Week 9 HW: Cell-Free Systems
Week 9 Cell-Free Systems;
General homework questions
1. Explain the main advantages of cell-free protein synthesis over traditional in vivo methods, specifically in terms of flexibility and control over experimental variables. Name at least two cases where cell-free expression is more beneficial than cell production.
Flexibility: substrates, cofactors, inhibitors and many other factors can be directly manipulated as CFPS doesn’t really on a cell’s ability to stay alive. Toxic or “non-physiological” substances (D-a.a., HCN etc.).
Efficiency: there is no need for cloning, long growth phases (time related factors) or promoting substances to increase yield (eq: cytokines) to assure synthesis. Also, there’s less work needed- no codon-optimization needed, chaperones, so there can be more reactions ran in parallel with different conditions.
CFPS can lead to endless possibilities when it comes to amino acid/protein synthesis, not only being able to extend a bit over the 20 natural a.a., but actually building up proteins with who knows with what actions formed from any a.a. imaginable (H-,COOH-,NH2-,R-C), also, the synthesized proteins can have any propriety or cytotoxicity attributed as there’s no cell membrane (or viability in general) to care about.. More complex labeling is also easier as tRNA charging, energy mix and modification enzymes can be directly tuned, without interfering the cell’s native metabolism.
All system energy is channeled to one/ more proteins of interest and just that.
2. Describe the main components of a cell-free expression system and explain the role of each component.
Cell extract composed of the molecular machinery and cofactors need for reactions: Ribosome ribonucleic complex, RNA polymerase, other desired proteins. RNA p. is simply the enzyme that makes up corresponding tRNA to given DNA (TX); the RRNC sustains TL reactions by translating the tRNA in proteins; other proteins may be added to enhance target protein synthesis structure, function or TX-TL sustainability. Eq: translation factors (IF, EF, RF) are essential GTP-dependent proteins driving bacterial protein synthesis.
Template DNA/mRNA, amino acids and nucleotides/ nucleoside triphosphates (ATP, GTP, CTP, UTP) are also provided in most cases for obvious reasons.
Other components include: salt glutamates (K, Mg) and other ions to stabilize ribosome function and RNA structure.
3. Why is energy provision regeneration critical in cell-free systems? Describe a method you could use to ensure continuous ATP supply in your cell-free experiment.
Protein synthesis is energetically expensive, each peptide bond and each step of translation consume ATP and GTP; without constant regeneration, ATP and GTP are rapidly depleted, leading to early unwanted termination of TX-TL reactions and very low yields. n traditional cell-free protein synthesis reactions, the energy source (typically phosphoenolpyruvate (PEP) or creatine phosphate) is the most expensive substrate. However, for most biotechnology applications glucose is the preferred commercial substrate.
Article demonstrates how G6P (glucose-6-phosphate) can be used as an cheaper alternative to PEP; G6P –(phosphoglucose isomerase; -2Na)-> fructose-6-phosphate –(phosphofructokinase-1)-> fructose-1,6-bisphosphate –(aldolase- cleave/ break down)-> G3P ->..-> phosphoenolpyruvate (PEP) –(Pyruvate kinase)-> ADP->ATP, pyruvate;
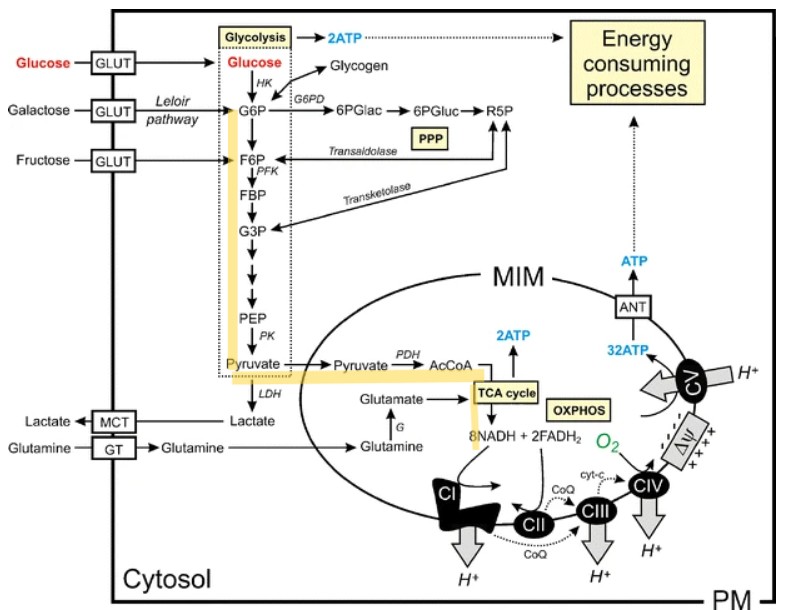

pretty much this but less hardcore, because CFS. note: there are more reactions that take place such as 3PGA ⇌ 2PGA via phosphoglycerate mutase
So, from what I’ve read, there’s an undeniable need for a substrate and some enzymes to catalyze reactions to ATP, the further you go from phosphorylation, the more enzymes are needed.
