Week 11 HW: Bioproduction & Cloud Labs
Week 11 Bioproduction, cloud-autonomous laboratories;
A: The 1,536 Pixel Artwork Canvas | Collective Artwork
*I guess I’m more the destructive type as I had more fun deleting pixels than I had painting the canvas (only after the experiment ended tho!!) ;)


–


It was such a nice and unexpected initiative, I also liked hearing about the million dollar project and r/place. I’m looking forward to what’s going to happen next in the context of the “1536 Pixel Artwork” :)
I would have liked a shorter cooldown as I’m not really the most patient person ever, and a more diverse colour palette.
B: Cell-Free Protein Synthesis | Cell-Free Reagents
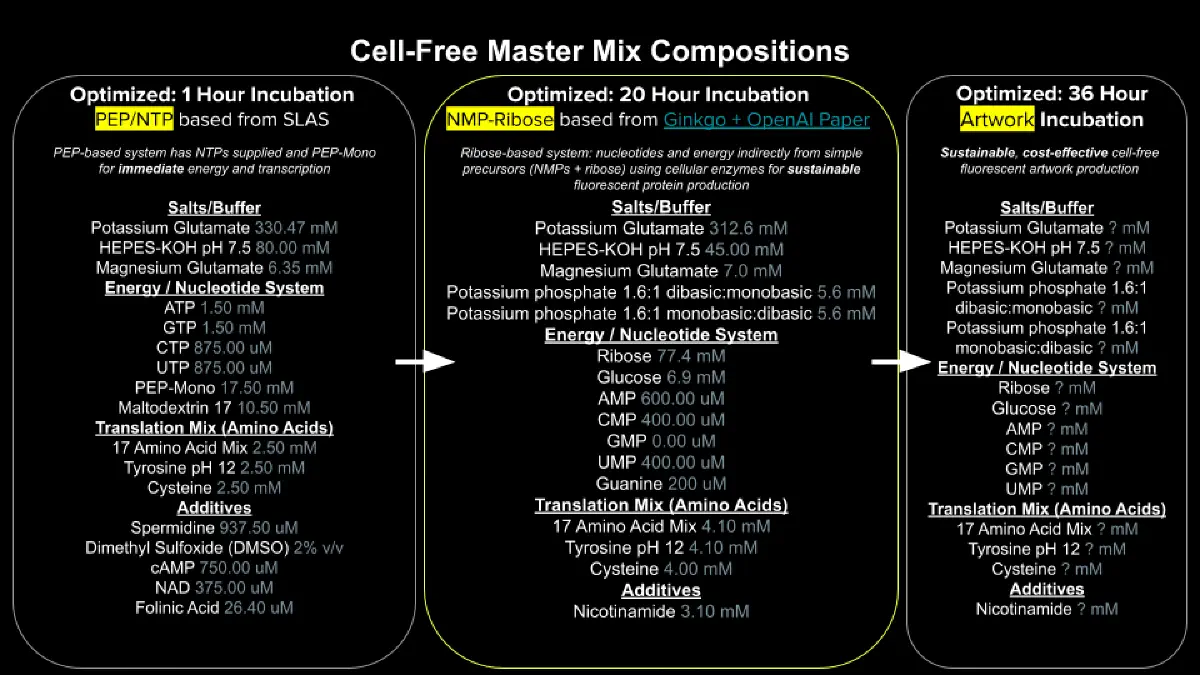

- Each component’s role in the cell-free reaction.
E. coli Lysate
BL21 (DE3) Star Lysate (includes T7 RNA Polymerase)
- core Tx/Tl mechanism drivers: ribosomes, tRNA, relevant enzymes and sometimes E. coli assigned cofactors; T7 specific polymerase that attaches to given DNA and transcribes it into mRNA (for high-copy T7 promoters).
Salts/Buffer
Potassium Glutamate
- determines ionic “strength”, mimics intracellular conditions to stabilize ribosomes and other protein enzymes.
