Week 2 HW: DNA Read, Write & Edit
Week 2 Gel art, DNA processing;
I:
I used rcdonovan to iterate a design (after unselecting Ndel, PvulI and Xhol enzymes), that I could later modify to my liking. I used ↑and ↓ to change specific ladders.
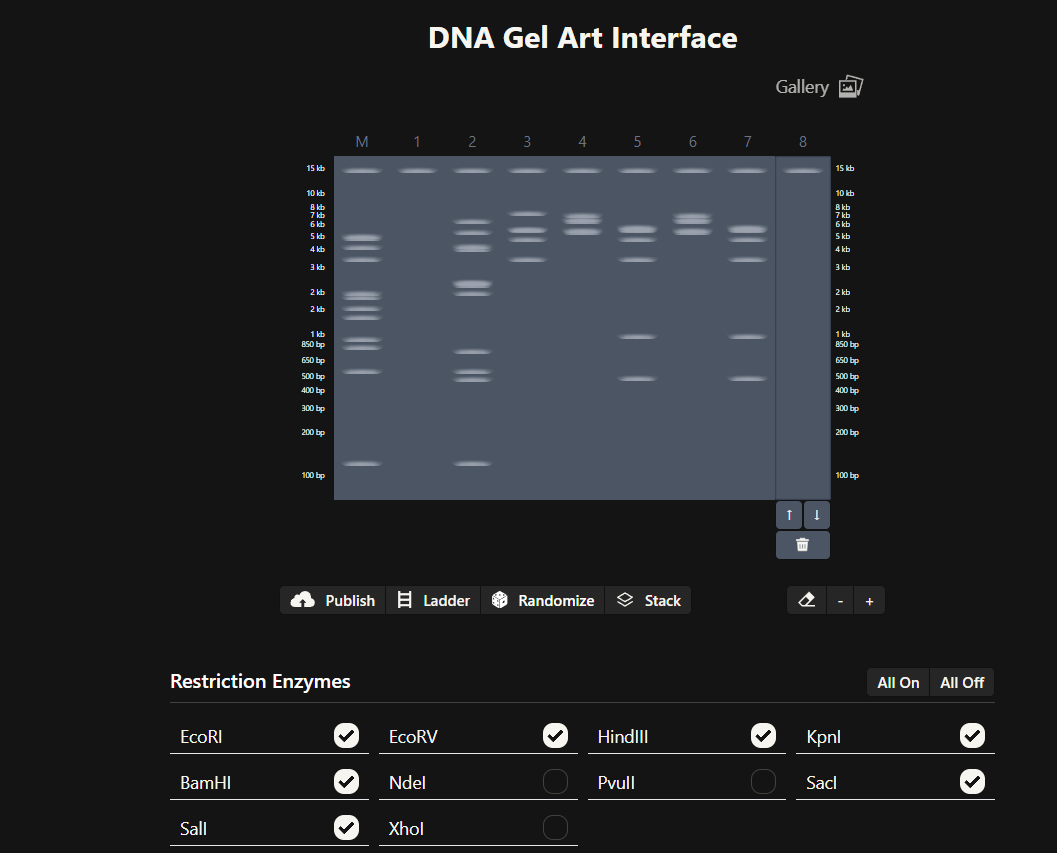

