from opentrons import types
metadata = {
'author': 'Casian Veselin',
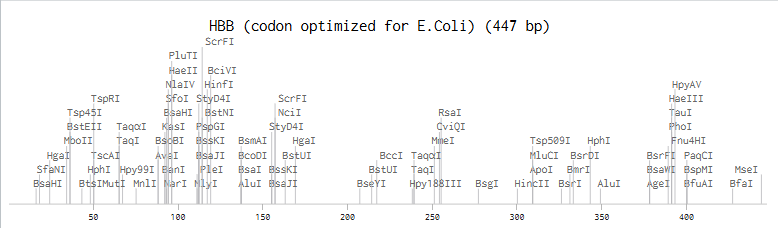
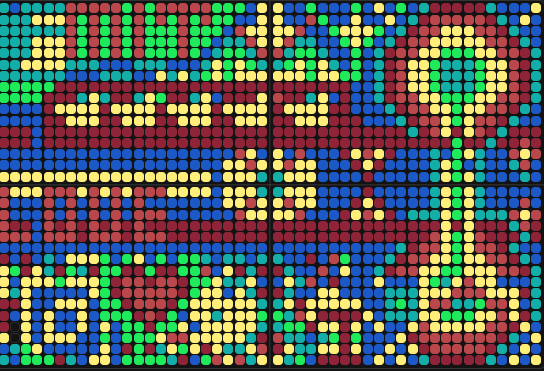
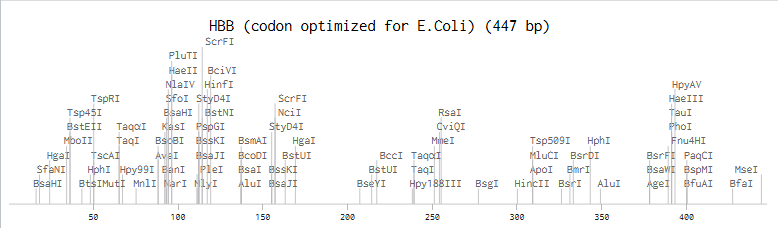
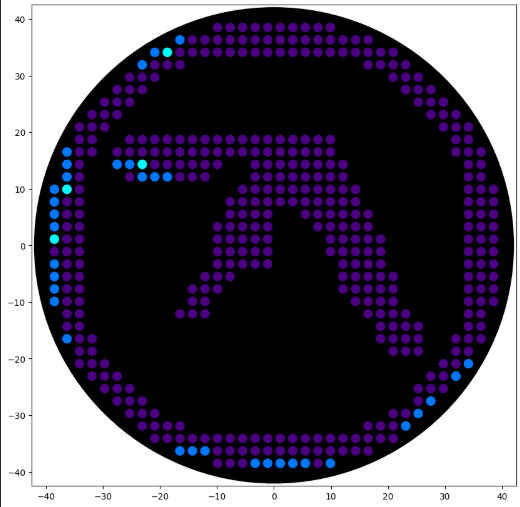
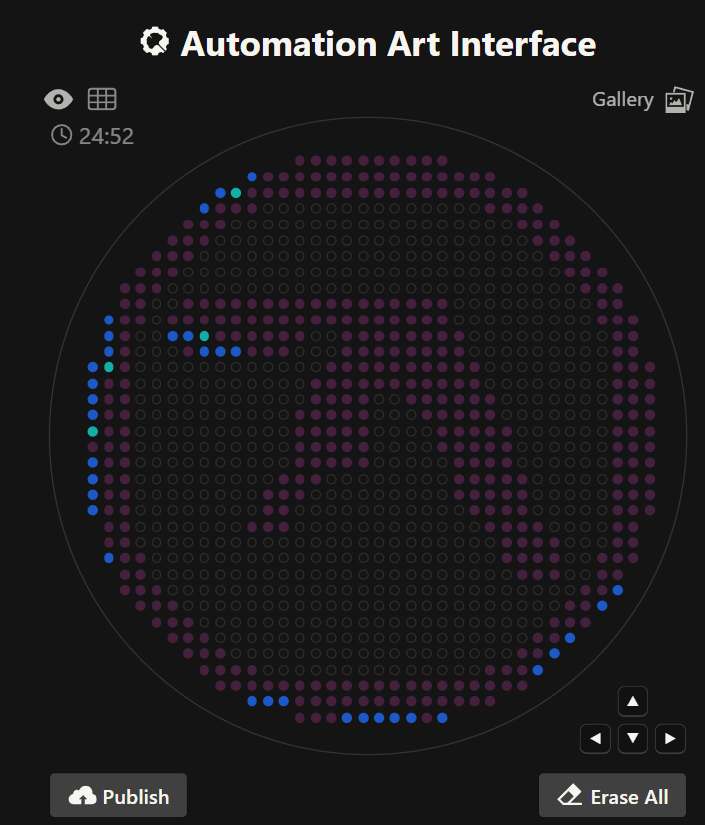
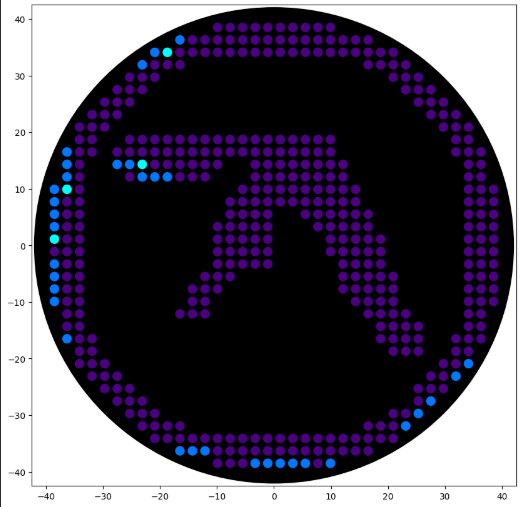
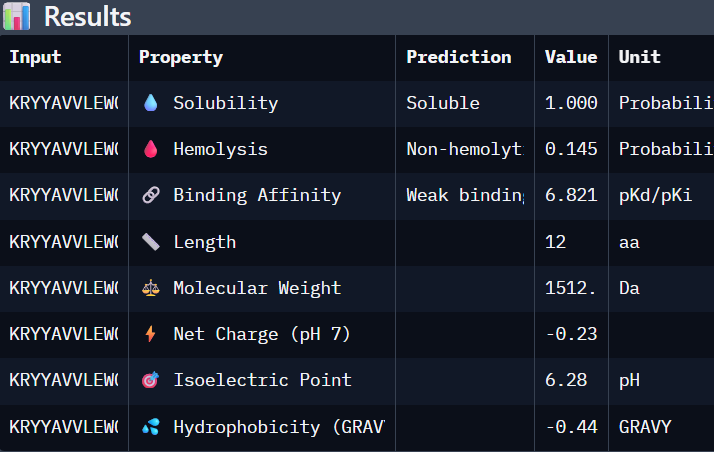
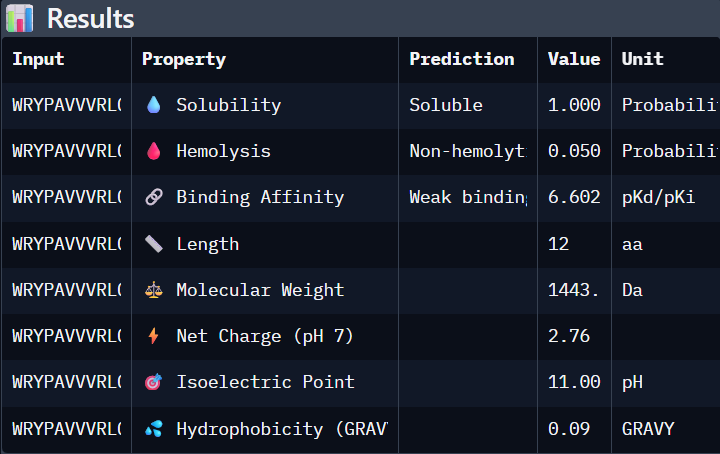
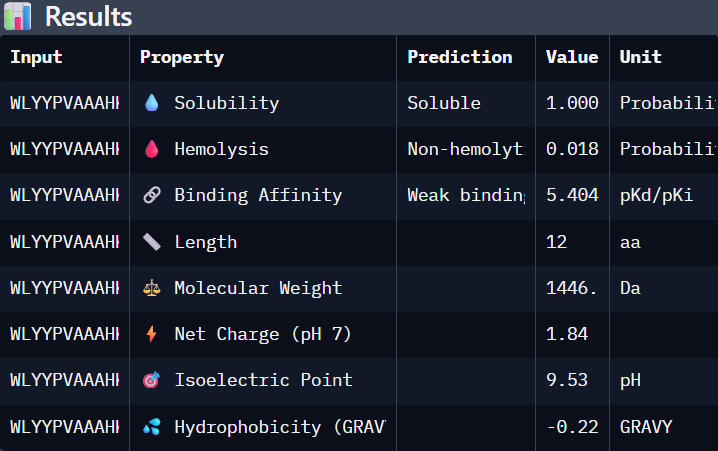
'protocolName': 'Aphex Twin logo ',
'source': 'HTGAA 2026 Opentrons Lab',
'apiLevel': '2.20'
}
# --- Robot Deck Setup ---
TIP_RACK_DECK_SLOT = 9
COLORS_DECK_SLOT = 6
AGAR_DECK_SLOT = 5
PIPETTE_STARTING_TIP_WELL = 'A1'
# Mapping each protein set to a specific well on the 96-aluminum block
# Updated mapping using standard Matplotlib colors
well_colors = {
'A1': '#0077ff', # Electra2 (Blue)
'A2': '#4b0082', # mKate2 (Deep Purple)
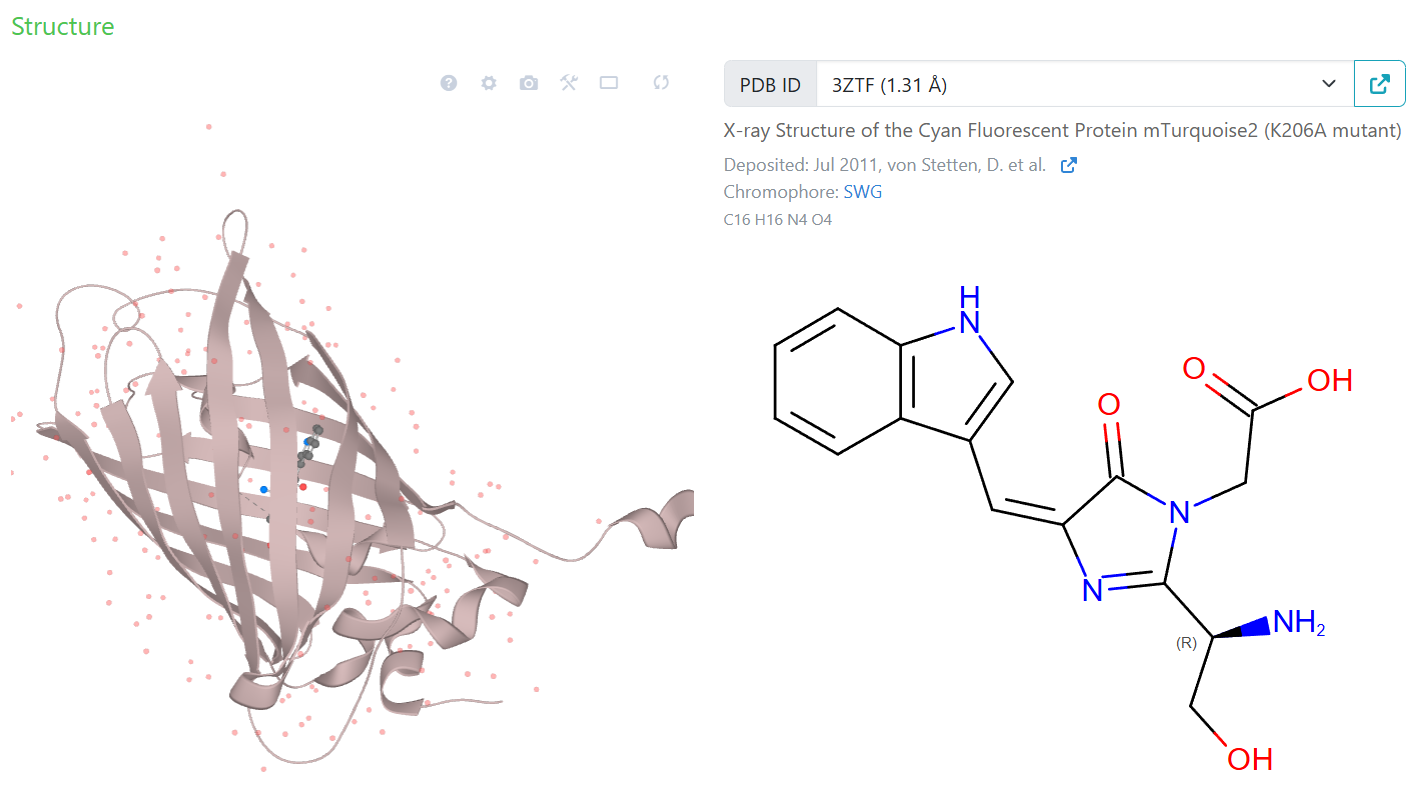
'C1': '#00ffff' # mTurquoise2 (Cyan)
}
def run(protocol):
# --- Load Labware ---
tips_20ul = protocol.load_labware('opentrons_96_tiprack_20ul', TIP_RACK_DECK_SLOT, 'Opentrons 20uL Tips')
pipette_20ul = protocol.load_instrument("p20_single_gen2", "right", [tips_20ul])
temperature_module = protocol.load_module('temperature module gen2', COLORS_DECK_SLOT)