4. Compare prokaryotic versus eukaryotic cell-free expression systems. Choose a protein to produce in each system and explain why.
| CFS / POV | prokaryotic | eukaryotic |
|---|---|---|
| speed (ΔT) | really fast (hours) | slower (complex processing needed) |
| yield (μ) | high | moderate |
| post-TL modifying | mostly absent | possible (glycolysis, phosphorylation) |
| protein folding | high risk of error with complex proteins | better |
| protein synthesis | GFP | antibody |
| why | small, robust, no post-TL changes needed, more efficient to syn. in prok. | needs complex folding mechanisms and assembly |
5. How would you design a cell-free experiment to optimize the expression of a membrane protein? Discuss the challenges and how you would address them in your setup.
Membrane proteins need a hydrophobic medium to fold properly and remain soluble. So I would most probably adjust to imposed settings- add lipids/liposomes, nanodiscs (membrane patches with scaffold proteins). But,
a. proteins may not insert into lipids spontaneously (=> decreased yield, waste build-up)- mimic cellular insertion mechanism.
b. protein synthesis requires ribosomes and hydrophobic a.a. => proteins aggregate and result in inclusion bodies (useless)- detergents (eg. DDM) can form shielding micelles around protein hydrophobic parts to keep them soluble.
6. Imagine you observe a low yield of your target protein in a cell-free system. Describe three possible reasons for this and suggest a troubleshooting strategy for each.
- Genetic material template degraded, contaminated with salts or not enough. Scale up concentration template.
- Non-optimal reaction conditions, there are a lot of factors that impact CFS (as well as CDS, but cells can mediate themselves considerably)- ions, temperature, energy, pH, redox states, other reagent wrong concentration; follow protocol or change protocol, accordingly.
- Degradation of protein; depends a lot..
Homework question from Kate Adamala
Design an example of a useful synthetic minimal cell as follows:
1. Pick a function and describe it.
a. What would your synthetic cell do? What is the input and what is the output?
Optimize immune responses towards Helicobacter pylori. It detects CagA or VacA oncoproteins specific to pylori bacteria via surface-bound antibodies/ Toll-like receptors: antibody-induced clustering of surface receptors (eg. BCR, Fc, or RTKs) => activates associated tyrosine kinases to initiate downstream signaling pathways, which directly phosphorylate translation components => protein synthesis.
b. Could this function be realized by cell-free Tx/Tl alone, without encapsulation?
No, the DNA wound get transcribed randomly and lead to autoimmune disease activation; there’s no trigger protein or reusability.
c. Could this function be realized by genetically modified natural cell?
Maybe, but it would be too hard to modify everything as I want it to be and also keep alive and efficiently running.
d. Describe the desired outcome of your synthetic cell operation.
H. pylori survives by using its virulence factors (such as CagA and VacA) to evade immune detection and modulate the response. It can induce regulatory T cells (important for avoiding autoimmune responses in humans) that suppress the effective Th1/Th17 responses, leading to chronic inflammation rather than elimination. So, by attaching specific CagA/ VacA receptors on prostatic cell’s surface, it can only get simulated in the presence of a pylori infection and help the immune system synthesize ligands that bind to a part of regulatory cytokines such as IL-10, TGF-beta (downregulate T cell activity) and proliferation, promoting (immune-boosts) cytokines such as IFN-gamma, IL-6 or IL-2 (doesn’t provide inflammatory response).
2. Design all components that would need to be part of your synthetic cell.
a. What would be the membrane made of?
Chitin to withstand gastric acid low pH (<2), inspired by fungi cell wall.
b. What would you encapsulate inside? Enzymes, small molecules.