- I made an account on benchling, after importing the lambda DNA, I selected “digests” to make a total of 8 different ladders. I begun playing around with restriction enzymes. After which, I ended up with the following:
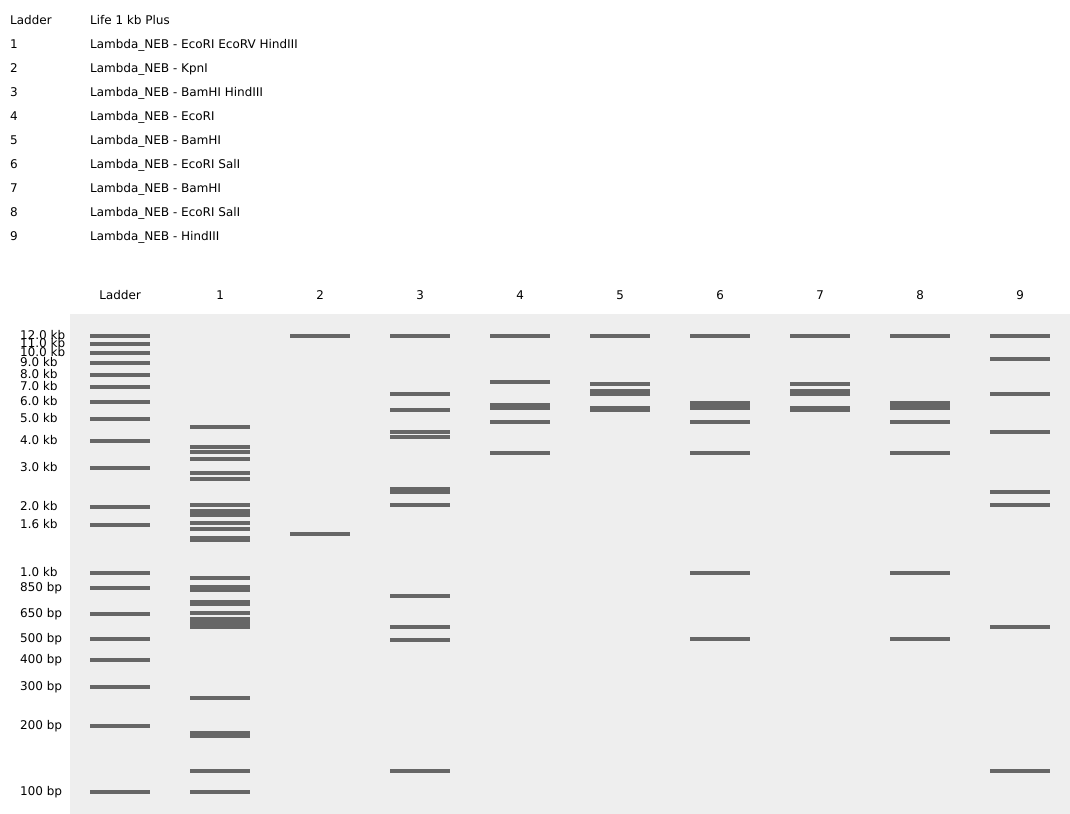

Ladder
- EcoRI, EcoRV, HindIII;
- KpnI;
- BamHI, HindIII;
- EcoRI;
- BamHI;
- EcoRi, SalI;
- BamHI;
- EcoRI, SalI;
- HindIII;
The gel art is supposed to resemble the letter “m”.
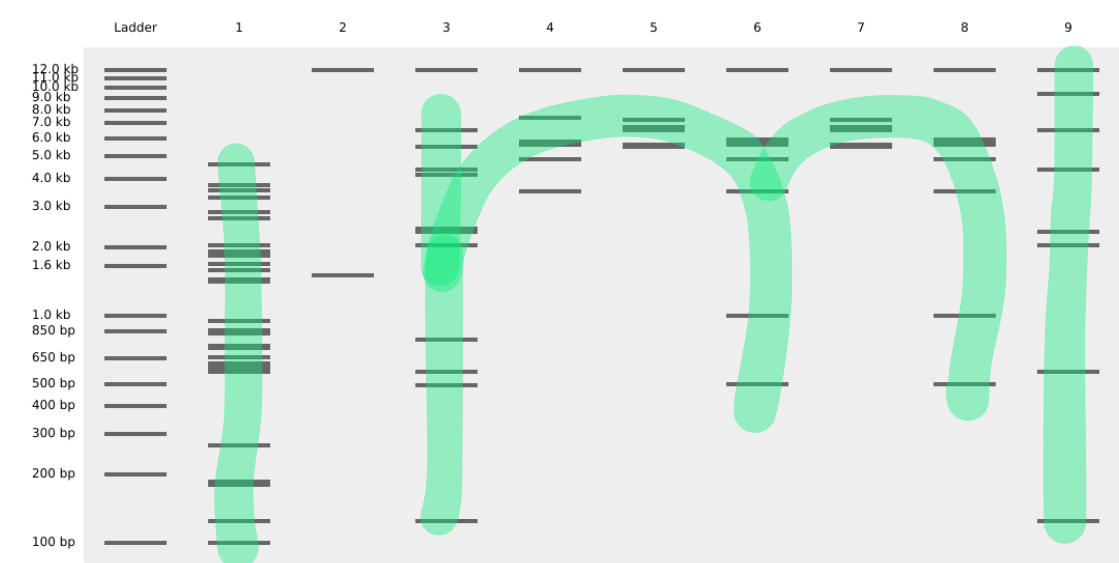