color_plate = temperature_module.load_labware('opentrons_96_aluminumblock_generic_pcr_strip_200ul', 'Cold Plate')
agar_plate = protocol.load_labware('htgaa_agar_plate', AGAR_DECK_SLOT, 'Agar Plate')
center_location = agar_plate['A1'].top()
pipette_20ul.starting_tip = tips_20ul.well(PIPETTE_STARTING_TIP_WELL)
# --- Data ---
# Grouping all point sets into a dictionary for clean iteration
all_designs = {
'#4b0082': [(-9.9, 38.5),(-7.7, 38.5),(-5.5, 38.5),(-3.3, 38.5),(-1.1, 38.5),(1.1, 38.5),(3.3, 38.5),(5.5, 38.5),(7.7, 38.5),(9.9, 38.5),(-14.3, 36.3),(-12.1, 36.3),(-9.9, 36.3),(-7.7, 36.3),(-5.5, 36.3),(-3.3, 36.3),(-1.1, 36.3),(1.1, 36.3),(3.3, 36.3),(5.5, 36.3),(7.7, 36.3),(9.9, 36.3),(12.1, 36.3),(14.3, 36.3),(16.5, 36.3),(-16.5, 34.1),(-14.3, 34.1),(-12.1, 34.1),(-9.9, 34.1),(-7.7, 34.1),(-5.5, 34.1),(-3.3, 34.1),(-1.1, 34.1),(1.1, 34.1),(3.3, 34.1),(5.5, 34.1),(7.7, 34.1),(9.9, 34.1),(12.1, 34.1),(14.3, 34.1),(16.5, 34.1),(18.7, 34.1),(20.9, 34.1),(-20.9, 31.9),(-18.7, 31.9),(-16.5, 31.9),(16.5, 31.9),(18.7, 31.9),(20.9, 31.9),(23.1, 31.9),(-25.3, 29.7),(-23.1, 29.7),(-20.9, 29.7),(20.9, 29.7),(23.1, 29.7),(25.3, 29.7),(-27.5, 27.5),(-25.3, 27.5),(-23.1, 27.5),(23.1, 27.5),(25.3, 27.5),(27.5, 27.5),(-29.7, 