Week 6 HW: Genetic circuits I
Week 6 Genetic circuits part I: Assembly Technologies;
Lab protocol questions:
1. What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
The Phusion HF PCR Master Mix is one of the “Media, Equipment and Consumables” from the protocol.
It consists of Phusion DNA Polymerase, deoxynucleotides and reaction buffer (containing MgCl2)
- Phusion DNA Polymerase- high fidelity (few errors) and robust performance (consistency).
- Deoxynucleotides- mix of standard nucleotides, but exclusive to DNA: A,C,T,G (“deoxy-“).
- Reaction buffer- used for maintaining optimal biochemical proprieties (such as pH) and ensure maximum enzyme activity (also includes essential ions for primer annealing).
2. What are some factors that determine primer annealing temperature during PCR?
Primer annealing represents the stage of primer bind to specific single-stranded DNA;
Mainly, annealing temperature is determined by individual primer melting temperature and chemical environment of reaction. Generally, it is determined approx. melting point of primer -5°C.
3. There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
PCR
- Description: Based on used DNA sequence, PCR determines cell-independent synthesis of copies of DNA.
Restriction enzyme digest
- Description: using restriction enzymes, there isn’t really much “creating” done, just cutting. Special enzymes are incubated along DNA sequences and act upon it, cutting it into smaller pieces based on presence and identification of special codon/ nucleotide short sequences. After undergoing electrophoresis, the different DNA segments spread and get predictably “digested” based on molecular weight.
| PCR | R.E. digest | |
|---|---|---|
| Media | nuclease-free water, primers, taq polymerase, dNTP mix, buffer | nuclease-free water, chosen enzyme, DNA dye (preferably fluorescent), TAE buffer |
| Equipment | PCR machine setup, PCR tubes, micropipettes | electrophoresis setup, (..and a PCR setup) |
| Accesbility (1-5) | 3.5 | 2 |
| Cost- protocol/sample (avg.) | 5€ | 4€ |
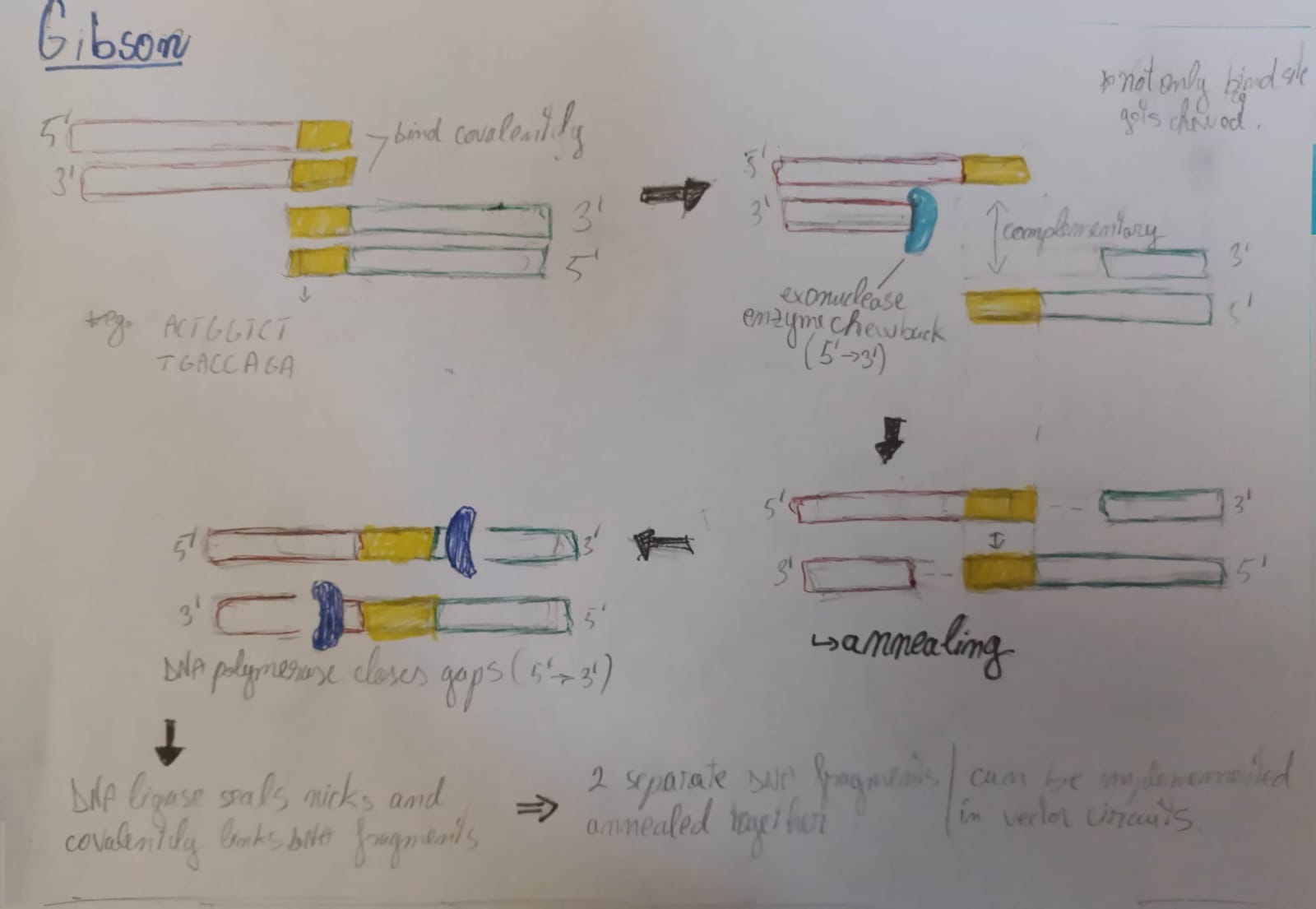
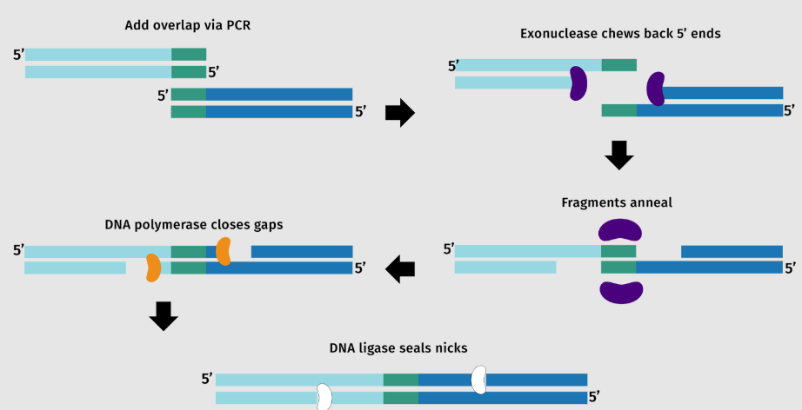
Both serve the purpose of vector linearization, making Gibson assembly mechanism possible.
4. How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
First of all, the sequences need to have blunt identical ends so that it can be cut and ligated. And so, DpnI digestion may be applied to ensure that methylated parental DNA templates are selectively degraded; gel electrophoresis could also serve as a way to check DNA purity.

5. How does the plasmid DNA enter the E. coli cells during transformation?
Through pores, ruptures in cell membrane created via thermic shock or electroporation (under the act of diffusion).
