of growing up down the road from SymbioticA, research centre for biological arts, I found myself at an impass when the research centre was (unfairly) shutdown by the university at the start of 2023. As a bioartist working with DNA origami, The difficulty I faced after the closure of the research centre wasn’t procuring laboratory equipment; after all, lab equipment is often relatively simple to DIY, and components wise are generally cheap without the inflation tacked on by “Big Science Suppliers”. After all, Paul Rothemond developed the first DNA origami structures at his own home using his kitchen stove as the thermocycler.
No, the difficulty I found was in the procurement of, and pricing of, laboratory reagents from companies like sigma or even twist biosciences, whom would not allow me to order DNA to a residential address despite having the qualifications and the sequence itself being non-biohazardous and thus not requiring even a PC -1 Lab for its use.
If a global biotech future is to be truly accessible, something needs to be done about the cost and portability of dna synthesis. It is all well and good for scientists and researchers integrated into universities or private laboratories to be able to test and develop DNA nanotechnology using commercially available ssDNA synthesis, but for biohack spaces and the informed civilian/bioartist at home, the cost ssDNA oligos scales to the point at which it becomes financially prohibitive to use techniques like rapid prototyping or even just having to do multiple experiments because the authors of the protocol you are following missed out an important step and all of a sudden your experiment failed, oligos gone, and having to reach for the wallet.
DNA synthesis is conceptually simple, the reagents are cheap, and until the rise of 3D printers there was no way for a hobbyist to produce a DNA synthesis device. That being said, several “DIY” DNA synthesis devices have been produced by scientists, however these exist outside of a budget which is feasible for community biohack spaces ($15000 USD and up)
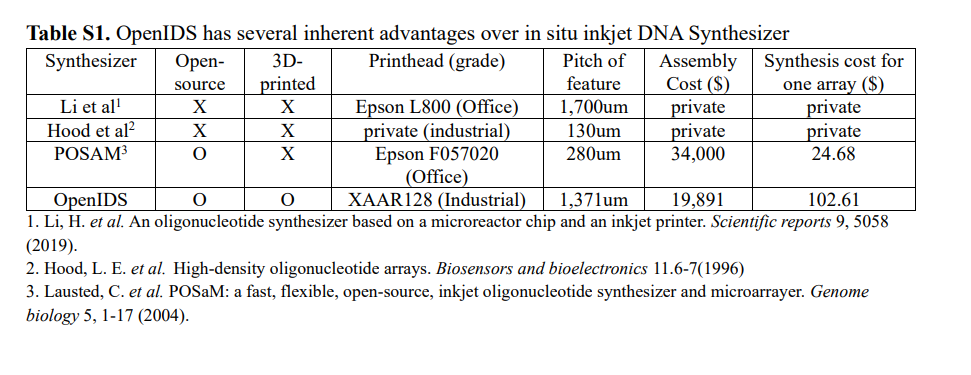

Kim, J., Kim, H. & Bang, D. An open-source, 3D printed inkjet DNA synthesizer. Sci Rep 14, 3773 (2024). https://doi.org/10.1038/s41598-024-53944-x
As such, my aim for this course is to produce a truly DIY and truly low-cost DNA synthesis pathway and/or device which allows for hobbyist biohack spaces to generate ssDNA sequences with an error rate of 5-10%.
The governance question for DIY DNA synthesis is a tricky one, and I draw some inspiration from Georg Tremmels work BLP-2000D in answering this, how do we facilitate the synthesis of DNA to address the inequality of access to biotech while still taking biosecurity seriously and ensuring that our DIY protocol does not enable illicit use cases? To be more clear, I break this down into three overarching governance goals
- How do we prevent harm and misuse from DIY DNA synthesis?
- How to create a compliance system within our DIY protocol?
- How do we enable equal access to DNA synthesis?
- How can we reduce the cost of DNA synthesis so that non-grant-funded isntitutions can experiment with ssDNA oligos?
Governance actions:
- Capability-limited DNA synthesis
- design the tool so it is optimized for oligos of limited length
- prevents the production of protein/gene sequences as well as preventing the production of viral genomes for instance.
- design the tool so it is optimized for oligos of limited length
- Background checks and restricting access to the DIY DNA synthesis protocol without being accepted (still free)
- design software for interfacing with the DIY DNA synthesis device which screens the oligos being produced or asks the user to write a short description about the usecase of the oligo and then check whether their description and the oligo sequence match.