25.3),(-27.5, 25.3),(-25.3, 25.3),(25.3, 25.3),(27.5, 25.3),(29.7, 25.3),(-31.9, 23.1),(-29.7, 23.1),(-27.5, 23.1),(27.5, 23.1),(29.7, 23.1),(31.9, 23.1),(-34.1, 20.9),(-31.9, 20.9),(-29.7, 20.9),(29.7, 20.9),(31.9, 20.9),(34.1, 20.9),(-34.1, 18.7),(-31.9, 18.7),(-25.3, 18.7),(-23.1, 18.7),(-20.9, 18.7),(-18.7, 18.7),(-16.5, 18.7),(-14.3, 18.7),(-12.1, 18.7),(-9.9, 18.7),(-7.7, 18.7),(-5.5, 18.7),(-3.3, 18.7),(-1.1, 18.7),(1.1, 18.7),(3.3, 18.7),(5.5, 18.7),(7.7, 18.7),(9.9, 18.7),(31.9, 18.7),(34.1, 18.7),(-34.1, 16.5),(-31.9, 16.5),(-27.5, 16.5),(-25.3, 16.5),(-23.1, 16.5),(-20.9, 16.5),(-18.7, 16.5),(-16.5, 16.5),(-14.3, 16.5),(-12.1, 16.5),(-9.9, 16.5),(-7.7, 16.5),(-5.5, 16.5),(-3.3, 16.5),(-1.1, 16.5),(1.1, 16.5),(3.3, 16.5),(5.5, 16.5),(7.7, 16.5),(9.9, 16.5),(31.9, 16.5),(34.1, 16.5),(36.3, 16.5),(-34.1, 14.3),(-20.9, 14.3),(-18.7, 14.3),(-16.5, 14.3),(-14.3, 14.3),(-12.1, 14.3),(-9.9, 14.3),(-3.3, 14.3),(-1.1, 14.3),(1.1, 14.3),(3.3, 14.3),(5.5, 14.3),(7.7, 14.3),(9.9, 14.3),(12.1, 14.3),(34.1, 14.3),(36.3, 14.3),(-34.1, 12.1),(-25.3, 12.1),(-16.5, 12.1),(-14.3, 12.1),(-12.1, 12.1),(-3.3, 12.1),(-1.1, 12.1),(1.1, 12.1),(3.3, 12.1),(5.5, 12.1),(7.7, 12.1),(9.9, 12.1),(12.1, 12.1),(34.1, 12.1),(36.3, 12.1),(-34.1, 9.9),(-5.5, 