Week 5 HW: Protein Design II
Week 5 Protein Design part II;
Superoxide dismutase 1 (SOD1) – cytosolic antioxidant enzyme that converts superoxide radicals (O2-) into H2O2 and O2. In its native state, it forms a stable homodimer and binds Co and Zn.
Mutations in SOD1 cause familial Amyotrophic Sclerosis (ALS). A4V leads to most aggressive forms of disease, as it destabilizes the N-terminus, perturbs folding energetics and promotes toxic aggregation.
A. SOD1 Binder Peptide Design
Part 1. Generate binders with PepMLM
P00441:
sp|P00441|SODC_HUMAN Superoxide dismutase [Cu-Zn] OS=Homo sapiens OX=9606 GN=SOD1 PE=1 SV=2
A.a. seq: MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
Induced mutation: (A4V)
MATKVVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
Using: PepMLM colab:
from hugging face
- set the length to 12 in cells [3] and [4];
- ran all codes;
| No. | Binder | Pseudo Perplexity |
|---|---|---|
| 0. | FLYRWLPSRRGG | (control) |
| 1. | WHSGATALAWKK | 9.649391 |
| 2. | KRYYAVVLEWGE | 30.275303 |
| 3. | WRYPAVVVRLGK | 14.739761 |
| 4. | WLYYPVAAAHKK | 12.779286 |
Part 2. Evaluate Binders with AlphaFold3
Using alphafold server:
Seed: 1935780713
ipTM score: 0.32
1. WHSGATALAWKK
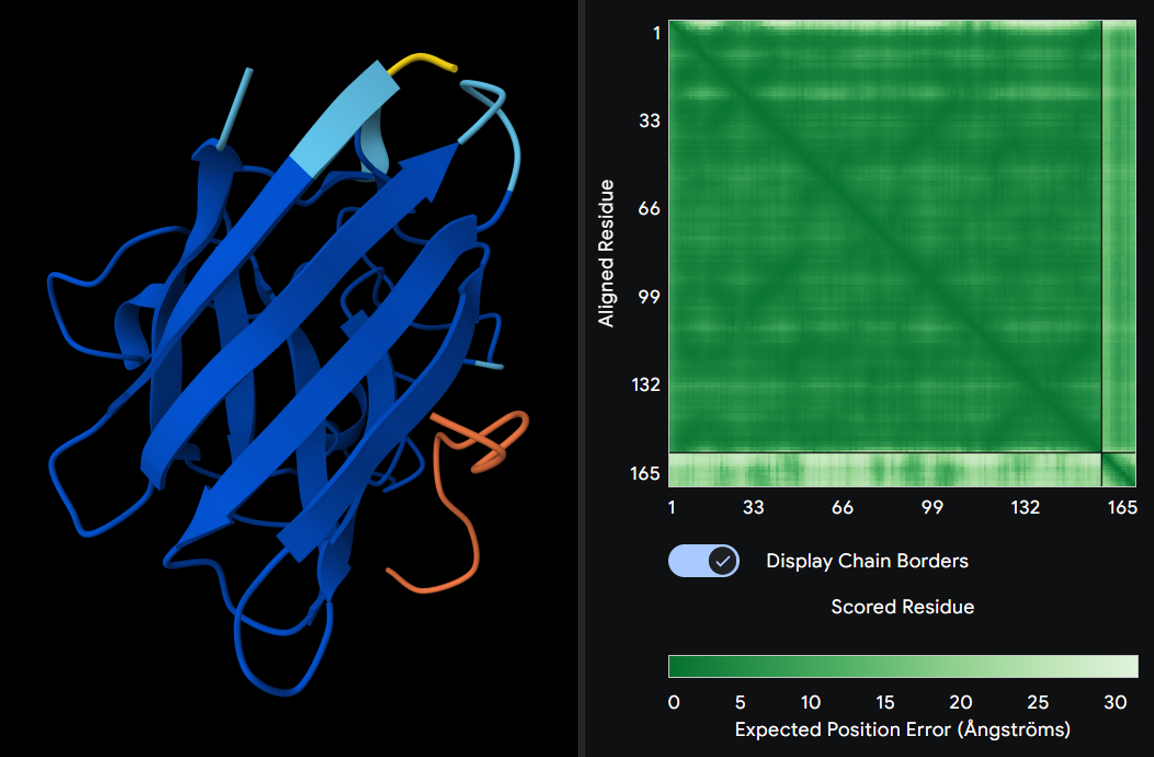

Seed: 1320704135
ipTM score: 0.4
2. KRYYAVVLEWGE
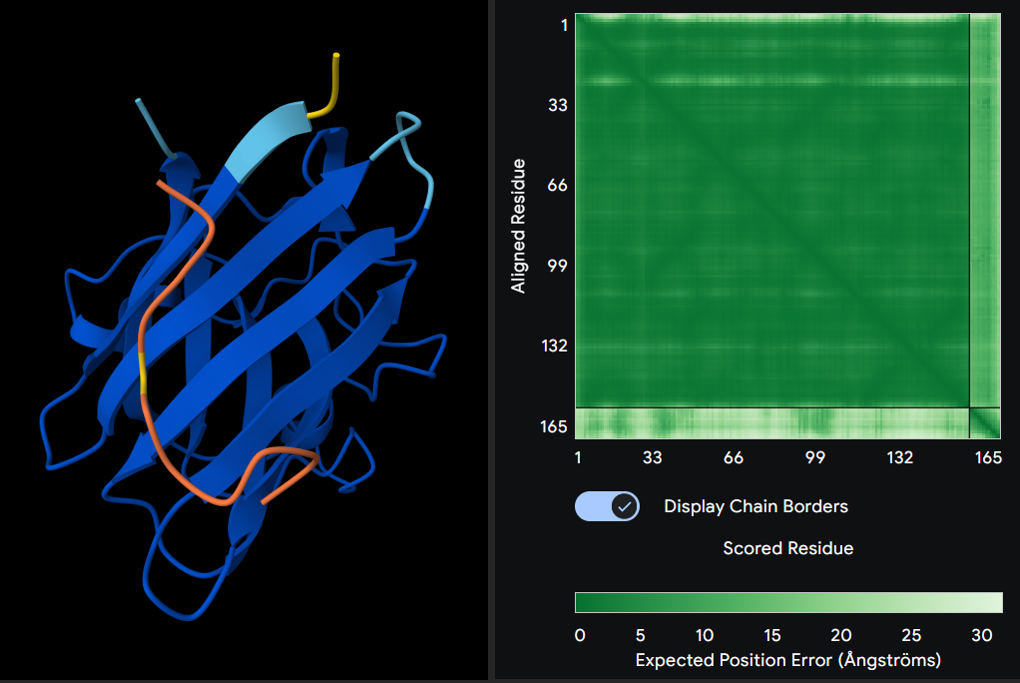

Seed: 105903932
ipTM score: 0.38
3. WRYPAVVVRLGK
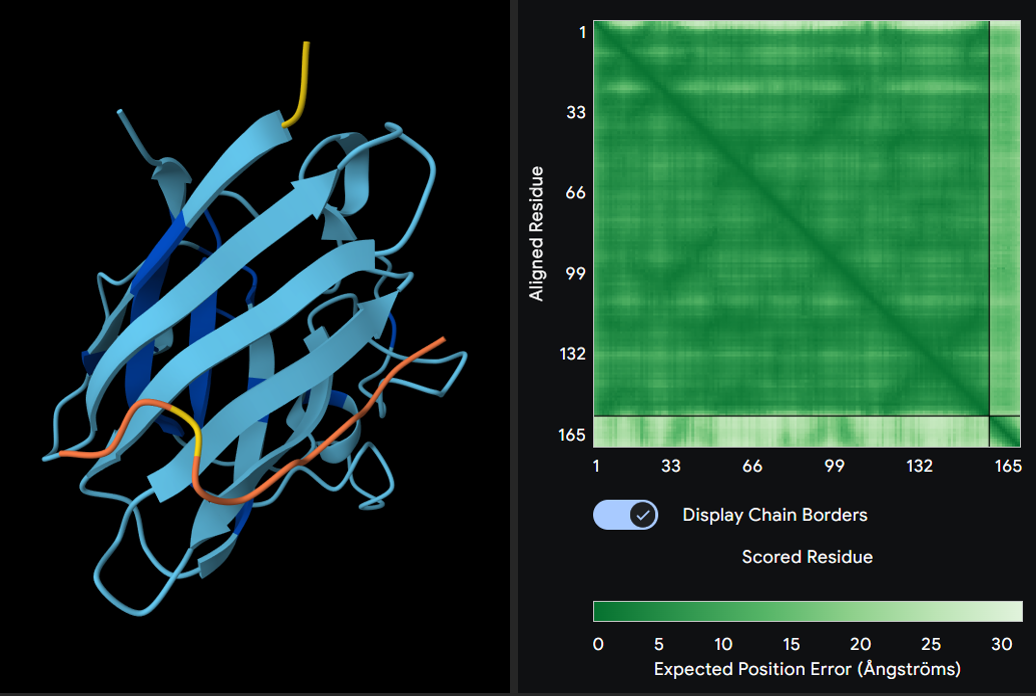

Seed: 1939890175
ipTM score: 0.31
4. WLYYPVAAAHKK
