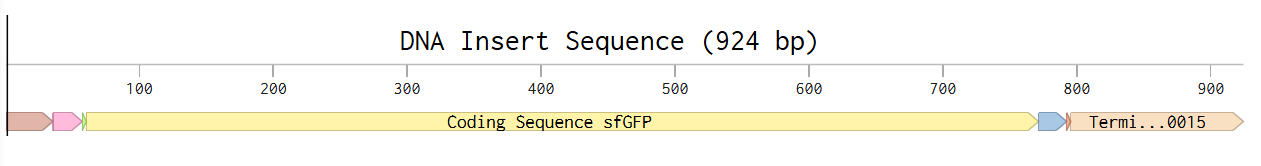
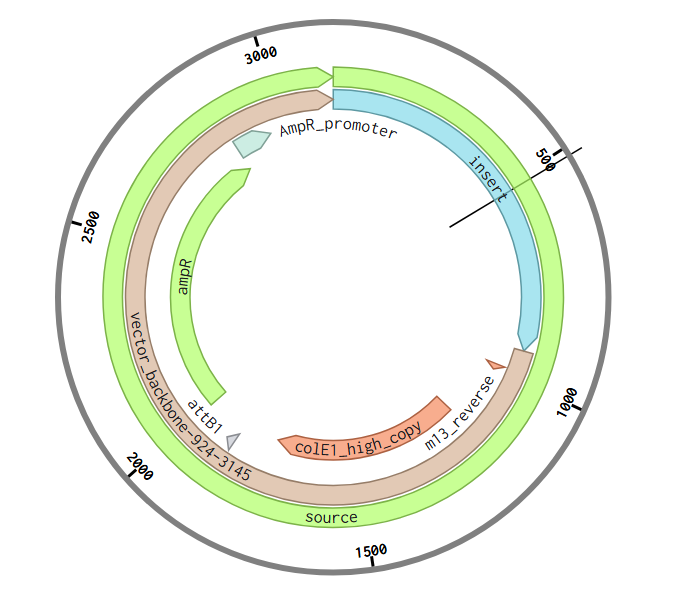
Week 2 HW: DNA Read, Write, and Edit
Part 1: Benchling & In-silico Gel Art
My node (W&M) is using one of our phages’(Kampy) DNA for the Gel Art
- DNA - https://phagesdb.org/phages/Kampy/
- Our lab’s restriction enzymes:
- AfIII
- Age1
- AluI
- ApaI
- BamHI-HF