9.9),(-3.3, 9.9),(-1.1, 9.9),(1.1, 9.9),(3.3, 9.9),(5.5, 9.9),(7.7, 9.9),(9.9, 9.9),(12.1, 9.9),(14.3, 9.9),(34.1, 9.9),(36.3, 9.9),(38.5, 9.9),(-36.3, 7.7),(-34.1, 7.7),(-7.7, 7.7),(-5.5, 7.7),(-3.3, 7.7),(-1.1, 7.7),(1.1, 7.7),(3.3, 7.7),(5.5, 7.7),(7.7, 7.7),(9.9, 7.7),(12.1, 7.7),(14.3, 7.7),(34.1, 7.7),(36.3, 7.7),(38.5, 7.7),(-36.3, 5.5),(-34.1, 5.5),(-7.7, 5.5),(-5.5, 5.5),(-3.3, 5.5),(-1.1, 5.5),(5.5, 5.5),(7.7, 5.5),(9.9, 5.5),(12.1, 5.5),(14.3, 5.5),(16.5, 5.5),(34.1, 5.5),(36.3, 5.5),(38.5, 5.5),(-36.3, 3.3),(-34.1, 3.3),(-9.9, 3.3),(-7.7, 3.3),(-5.5, 3.3),(-3.3, 3.3),(-1.1, 3.3),(7.7, 3.3),(9.9, 3.3),(12.1, 3.3),(14.3, 3.3),(16.5, 3.3),(34.1, 3.3),(36.3, 3.3),(38.5, 3.3),(-36.3, 1.1),(-34.1, 1.1),(-9.9, 1.1),(-7.7, 1.1),(-5.5, 1.1),(-3.3, 1.1),(-1.1, 1.1),(9.9, 1.1),(12.1, 1.1),(14.3, 1.1),(16.5, 1.1),(18.7, 1.1),(34.1, 1.1),(36.3, 1.1),(38.5, 1.1),(-38.5, -1.1),(-36.3, -1.1),(-34.1, -1.1),(-9.9, -1.1),(-7.7, -1.1),(-5.5, -1.1),(-3.3, -1.1),(-1.1, -1.1),(9.9, -1.1),(12.1, -1.1),(14.3, -1.1),(16.5, -1.1),(18.7, -1.1),(34.1, -1.1),(36.3, -1.1),(38.5, -1.1),(-36.3, -3.3),(-34.1, -3.3),(-9.9, -3.3),(-7.7, -3.3),(-5.5, -3.3),(-3.3, -3.3),(-1.1, -3.3),(12.1, -3.3),(14.3, -3.3),(16.5, -3.3),(18.7, -3.3),(34.1, -3.3),(