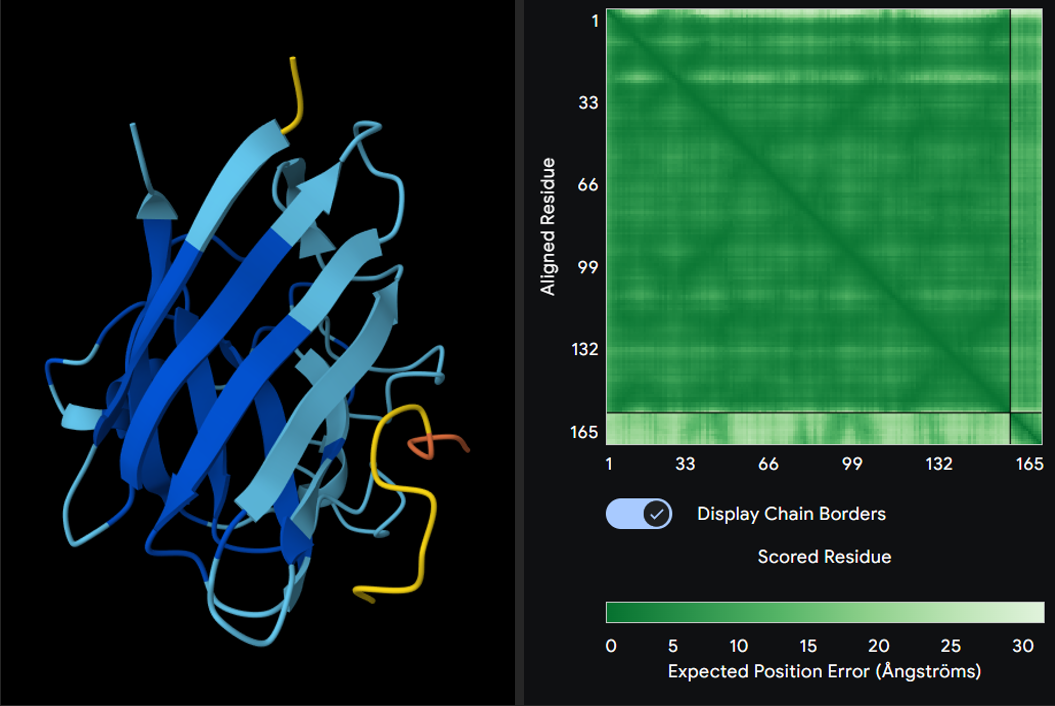

Seed: 25762303
ipTM score: 0.51
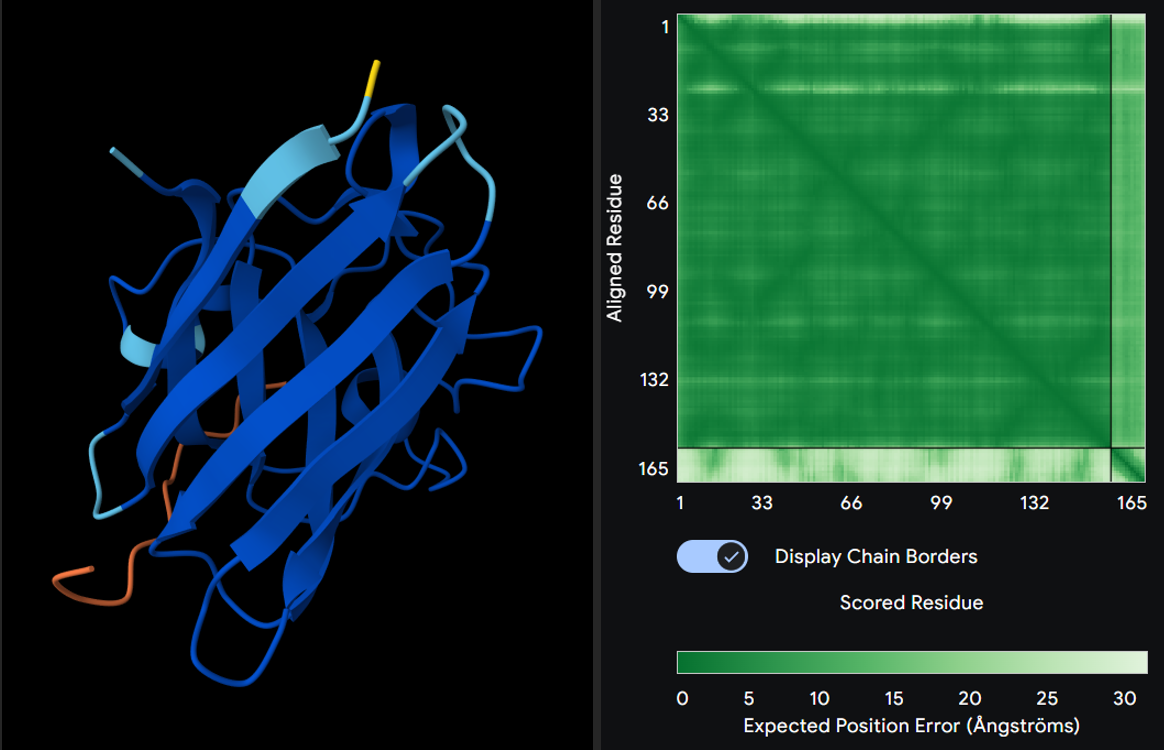
3. ipTM score
-> interface predicted template modeling. ipTM measures the accuracy of the predicted relative positions of the subunits within the complex. Values higher than 0.8 represent confident high-quality predictions, while values below 0.6 suggest likely a failed prediction. ipTM values between 0.6 and 0.8 are a gray zone where predictions could be correct or incorrect.