- Partner with maker spaces who are wanting to/already have biohack/community lab spaces and work with them to produce cheap DIY DNA synthesis devices which are embedded into their infrastructure.
| Aspect | Action 1: Capability-limited DIY | Action 2: Screening/Attestation layer | Action 3: Community biofoundry partnership |
|---|---|---|---|
| Purpose | Design the tool so it is optimized for short oligos only, discouraging scaling into gene/protein synthesis | screening happens at companies (sometimes) when you order sequences — independent synthesis could bypass those chokepoints entirely. As such, integrate a voluntary, screening-and-attestation layer; either at the software level or at the level of initial access to the DIY DNA synthesis protocol (background check) to prevent DIY DNA synthesis from falling into “the wrong hands” | Allow for DIY DNA synthesis while offsetting the regulation and discression of access to the protocol/device to community spaces rather than residential open access |
| Design | Actors:Open-source device designers implement hard limits on oligo length produced, through hard-coding or inherent error rate increasing when length of ssDNA production increases | Actors: Government/International bodies generate lists of banned DNA/protein sequences; device designers and lab managers ensure proper implementation of these bans. | Actors: Community biolabs partner with device designers to implement DIY DNA synthesis program/protocols. COmpanies can ship easier to community biolabs than home residencies |
| Assumptions | Limiting oligo length meaningfully reduces misuse risk (it reduces some capability, but not all). That constraints won’t just be modified out by advanced users. That “short-only” still meets enough legitimate use cases to be worth building. | That robust “risk checks” can be defined without outlining and inspiring misuse cases | That communities can sustain this operationally (staffing, governance fatigue, liability). That hub models remain accessible rather than becoming exclusive clubs. That suppliers will support delivering reagents for DIY DNA synthesis. |
| Risks | Failure: informed and inclined users would be able to circumvent this through convoluted steps to merge fragments of shorter ssDNA oligos into longer sequences. Additionally People bypass or fork the design to accomodate longer ssDNA synthesis.** Success:** If this system becomes widely adopted regulators and media may clamp down or point fingers when a single biosecurity risk incident occurs, even if unrelated. | Failure: Easy to lie or create false persona to pass the initial background checks.** Success:** Access to ssDNA synthesis is still being arbitrarily gatekept just the body which gatekeeps becomes an open-source organization rather than commercial/institutional bodies. Might normalize a world where creative/research autonomy requires constant permissioning, | Failure: Geographically uneven access to community biolabs, leading to our goal of open-access to all interested being inhibited by logistical or local government fund distributions. Success: If becomes too good/popular access in community labs may be bottlenecked by realistic physical limitations of access or having to pay for memberships to community labs |
| Policy Goal | Option 1: Capability-limited DIY | Option 2: Screening/Attestation layer | Option 3: Community biofoundry partnership |
|---|---|---|---|
| Enhance Biosecurity — prevent incidents | 2 | 1 | 2 |
| Enhance Biosecurity — help respond | 3 | 2 | 1 |
| Foster Lab Safety — prevent incidents | 2 | 2 | 1 |
| Foster Lab Safety — help respond | 3 | 2 | 1 |
| Protect environment — prevent incidents | 2 | 2 | 1 |
| Protect environment — help respond | 3 | 2 | 1 |
| Minimize costs & burdens | 1 | 2 | 3 |
| Feasibility | 2 | 2 | 2 |
| Not impede research | 1 | 2 | 2 |
| Promote constructive applications | 1 | 2 | 1 |
Describe which governance option, or combination of options, you would prioritize, and why. Outline any trade-offs you considered as well as assumptions and uncertainties
Audience for this recommendation I’m writing this recommendation primarily for community biolab / biohackspace leadership (e.g., makerspace boards, lab managers, and their insurers) and secondarily for national regulators and public funders (in an Australian context: agencies that set biosecurity policy and the grant bodies that fund community science infrastructure). These audiences are directly positioned to implement practical access pathways while maintaining duty-of-care, lab safety, and incident response capacity.
Governance recommendation: Based on our initial goal with this project for expanding access to DNA synthesis without contributing to global biosecurity concerns with biowarfare/misuse of DNA nanotechnology, I would prioritize combination of Option 1 + Option 3, with the implimentation of the software-level screening of DNA sequences to be synthesized from Option 2.