36.3, -3.3),(38.5, -3.3),(-36.3, -5.5),(-34.1, -5.5),(-12.1, -5.5),(-9.9, -5.5),(-7.7, -5.5),(12.1, -5.5),(14.3, -5.5),(16.5, -5.5),(18.7, -5.5),(20.9, -5.5),(34.1, -5.5),(36.3, -5.5),(38.5, -5.5),(-36.3, -7.7),(-34.1, -7.7),(-14.3, -7.7),(-12.1, -7.7),(-9.9, -7.7),(12.1, -7.7),(14.3, -7.7),(16.5, -7.7),(18.7, -7.7),(20.9, -7.7),(34.1, -7.7),(36.3, -7.7),(38.5, -7.7),(-36.3, -9.9),(-34.1, -9.9),(-14.3, -9.9),(-12.1, -9.9),(14.3, -9.9),(16.5, -9.9),(18.7, -9.9),(20.9, -9.9),(34.1, -9.9),(36.3, -9.9),(38.5, -9.9),(-36.3, -12.1),(-34.1, -12.1),(-16.5, -12.1),(-14.3, -12.1),(-12.1, -12.1),(16.5, -12.1),(18.7, -12.1),(20.9, -12.1),(23.1, -12.1),(34.1, -12.1),(36.3, -12.1),(-36.3, -14.3),(-34.1, -14.3),(18.7, -14.3),(20.9, -14.3),(23.1, -14.3),(25.3, -14.3),(34.1, -14.3),(36.3, -14.3),(-34.1, -16.5),(-31.9, -16.5),(18.7, -16.5),(20.9, -16.5),(23.1, -16.5),(25.3, -16.5),(31.9, -16.5),(34.1, -16.5),(36.3, -16.5),(-34.1, -18.7),(-31.9, -18.7),(20.9, -18.7),(23.1, -18.7),(25.3, -18.7),(31.9, -18.7),(34.1, -18.7),(-34.1, -20.9),(-31.9, -20.9),(-29.7, -20.9),(29.7, -20.9),(31.9, -20.9),(-31.9, -23.1),(-29.7, -23.1),(-27.5, -23.1),(27.5, -23.1),(29.7, -23.1),(-29.7, -25.3),(-27.5, -25.3),(-25.3, -25.3),(25.3, -25.3),(27.5, -25.3),(29.7, -25.3),(-27.5, -27.5),(-25.3, -27.5),(-23.1, -27.5),(23.1, 