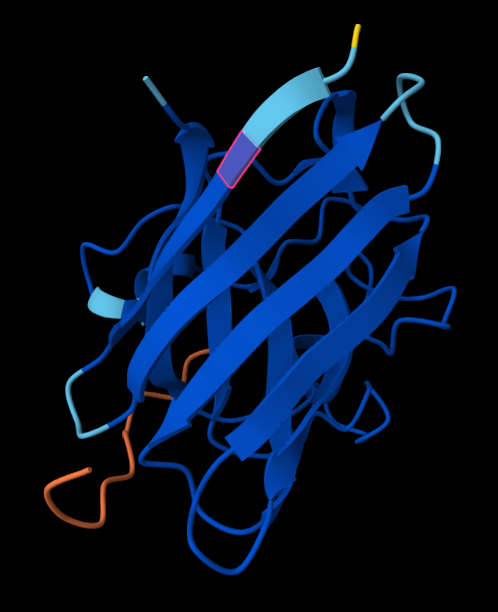
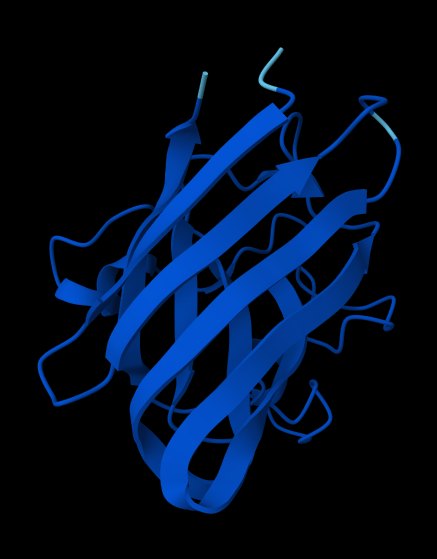
- where does the peptide bind?
Neither peptide seems to be closely bonded to SOD1. In reality, they may be weak forces acting throughout structure, facilitating functional bonds.
- does it localize near the N-terminus, close to A4V?
Except to 2nd peptide, the other 3 (+control) are proposed to not act close to the N-terminus of SOD1; it also doesn’t engage the beta-barrel region. All are surface-bound, further than expected from protein than I expected.
4. Describe ipTM values, observe if any PepMLM-generated peptide matched or exceeds the known binder?
They’re low, all of them appear to be under the threshold that could guarantee a definitive, reliable interaction prediction. AlphaFold’s server specifies that a