HEPES-KOH pH 7.5
- reaction buffer; keeps pH optimal for enzymatic activity.
Magnesium Glutamate
- Mg2+ (cofactor) source, important for ribosome, polymerase activity and nucleotide interactions
Potassium phosphate monobasic
- KH2PO4: one potassium atom, retains 2 acidic hydrogens; acts as the more acidic than dibasic p.p., lowers pH is solution (acidity corrector) => buffer + phosphate for metabolic reactions and nucleotide synthesis. pH~4.1-4.5 at 25°C.
Potassium phosphate dibasic
- K2HPO4: 2 potassium atoms, one acidic hydrogen atom; more basic, can raise pH, same importance as p.p. monobasic. pH~8.5-9.6 at 25°C.
Energy / Nucleotide System
Ribose
- carbon 12 source used to regenerate nucleotides via ‘salvage pathways’.
Glucose
- primary energy source that gets worked down to ATP through glycolytic pathways by enzymes in the lysate.
Ribonucleic monophosphate (NMPs) – ribose sugar, phosphate group, nitrogenous base = AMP, CMP, GMP, UMP.
AMP
- precursor to ATP, incorporated into RNA as adenosine, purine base.
CMP
- incorporated into RNA as cytosine, pyrimidine base.
GMP
- can be converted to GTP- important for Tl and RNA synthesis, incorporated into RNA as guanine, purine base.
UMP
- incorporated into RNA as uracil, pyrimidine base.
Guanine
- base precursor that can be converted to GMP via salvage pathways, supplements nucleotide pools.
Translation Mix (Amino Acids)
17 Amino Acid Mix
- supplies most amino acids needed for protein synthesis. All a.a. except tyrosine, cysteine and tryptophan; (Phe, Leu, Ser, Pro, His, Gln, Arg, Ile, Met, Thr, Asn, Lys, Arg, Val, Ala, Asp, Glu, Gly)
Tyrosine
- poor solubility at neutral pH because of its hydrophobic aromatic ring. It’s often prepared in alkaline solution (pH>7) and added separately so it doesn’t precipitate.
Cysteine
- chemically unstable due to reactive thiol (-SH) group and gets oxidized to cystine, especially in oxygenated solutions. Is added separately to avid redox balance in reaction.
Additives
Nicotinamide
- precursor to NAD+, NADH; supports redox balance and metabolic energy regeneration.
Backfill
Nuclease Free Water
- backbone of solution, adjusts final concentration, volume and prevents nucleic acid degradation. Treated to remove DNases, RNases.
2. 1-hour PEP-NTP mix vs 20-hour NMP-ribose-glucose mix
| Mix: | 1-hour PEP-NTP | 20-hour NMP-ribose-glucose |
|---|---|---|
| Materials | ready-to-use NTPs | simpler precursors (NMPs) |
| Act | immediate | longer protein production |
| Energy source | PEP (phosphoenolpyruvate) | Glucose, Ribose |
The 1-hour system directly supplies high-energy molecules (PEP) and fully formed NTPs, enabling rapid but short-lived protein production with minimal metabolic processing. In contrast, the 20-hour system uses cheaper precursors (NMPs, ribose, glucose) and relies on endogenous metabolic pathways in the lysate to regenerate NTPs and ATP, allowing sustained, longer reactions. This makes the latter more cost-effective and stable over time but slower in initial output.
3. How can transcription occur if GMP is not included but Guanine is?
Guanine can be salvaged into GMP in some metabolic pathways that can be underwent in the lysate. Can, also, be phosphorylated to GTP.
C: Planning the Global Experiment | Cell-Free Master Mix Design
1. biophysical or functional property of each protein that affects expression or readout in cell-free systems
a. folding kinetics
b. ph sensitivity
c. oxygen dependence
d. maturation time
e. ATP regeneration (nucleotide source, a.a. incorporation)
f. amino acid supply
1. sfGFP
Famously robust and ultra-fast folding. No need for cellular chaperons. Resistant to high salt concentrations and temperature fluctuations.