27.5),(25.3, -27.5),(-25.3, -29.7),(-23.1, -29.7),(-20.9, -29.7),(20.9, -29.7),(23.1, -29.7),(-23.1, -31.9),(-20.9, -31.9),(-18.7, -31.9),(-16.5, -31.9),(16.5, -31.9),(18.7, -31.9),(20.9, -31.9),(-20.9, -34.1),(-18.7, -34.1),(-16.5, -34.1),(-14.3, -34.1),(-12.1, -34.1),(-9.9, -34.1),(-7.7, -34.1),(-5.5, -34.1),(-3.3, -34.1),(-1.1, -34.1),(1.1, -34.1),(3.3, -34.1),(5.5, -34.1),(7.7, -34.1),(9.9, -34.1),(12.1, -34.1),(14.3, -34.1),(16.5, -34.1),(18.7, -34.1),(20.9, -34.1),(-9.9, -36.3),(-7.7, -36.3),(-5.5, -36.3),(-3.3, -36.3),(-1.1, -36.3),(1.1, -36.3),(3.3, -36.3),(5.5, -36.3),(7.7, -36.3),(9.9, -36.3),(12.1, -36.3),(14.3, -36.3),(16.5, -36.3),(-9.9, -38.5),(-7.7, -38.5),(-5.5, -38.5),(7.7, -38.5)],
'#0077ff': [(-16.5, 36.3),(-20.9, 34.1),(-23.1, 31.9),(-36.3, 16.5),(-36.3, 14.3),(-27.5, 14.3),(-25.3, 14.3),(-36.3, 12.1),(-23.1, 12.1),(-20.9, 12.1),(-18.7, 12.1),(-38.5, 9.9),(-38.5, 7.7),(-38.5, 5.5),(-38.5, 3.3),(-38.5, -3.3),(-38.5, -5.5),(-38.5, -7.7),(-38.5, -9.9),(-36.3, -16.5),(34.1, -20.9),(31.9, -23.1),(27.5, -27.5),(25.3, -29.7),(23.1, -31.9),(-16.5, -36.3),(-14.3, -36.3),(-12.1, -36.3),(-3.3, -38.5),(-1.1, -38.5),(1.1, -38.5),(3.3, -38.5),(5.5, -38.5),(9.9, -38.5)],