II:
With no access to a lab, I studied the protocol and took notes.
III:
3.1. I choose the beta hemoglobin subunit, constituent part of hemoglobin heterotetramer pigment, responsible for O2 transport in the human body. Hemoglobin is composed of four groups, 2 Alpha respectively 2 Beta, each having an iron-containing heme group (4 in total). The reason for my choice is that I find hemoglobin and its copper-containing homologue interesting.
sp|P68871|HBB_HUMAN Hemoglobin subunit beta OS=Homo sapiens OX=9606 GN=HBB PE=1 SV=2
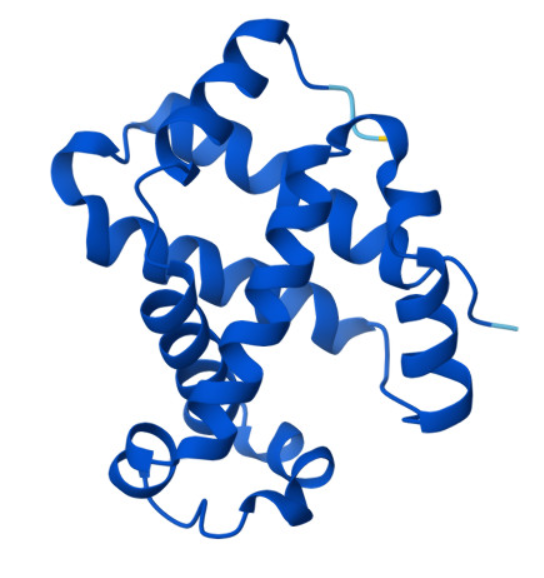