Option 3 is the most effective for upholding biosafety and biosecurity responsibilities while also enabling wider access to DNA synthesizers. Additionally, it concentrates hazardous steps (chemicals, waste, training) inside spaces that can maintain protocols, supervision, incident logs, and disposal systems.
Option 1 reduces worst-case misuse by constraining the device/protocol to short oligos, making it less directly suitable for gene/protein construction or full genomes, even if a motivated actor could theoretically assemble fragments later.
Option 2 should exist as risk friction, not as the main “permission” system: it can help align DIY synthesis with norms already present in commercial screening, without turning community science into constant surveillance or arbitrary denial.
Trade-offs considered Biosecurity vs autonomy: Screening of users/uses and controlled access reduces the risk for misuse, but can also recreate the exact exclusion mechanisms that currently block residential bioartists and community labs.
Centralization vs equitable access: By focusing on community biofoundry hubs the biosafety infrastructure is offset to the organization using our DIY DNA synthesizer, but risks becoming new gatekeepers or failing to solve the problem of socioeconomic and geographic inaccessibility.
Feasibility vs ideal governance: Hub models require staffing, training, and insurance. If this fails politically or financially, limiting the capability of the DIY DNA synthesizer still provides a partial harm-reduction pathway.
Assumptions
Limiting synthesis to short oligos meaningfully reduces immediate misuse capability relative to unconstrained synthesis.
Community labs can maintain baseline safety (training, chemical handling, waste disposal) better than unsupervised residential settings.
Suppliers are more likely to ship reagents to incorporated community labs than private residences.
The primary legitimate use cases for this project (education, prototyping, bioart research) are compatible with short oligo lengths and/or hub-based access.
Uncertainties Whether constraint mechanisms would be forked/removed by advanced users, undermining Option 1. Whether public perception/media narratives would treat any incident (even unrelated) as justification to restrict all DIY biology. Whether screening/attestation can be done without creating a harmful surveillance norm or excluding marginalized users. Whether community labs can sustainably staff, insure, and govern a “biofoundry-like” service without burnout or prohibitive cost.
Ethical concerns that arose this week Reflecting on class, the main ethical concerns for me weren’t abstract, several felt newly personal and operational given my experience being blocked from ordering benign sequences to a residential address:
Equity and epistemic injustice (who is allowed to do biotech): I was struck by how governance often defaults to “institutional legitimacy” (university affiliation, commercial lab address) rather than demonstrated competence and intent. This can entrench inequity while creating an illusion of safety.
Dual-use tension without clear boundary lines: Short oligos are not inherently safe or unsafe; risk emerges from context, assembly pathways, and intent. The ethical difficulty is preventing harm without treating all curiosity as suspect.
Privacy and surveillance: While screening sounds reasonable, history has shown that both humans and AI are capable of biased decisions. Additionally, by normalizing identity checks, intent declarations, and centralized logging of biological “interests,” which may therein become a simulacra of the institutional blockades we are aiming to circumvent with a low cost DIY solution.
Environmental and safety infrastructures: Even if the biological mechanisms are benign, reagents (solvents, acids, waste streams) carry real safety concerns and support for finding waste disposal centers/protocols should be included in the final product.
Governance actions to address those ethical issues A) Build “responsibility infrastructure,” not just restrictions
- Community-lab SOP bundles: standardized training modules, signage, spill response, storage, disposal logs, and incident response templates packaged with the protocol/device.
- Safety-by-default design: reagent volumes minimized; enclosed fluid paths; outline reagent waste protocols and guidance. B) Constraint capability in ways that are transparent and contestable Short-oligo length limits as a default (Option 1), plus explicit documentation of why this reduces certain misuse pathways. Make constraints visible and auditable (so communities can trust them), but avoid publishing “how to bypass” guides. C) Use screening as a low-friction, privacy-preserving layer Prefer local screening (software runs on-device) rather than central servers collecting sequences. Use attestation prompts focused on safety (e.g., “confirm you have chemical PPE and waste plan”) rather than intrusive background checks. If any logging exists, it should be opt-in, minimal, and controlled by the community lab, not an external entity.