Template DNA, enzymes responsible for the phosphorylation cascade from the antibody to nucleus/nuclei; enzymes for protein assembly and a synthetic ER + Golgi apparatus (for ILs syn, maturation and transport) to assure correct protein folding and efficiency.
c. Which organism your Tx/Tl system will come from? Is bacterial OK, or do you need a mammalian system for some reason? (hint: for example, if you want to use small molecule modulated promotors, like Tet-ON, you need mammalian)
Possibly long-lived plasma cells’ DNA to get a comprehensive idea of how the cytokines need to be synthesized and maybe fungi to ensure some membrane regeneration activity. But for efficiency, I think bacterial system templates can be used as well as ILs, IFNs aren’t that hard to translate, and the surface-attached antibodies/ other receptors can be placed then when the syn cell is fabricated.
d. How will your synthetic cell communicate with the environment? (hint: are substrates permeable? or do you need to express the membrane channel?)
ILs can mostly pass the cell membrane fairly easily, ligand proteins not so much as they’re big (at least they’re not polar) and get caught up in the lipid bilayer. In biological cells they get transported in a vesicle that pops outside of cell; the whole process is driven by ER ad GA.
3. Experimental details
a. List all lipids and genes. (bonus: find the specific genes; for example, instead of just saying “small molecule membrane channel” pick the actual gene.)
To be feasible: there are high chances of promoting autoimmune responses, but because it’s easier to transport I’ll go with IL-2 and IFN gamma, (no ligands involved). I still depend on GA and ER presence and function and other phosphorylation proteins/ small molecules, lipids and enzymes.
b. How will you measure the function of your system?
Sensitivity + specificity to oncoproteins; Synthesis capabilities.
Homework question from Peter Nguyen
Freeze-dried cell-free systems can be incorporated into all kinds of materials as biological sensors or as inducible enzymes to modify the material itself or the surrounding environment. Choose one application field — Architecture, Textiles/Fashion, or Robotics — and propose an application using cell-free systems that are functionally integrated into the material. Answer each of these key questions for your proposal pitch:
• Write a one-sentence summary pitch sentence describing your concept. • How will the idea work, in more detail? Write 3-4 sentences or more. • What societal challenge or market need will this address? • How do you envision addressing the limitation of cell-free reactions (e.g., activation with water, stability, one-time use)?
Homework question from Ally Huang
1. Provide background information that describes the space biology question or challenge you propose to address. Explain why this topic is significant for humanity, relevant for space exploration, and scientifically interesting. (Maximum 100 words)
Jupiter’s moon, Europa has liquid water on its surface and many substantial elements and may hide alien life. But, I think it may actually be able to host human life in the distant future by changing its atmosphere and overall proprieties. The main problems are extreme cold, intense radiation from Jupiter’s magnetosphere and low gravitational pull.
2. Name the molecular or genetic target that you propose to study. Examples of molecular targets include individual genes and proteins, DNA and RNA sequences, or broader -omics approaches. (Maximum 30 words)
• IgG antibodies and their enzymatic proprieties; sequenced mature plasma B cell DNA to replicate; • Methanogenic archaea - Methanococcus jannaschii (deep-sea hydrothermal vent dweller) structure and metabolic processes.
3. Describe how your molecular or genetic target relates to the space biology question or challenge your proposal addresses. (Maximum 100 words)
IgG is way simpler and maybe more efficient to synthesize using BioBits and be put into action. The main problem is its folding, as it requires complex modular assembly of two heavy and two light chains into the “Y” shape. I couldn’t find an a.a. seq specific for IgG with the variable domain specific for catalyzing H2O into O3, but sequencing a plasma B cell genome, CFS can replicate TX-TL and synthesize target protein (~150 kDa structure).
4. Clearly state your hypothesis or research goal and explain the reasoning behind it. (Maximum 150 words)
Europa’s atmosphere and overall hostability can be increased by increasing its atmosphere ability to absorb radiation and keep heat in by using a gas such as methane (CH4), or ozone (O3). They not only are greenhouse gases that keep warm from diffusing off the moon’s surface, but can also absorb radiation and a large enough quantity may actually make Europa way less dangerous. Using a methanogenic cell’s metabolic pathway as a template, CO2 can be indirectly turned into CH4; and endogenous ozone production present in immune sys. to turn H2O (deep under all the ice) into O3. A CFS can achieve this without the need of life sustaining factors.
5. Outline your experimental plan - identify the sample(s) you will test in your experiment, including any necessary controls, the type of data or measurements that will be collected, etc. (Maximum 100 words)
I need to be sure that CFS can get under the ice without usage of electromagnetic devices to warm up and have the substrate to be able to work. Also, it needs to synthesize a lot of proteins to sustain the CH4 metabolic pathway, or be able to act as a catalyzer for the H2O+O2-> H2O3 reaction, that later decomposes into O3 and H2O2 (that turns into O2 and H2O photochemically). I need to somehow experiment self-sustainability. ER action at -200°C or substituent efficiency and feasibility.