Next, I used uniprot to find HBB, homo sapiens, protein amino acid sequence:
uniprot-p68871
a.a. HBB seq:
MVHLTPEEKSAVTALWGKVNVDEVGGEALGRLLVVYPWTQRFFESFGDLSTPDAVMGNPKVKAHGKKVLGAFSDGLAHLDNLKGTFATLSELHCDKLHVDPENFRLLGNVLVCVLAHHFGKEFTPPVQAAYQKVVAGVANALAHKYH
3.2. Using Cusabio, I reverse translated the a.a. seq. into nucleotide seq.: reverse tranlation
nucleotide HBB seq:
ATGATGGTTCATTTAACTCCTGAAGAAAAATCTGCTGTTACTGCTTTATGGGGTAAAGTTAATGTTGATGAAGTTGGTGGTGAAGCTTTAGGTCGTTTATTAGTTGTTTATCCTTGGACTCAACGTTTTTTTGAATCTTTTGGTGATTTATCTACTCCTGATGCTGTTATGGGTAATCCTAAAGTTAAAGCTCATGGTAAAAAAGTTTTAGGTGCTTTTTCTGATGGTTTAGCTCATTTAGATAATTTAAAAGGTACTTTTGCTACTTTATCTGAATTACATTGTGATAAATTACATGTTGATCCTGAAAATTTTCGTTTATTAGGTAATGTTTTAGTTTGTGTTTTAGCTCATCATTTTGGTAAAGAATTTACTCCTCCTGTTCAAGCTGCTTATCAAAAAGTTGTTGCTGGTGTTGCTAATGCTTTAGCTCATAAATATCATTAA
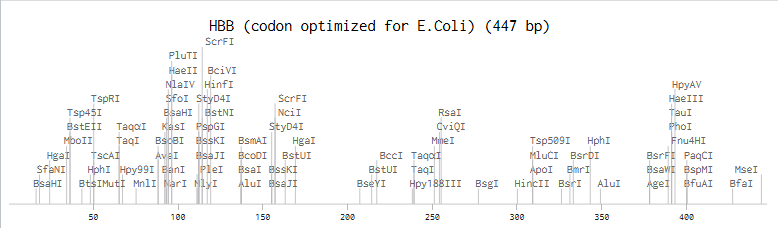
3.3. Because of its common usage as a vector “acceptor”, I chose to optimize the DNA sequence for Escherichia coli. I used Benchling’s option for optimization and obtained:
E. coli optimized:
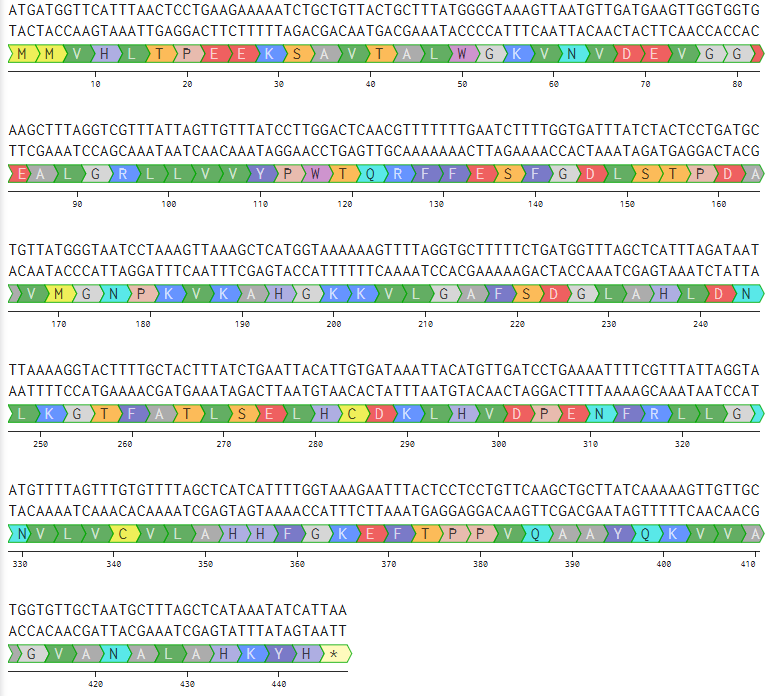
ATGATGGTGCATCTGACGCCGGAAGAAAAAAGTGCGGTGACCGCACTGTGGGGCAAGGTAAACGTCGATGAAGTCGGGGGCGAGGCTCTCGGGCGCCTGTTGGTGGTGTATCCCTGGACTCAACGGTTTTTTGAGAGCTTCGGAGACCTTTCTACCCCGGACGCGGTAATGGGAAACCCGAAAGTCAAAGCACACGGTAAGAAAGTGCTGGGCGCGTTTTCCGATGGCTTGGCTCATCTCGATAACCTGAAAGGTACGTTTGCCACCCTGTCGGAACTGCATTGTGATAAACTGCACGTTGACCCTGAAAATTTCCGTTTACTGGGCAATGTGCTTGTTTGCGTTTTAGCTCACCACTTTGGTAAAGAGTTCACACCACCGGTTCAGGCGGCCTACCAGAAGGTAGTTGCAGGTGTCGCCAATGCCCTAGCGCATAAATATCATTAA


- Also, its amino acid sequence is:
MMVHLTPEEKSAVTALWGKVNVDEVGGEALGRLLVVYPWTQRFFESFGDLSTPDAVMGNPKVKAHGKKVLGAFSDGLAHLDNLKGTFATLSELHCDKLHVDPENFRLLGNVLVCVLAHHFGKEFTPPVQAAYQKVVAGVANALAHKYH


3.4. For the DNA to successfully be transcribed and translated into my chosen protein, I could encode it into a plasmid, that I can insert into a bacteria such as E. Coli to naturally transcribe it using mRNA and later translate that RNA, using ribosomes, into the beta hemoglobin subunit. For increased efficiency I can use the optimized sequence to make up the vector.
3.5. A single gene can produce multiple proteins trough alternative processing of RNA transcript- alternative splicing: eukaryote DNA contains introns and exons, during splicing the exons are joined together while the introns get cut out. There is a chance for an exon to be cut out instead, resulting in a different sequence => different amino acids => completely different protein, or for an intron to be skipped and kept in the “mature” mRNA.
The same amino acid can be coded for by different codons (ex: ACG, ACA, ACT, ACC => Thr).