ipTM score under 0.8 indicated a relatively inaccurate prediction. Thus, the peptides may actually bind in some way to SOD1 (there would be higher chances of a surface-bound cumulated Van Der Waals interactions). There also is the possibility for an erroneous generation of a binder peptide by the google colab.
Comparing strictly ipTM scores, AlphaFold is more confident in my last tested peptide (WLYYPVAAAHKK).
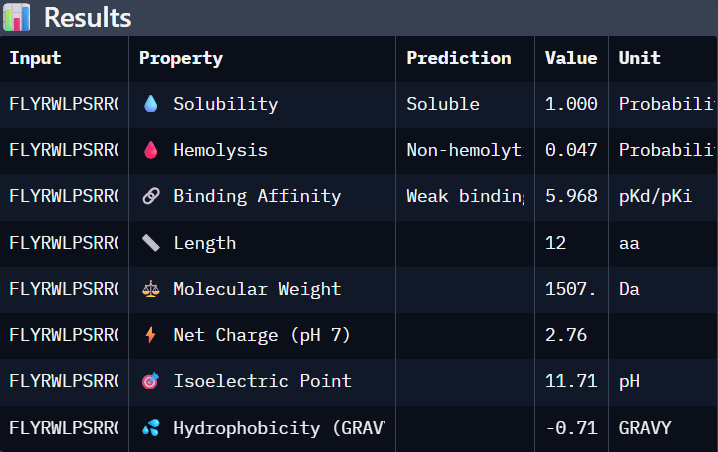
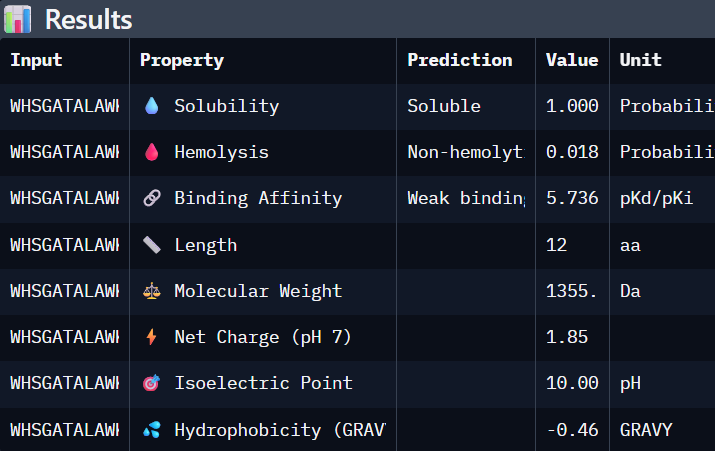
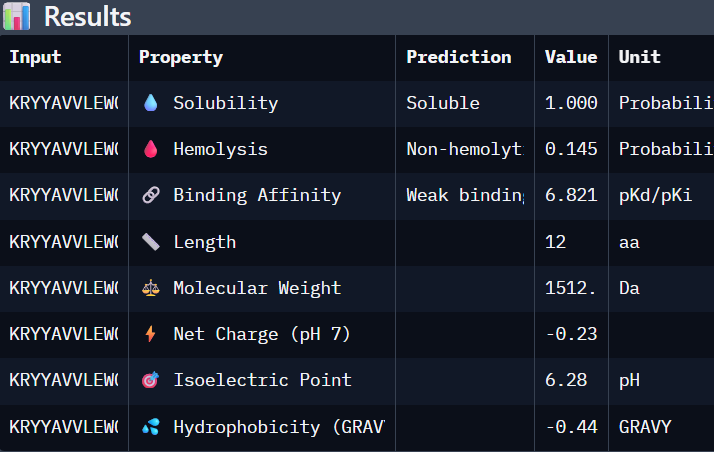
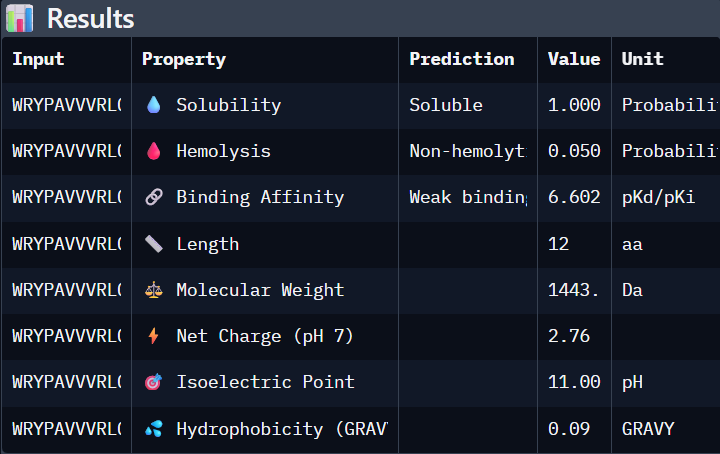
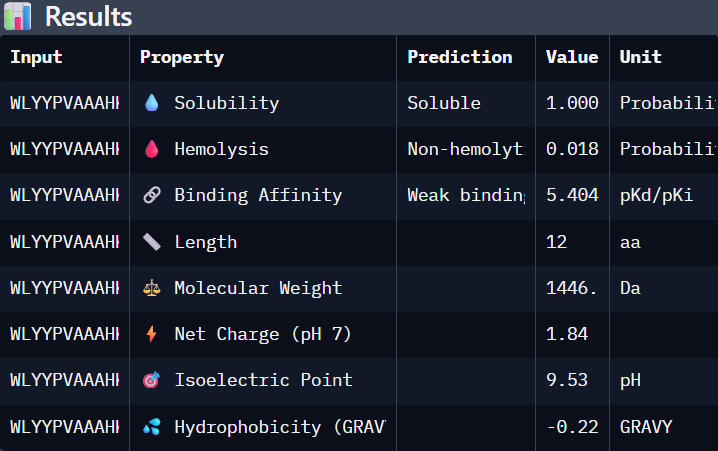
Part 3. Evaluate Properties of Generated Peptides in the PeptiVerse
Using PeptiVerse:
I selected “Calculate Basic Properties”, solubility, hemolysis, binding affinity and a neutral pH of 7.