D) Create equitable access lanes
1. Community biofoundry access model (Option 3): subsidize memberships or reagents, and host public workshops so hubs don’t become exclusive clubs.==
E) Clarify scope and boundaries honestly
1. Publish a clear statement of intended use cases (education, bioart prototyping) and explicitly outline what the protocol is NOT to be used for to provide legal protection (eg. pathogens, toxins, etc.).
2. Provide a “red flags” ethics guide that focuses on behavior and context rather than listing harmful sequences. Often informing users can be enough to prevent misuse.
Pre-Week 2 Homework
Homework Questions from Professor Jacobson:
Q: Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy?
A: error rate of polymerase is 1:10^6 (1 in every 1,000,000). Length of Human Genome is 3.2 Gbp (3,200,000,000), so when the error rate is applied to the human genome there would be roughly 3200 errors during cellular replication if not for embedded systems such as: the MutS repair system which detects DNA basepair mismatches and binds at the site, recruiting DNA repair machinery to correct the error; as well as inbuilt redundancy in some codons which allows for the existence non-harmful missense DNA mutations.
Q: How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice what are some of the reasons that all of these different codes don’t work to code for the protein of interest?
A:
pt 1. Average Human Protein is 1036 bp which is ~345 amino acids. then using the table of codon frequency in humans https://www.genscript.com/tools/codon-frequency-table we can calculate the average number of codons (i.e. redundancy) available to encode for an AA which is roughly 3.5 codons per amino acid. which means for the total length of the average human protein there is roughly 3.5^345 different ways to code for it.
pt 2. The nucleotide composition for codons may actually have important differences due to epigenetic alterations which the change in nucleotide composition opens or shuts for the specified gene. Additionally, the altered nucleotide composition would result in an altered mRNA composition which may have differing regulatory interactions than the normally encoding mRNA. Finally, the GC content of a gene can alter its tertiary structure, meaning that an alternative codon encoding the same protein but with a higher GC content could face problems with its transcription.
Homework Questions from Dr. LeProust:
Q: What’s the most commonly used method for oligo synthesis currently?
A: Phosphoramidite Chemical Synthesis
Q: Why is it difficult to make oligos longer than 200nt via direct synthesis?
A: There are error-correction layers embedded in genetic code and in DNA synthesis machinery which is not present in vitro, leading to errors occuring in longer direct synthesis, additionally, a side-effect of chemical synthesis is that the acid used to remove the protecting group at the "leading edge" can also cause depurination of A and G bases, resulting in a molecule which can undergo hydrolysis cleaving and ruining the synthesized product.
Q: Why can’t you make a 2000bp gene via direct oligo synthesis?
A: long ssDNA can form secondary structures such as hairpins and G-complexes, leading to physical inhibition of oligo synthesis. This problem is amplified when considering that genes often have secondary and tertiary structures embedded within them for regulatory genomic interactions. Additionally, due to the afformentioned ineffficiency, undesired depurination and hydrolysis of synthesized oligos, and inability for verification that each nucleotide succesfully the increased number of reaction cycles required for the 2000bp oligo makes the production of a usable product virtually 0%. Finally, physical realities of the silicon chip reaction chambers means that the long ssDNA strands can clog up the chambers/pores, leading to failed synthesis and probably then require cleaning with a very very very very small chimney sweeper.
Homework Question from George Church: [Given slides #2 & 4 (AA:NA and NA:NA codes)] What code would you suggest for AA:AA interactions? wait maybe edit this so its AA interactions between proteins
A: The wording of this question is a bit ambiguous and so I am going to answer it assuming that the question is asking about the reconstruction of a gene from its amino acid sequence, as found in the reference on slide 4 (Acevedo-Rocha CG, Budisa N. Xenomicrobiology: a roadmap for genetic code engineering. Microb Biotechnol. 2016 Sep;9(5):666-76. doi: 10.1111/1751-7915.12398. Epub 2016 Aug 4. PMID: 27489097; PMCID: PMC4993186.)
In order to properly determine the NA code for an AA chain one would require a branching code, wherein the sequence branches at each AA which can be encoded by multiple codons. For instance, Arginine (R) can be encoded by AGG, AGA, CGA, CGC, CGG, and GCU. As such, the code for decoding an AA would require 6 branching paths every time Arginine occurs. While this may seem absurd, it is important to remember that the physical NA composition of a gene, although encoding redundancy, may also have altered physical/chemical structure, leading to altered epigenetic activation or protein interaction due to an altered physical/chemical landscape at the molecular scale.