'#00ffff': [(-18.7, 34.1),(-23.1, 14.3),(-36.3, 9.9),(-38.5, 1.1)]
}
# --- Helper Functions ---
def location_of_color(color_name):
for well, protein in well_colors.items():
if protein.lower() == color_name.lower():
return color_plate[well]
raise ValueError(f"No well found for {color_name}")
def dispense_and_detach(pipette, volume, location):
assert(isinstance(volume, (int, float)))
above_location = location.move(types.Point(z=location.point.z + 5))
pipette.move_to(above_location)
pipette.dispense(volume, location)
pipette.move_to(above_location)
# --- Main Execution ---
for protein_name, points in all_designs.items():
if not points:
continue
pipette_20ul.pick_up_tip()
for i, (x, y) in enumerate(points):
# Aspirate every 20 drops (full pipette capacity)
if i % 20 == 0:
volume_needed = min(20, len(points) - i)
pipette_20ul.aspirate(volume_needed, location_of_color(protein_name))
target_location = center_location.move(types.Point(x=x, y=y))
dispense_and_detach(pipette_20ul, 1, target_location)
# Drop tip after completing one protein color to prevent mixing
pipette_20ul.drop_tip()
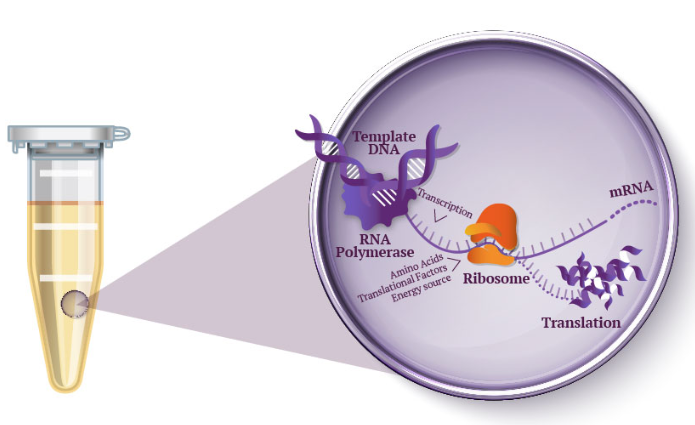
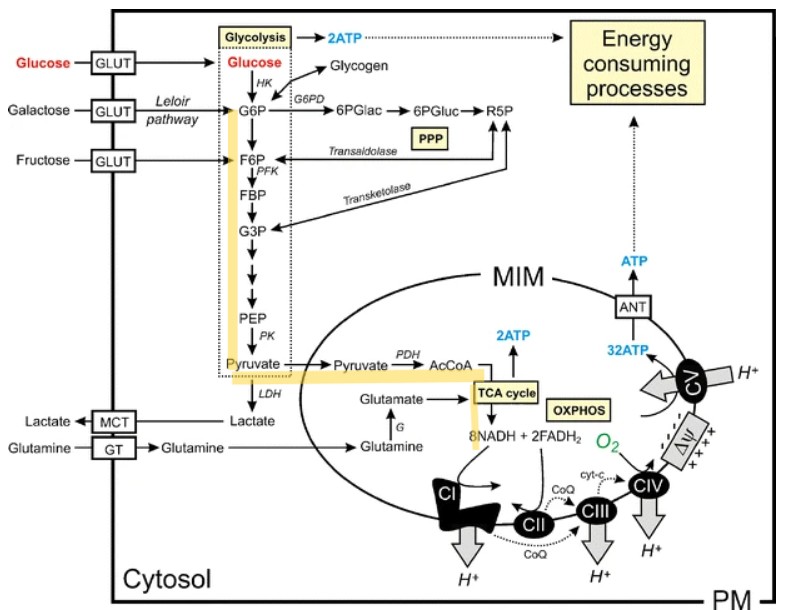
This paper explores enhancing lipid production in plant cells through automated, high-throughput genome engineering and phenotyping, focusing on maize (Zea mays) and Nicotiana benthamiana. Rather than industrial manufacturing, the research develops a conceptual framework for accelerating biological Design-Build-Test-Learn (DBTL) cycles to improve the yield of lipid macromolecules. The system measures editing efficiency by targeting the photosynthetic gene HCF136; its knockout produces distinct changes in chlorophyll fluorescence intensity, serving as a high-throughput proxy for successful genomic modification and subsequent analysis of lipid metabolic pathways.
The study utilizes the iBioFAB biofoundry, an automated platform integrated with Matrix-Assisted Laser Desorption/Ionization-Mass Spectrometry (MALDI-MS) to expedite genome characterization. These biofoundries operate as specialized informatics-driven workstations that combine robotics with high-throughput instrumentation to scale repetitive protocols. By automating the transition from genome editing to cellular effect characterization, the system reduces human error and significantly optimizes the time required to identify high-yielding lipid variants, transforming traditionally manual plant engineering into a scalable, iterative process.