- T -> U
VI:
4.1. I set up my accounts.
4.2. I followed the instructions gave on HTGAA site.
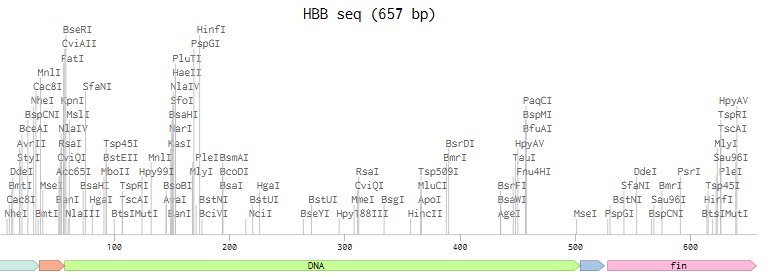

Optimized+ necessary exons:
TTTACGGCTAGCTCAGTCCTAGGTATAGTGCTAGCCATTAAAGAGGAGAAAGGTACCATGATGGTGCATCTGACGCCGGAAGAAAAAAGTGCGGTGACCGCACTGTGGGGCAAGGTAAACGTCGATGAAGTCGGGGGCGAGGCTCTCGGGCGCCTGTTGGTGGTGTATCCCTGGACTCAACGGTTTTTTGAGAGCTTCGGAGACCTTTCTACCCCGGACGCGGTAATGGGAAACCCGAAAGTCAAAGCACACGGTAAGAAAGTGCTGGGCGCGTTTTCCGATGGCTTGGCTCATCTCGATAACCTGAAAGGTACGTTTGCCACCCTGTCGGAACTGCATTGTGATAAACTGCACGTTGACCCTGAAAATTTCCGTTTACTGGGCAATGTGCTTGTTTGCGTTTTAGCTCACCACTTTGGTAAAGAGTTCACACCACCGGTTCAGGCGGCCTACCAGAAGGTAGTTGCAGGTGTCGCCAATGCCCTAGCGCATAAATATCATTAACATCACCATCACCATCATCACTAACCAGGCATCAAATAAAACGAAAGGCTCAGTCGAAAGACTGGGCCTTTCGTTTTATCTGTTGTTTGTCGGTGAACGCTCTCTACTAGAGTCACACTGGCTCACCTTCGGGTGGGCCTTTCTGCGTTTATA
I downloaded the sequence as a FASTA file;
4.3.& 4.4.& 4.5. Using Twist, I managed to import my sequence, transform it into pTwist Amp High copy vector. Downloaded it as GenBank construct and imported it to benchling. twist plasmid