2. mRFP1
Reduced photostability and slow maturation of chromophore (=> can delay fluorescence signal), less bright. Because CFS have limited working time, a big part of the protein may remain invisible before energetic resources run dry.
3. mKO2
Bright, maturates fast, sensible to pH.
4. mTurquoise2
High quantum yield (efficiency of converting absorbed light into emitted light), good folding efficiency, photostable, overall stable monomeric structure (=> doesn’t tend to form aggregates)
5. mScarlet_I
Rapid maturation, really bright (=> helps in decreasing red protein-specific readout delay; corelates translation rate to fluorescent signal almost perfectly), also folds efficiently.
6. Electra2
Rapid maturation, not that stable or bright, but tolerant to salts.
Adjusting cell-free mastermix for mTurquoise2
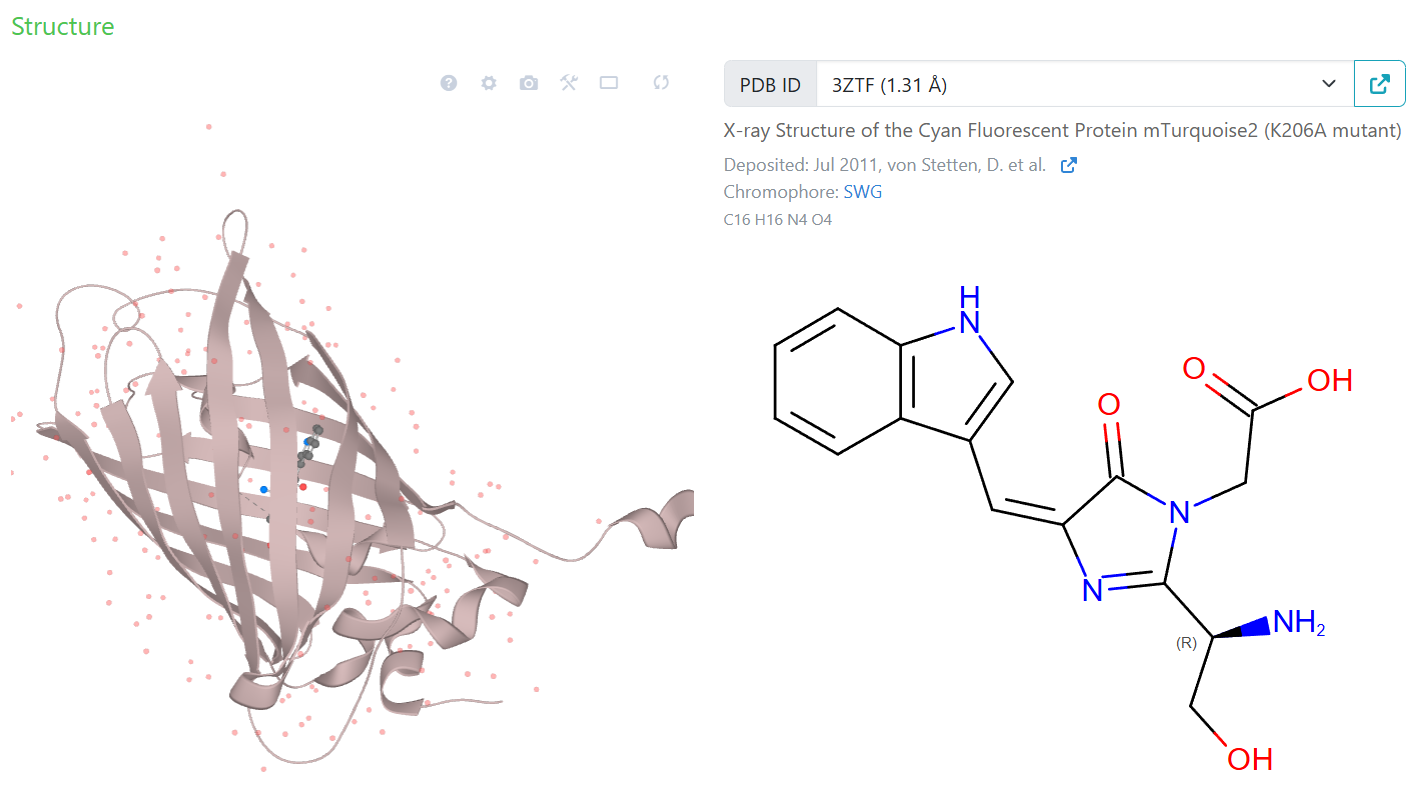

Add oxygen for chromophore formation?