Part 4: Generate Optimized Peptides with moPPIt
Using moPPIt colab
from hugging face
| Bind site | Binder | Hemolysis | Solubility | Affinity | Motif |
|---|---|---|---|---|---|
| 2-14 | NTKTCGERQQKV | 0.9684631042182446 | 0.9166666865348816 | 6.650517463684082 | 0.4602578580379486 |
| 14-26 | GYRKYFKEQFGS | 0.8697487562894821 | 0.8333333134651184 | 5.412135601043701 | 0.6805304288864136 |
| 81-93 | GKVCQRYFKKSE | 0.9514753371477127 | 0.8333333134651184 | 7.647891998291016 | 0.5069522857666016 |
| 93-105 | SKFKCEKISTKD | 0.9312182739377022 | 0.8333333134651184 | 6.648401260375977 | 0.5531454682350159 |
• Compared to PepMLM’s generated peptides, these ones are not as soluble (at least, based on the code’s prediction), more hemolytic. But overall, the ability to generate a peptide targeting a specific site/ surface may lead to improved results and better understanding of enzymatic activity of SOD1.
• I would probably use multiple softwares before advancing to wet lab testing and possibly clinical studies any generated peptide due to lack of confidence in generative models and chance of hemolysis. I find the second peptide to be the most safe, and even with its relatively small affinity score, there’s still chances of directed, promising protein binding.
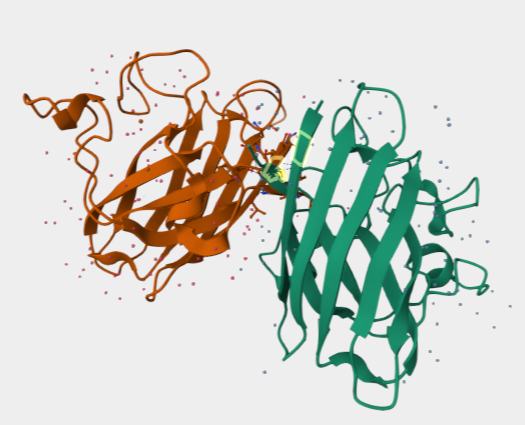
I was curious, so using alphafold:
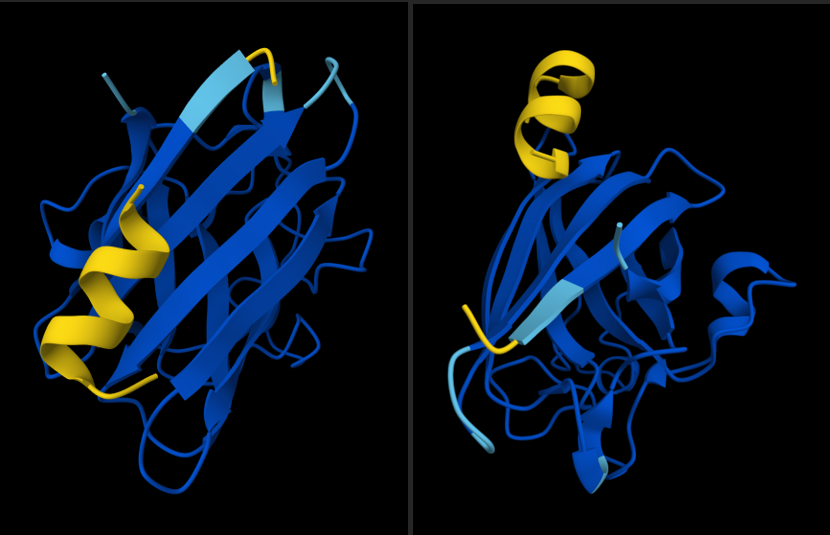

ipTM = 0.5 pTM = 0.89