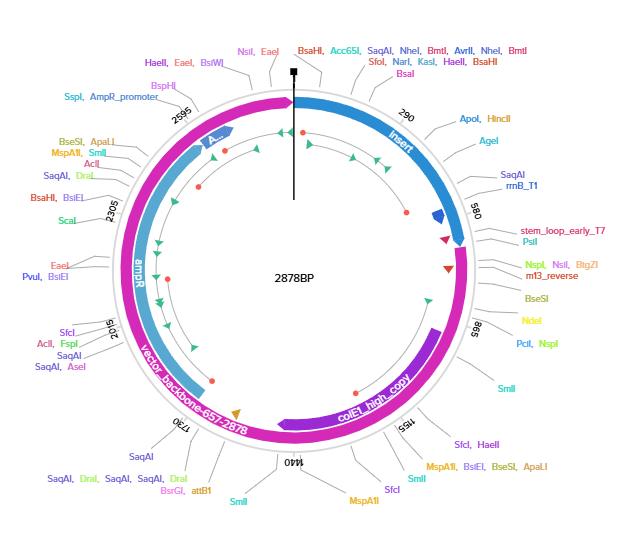

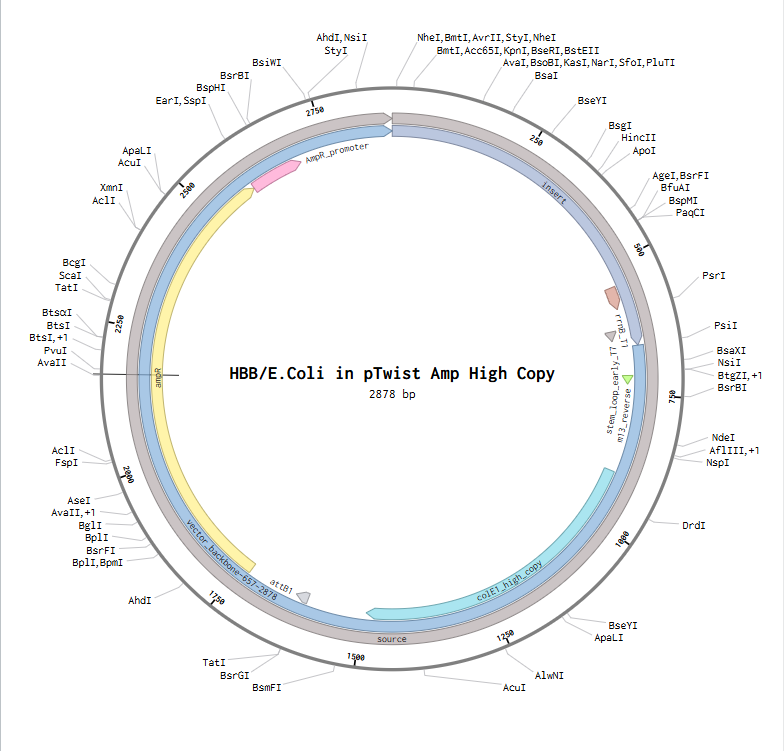

V:
5.1. My desired DNA to sequence is human reticular connective tissues’. As it might hide the pathway to synthesizing type 0 negative human blood in labs all around the world, making human blood donations unnecessary, furthermore, its DNA could play an important role in bone marrow synthesis.
tech: I would use Illumina sequencing-by-synthesis because of its high accuracy and high throughput (whole genome possible). I identified it to be second generation seq. tech. The input is the extracted genomic DNA from reticular connective tissue, the steps I’d follow are:
a) extraction, purification- get the DNA separated from fats, sugars, proteins and any other contaminates;
b) fragmentation into ~225 bp parts via enzymes; followed by end repair
c) adapter ligation- attached synthetic DNA sequences to both ends of each reticular DNA fragment for easier reading;
d) run genetic material through PCR machine for higher quality DNA, avoid nucleotide pools and get more material so I have more room for error.
NGS- second generation sequencing by synthesis technology
- Works by attaching a fluorescent dye to each nucleotide via polymerase enzyme and uploading information to a 4 color image of chip after each chemical flow and identify sequence after.
The primary output is a DNA nucleotide sequence.
5.2. The DNA sequence/ material I would synthesize is the phytogene found in coffee plants responsible for caffeine synthesis from xanthosine. It is not more efficient or cheaper than modern harvesting methods of caffeine or chemical lab synthesis, but it does have the advantage of less intermediary steps and purer resultants Caffeine can be used in medicine as treatment for sleep apnea, high yield purity is needed. In vector: synthesized DNA, attached to a Promoter, 7x His Tag, DNA seq to synthesize some chemical to incentivise bacteria to keep plasmid (e.q. gene made protein to make cell immune to antibiotic found in substrate), proposed coffee plant DNA, terminator. In substrate: xanthine+ ribofuranose => xanthosine, selected antibiotic.
tech: Twist to code for and buy plasmid. Later injected and accepted by bacteria (ex: E. Coli).
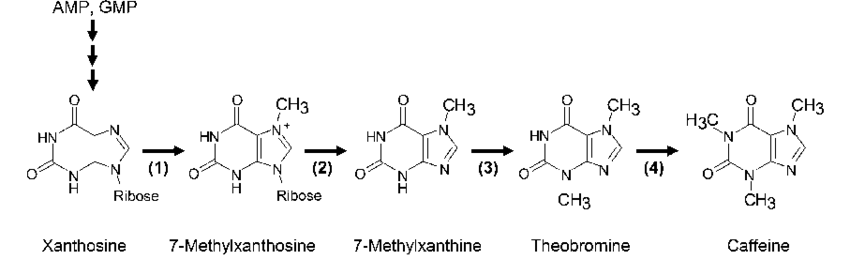
 plant metabolic pathway of creating “natural” caffeine
plant metabolic pathway of creating “natural” caffeine
5.3. The DNA I would edit would be found in bacteriophages. As virus’ genomes are the shortest; and because of their capability to reproduce fast, I could engineer virus DNA to fight against/ kill TB prokaryote. In third world countries, TB treatment is available, but not present enough to actually keep all population safe. An inoffensive virus spreading trough population only targeting TB could solve this century old problem.
tech: DRAG & DROP genome insertion mechanism- so that I’m not required to modify both strands of DNA and the whole genome, in turn.