D. Build-A-Cloud-Lab
I played around a bit on simulation tool to try and come up with a design. (I have no idea what I did there)
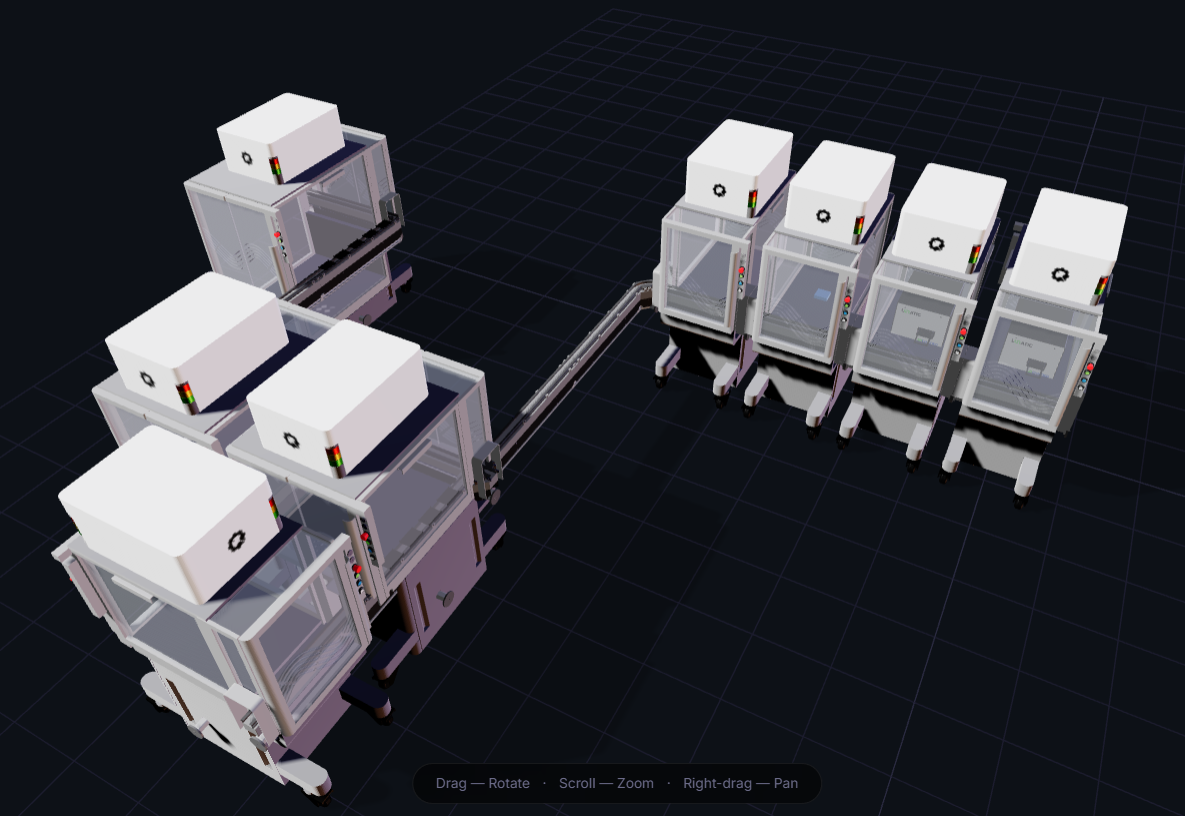


Bibliography
AI tool It was, in fact, really helpful :)