6. Describe another assembly method in detail (such as Golden Gate Assembly)
1. Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
2. Model this assembly method with Benchling or Asimov Kernel!
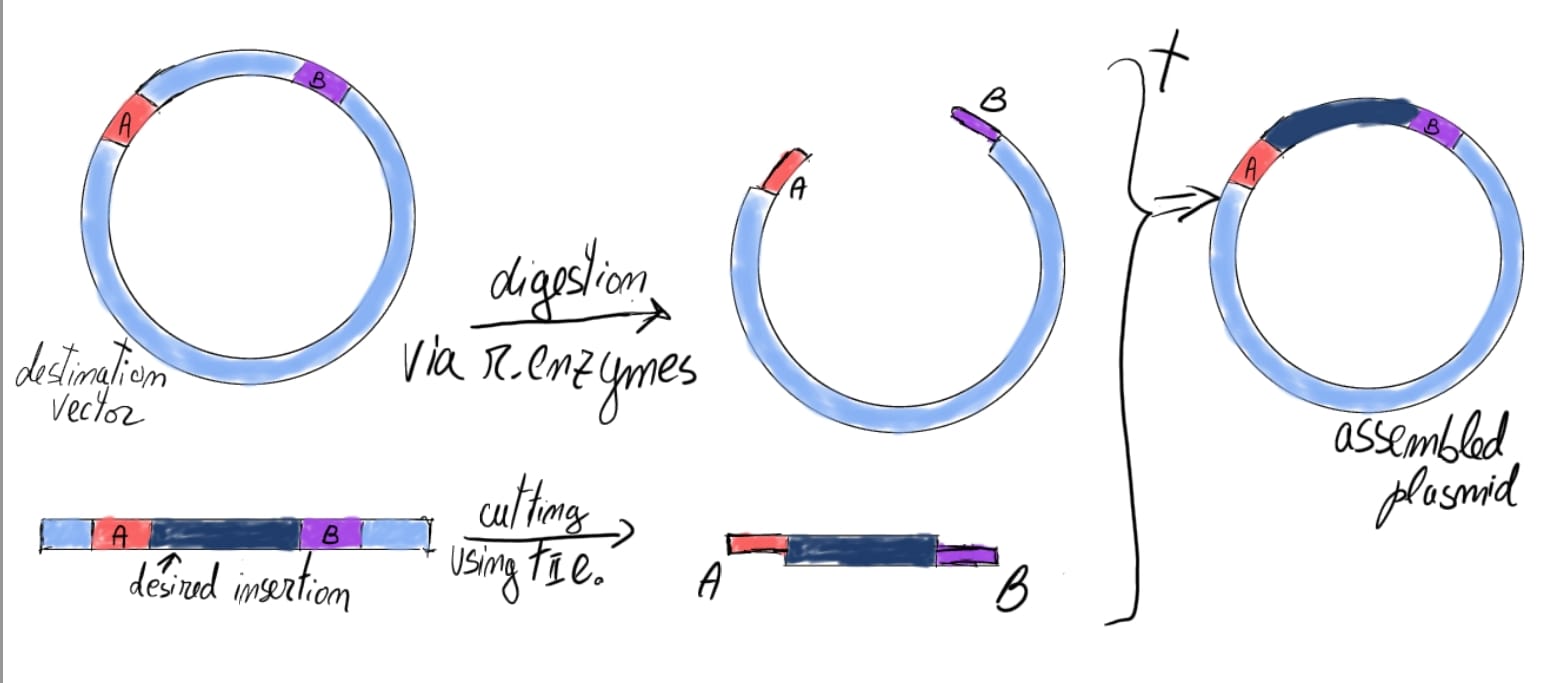
Golden Gate Assembly
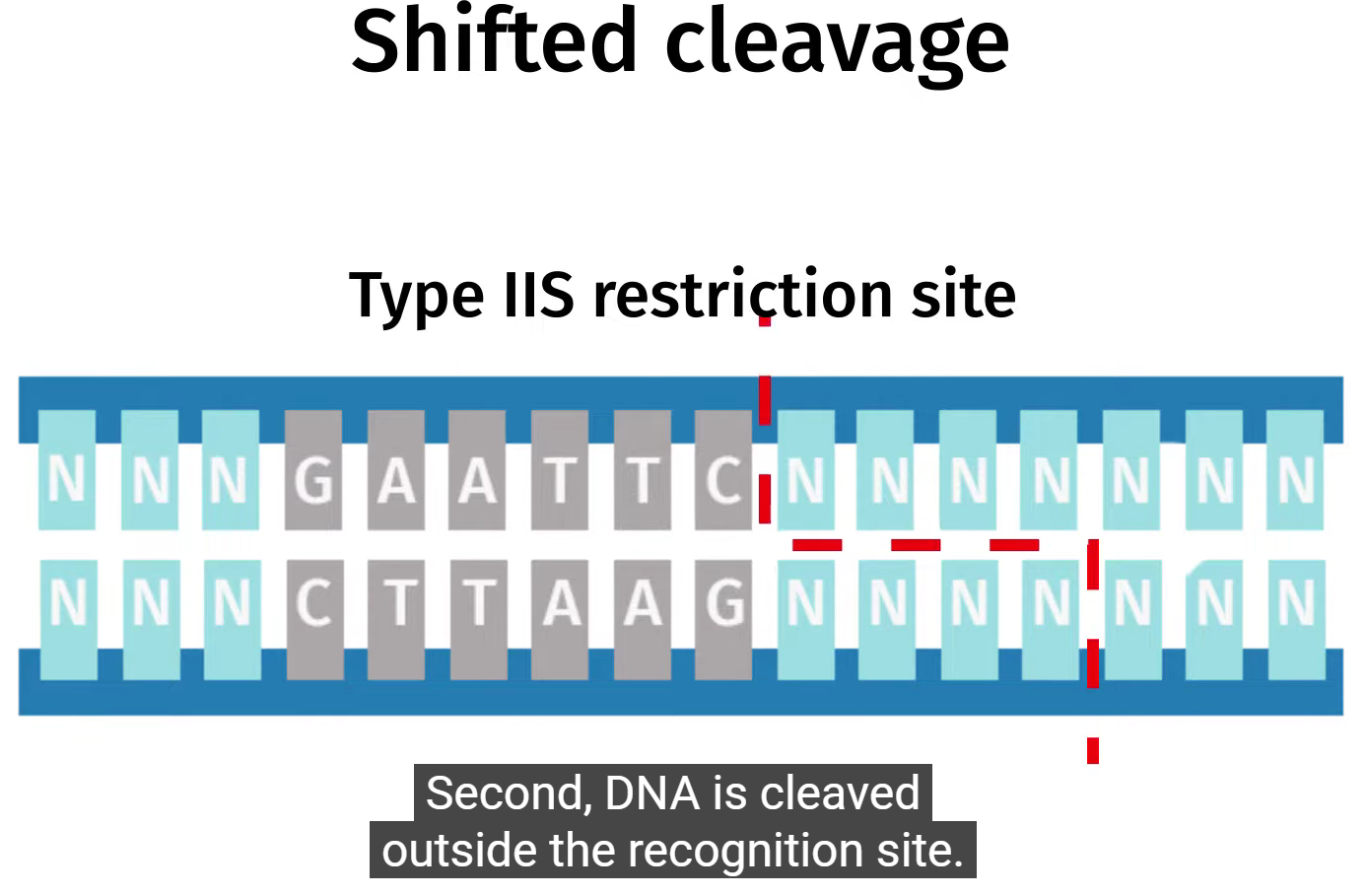
Golden Gate assembly is a highly efficient DNA cloning method. It uses type II enzymes that “cut” DNA consequent to recognized nucleotide sequence (usually palindromic), mostly after 4-5 nucleotides-post recognition site; and are dependent of Mg2+ as a cofactor. They act on both the plasmid and fragment pretty much the same; thus, the digested insertion fragment covalently binds to plasmid overhang complementary DNA strand.
Compared to Gibson a. it doesn’t require polymerases to build up lacking DNA strand fragments, but it does usually include ligase enzymes to ensure vector stability and successful annealing. G.G. assembly is preferred when working with many insertion fragments (30-50) because of its simultaneous activity of ligases and restriction enzymes.

Type II restriction enzyme example:
BSAI (takes 5-15 mins to digest PCR sample)
5’ ..GGTCTC (N1) | .. 3'
3’ ..CCAGAG (N5) | .. 5'
=> 4 base overhang