- BbsI
- BsaI-HF
- BglI
- BstXI
- DpnI
- Eco0109I
- Eco47III
- EcoICRI
- EcoRI-HF
- EcoRV
- HindIII
- I-SceI
- KpnI
- NotI-HF
- PspAI
- PstI-HF
- SpaI-HF
- XbaI
- XhoI
Part 2: Gel Art - Restriction Digests and Gel Electrophoresis
Our design
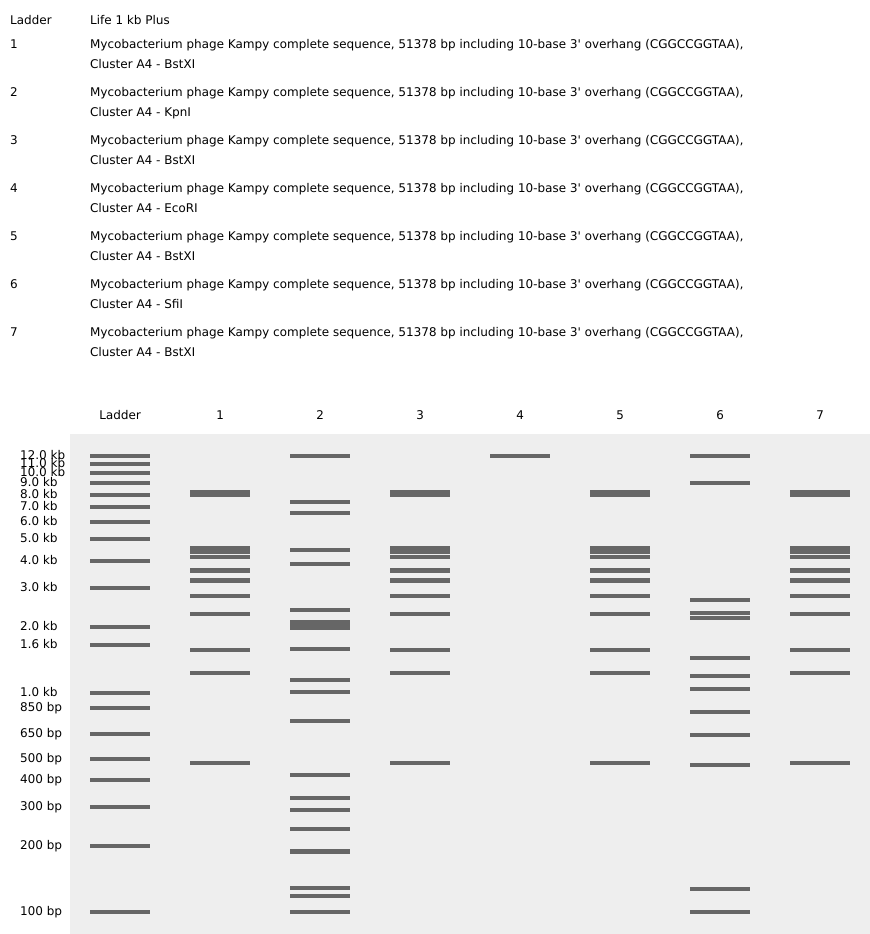


Our gel results
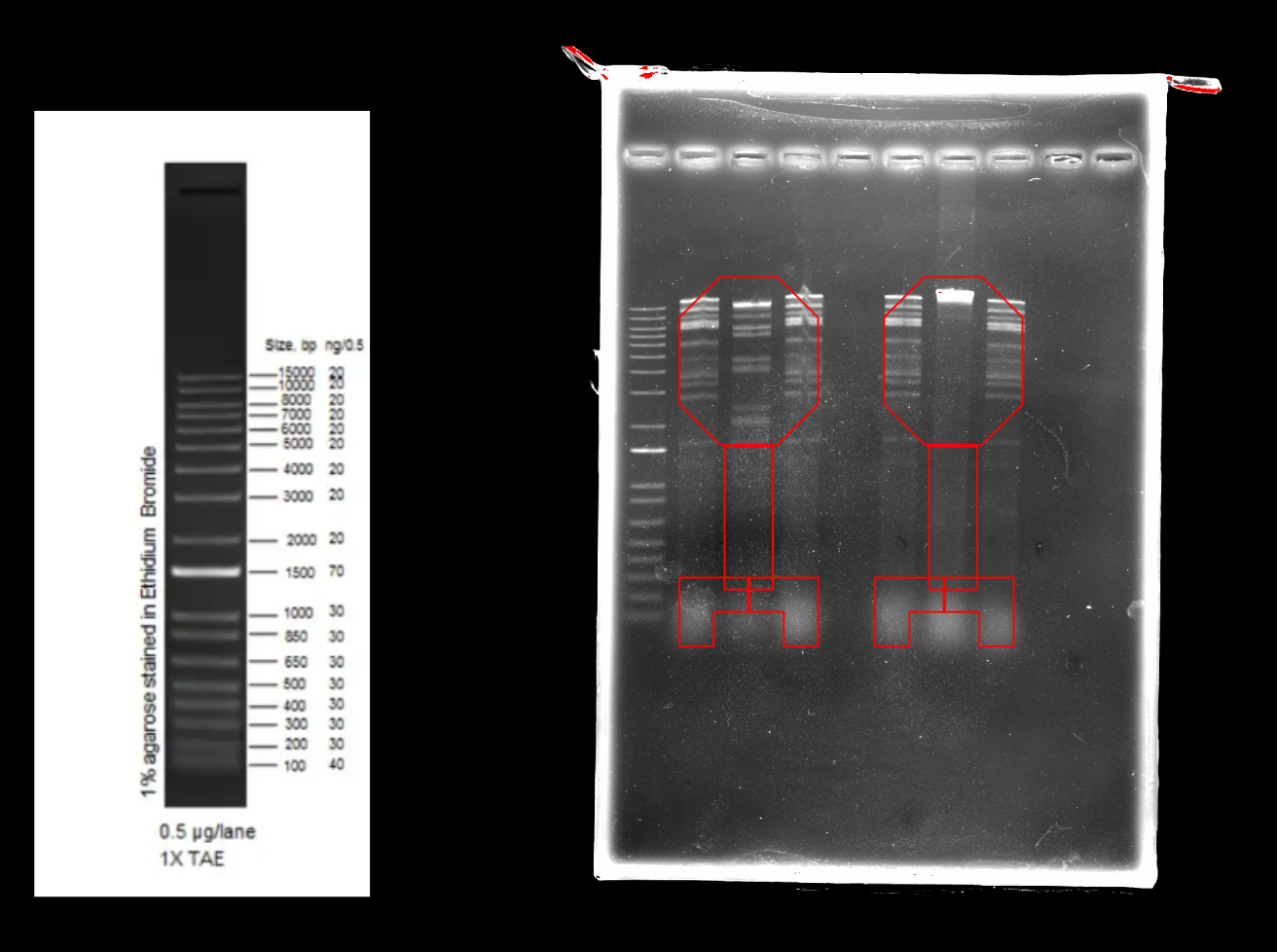

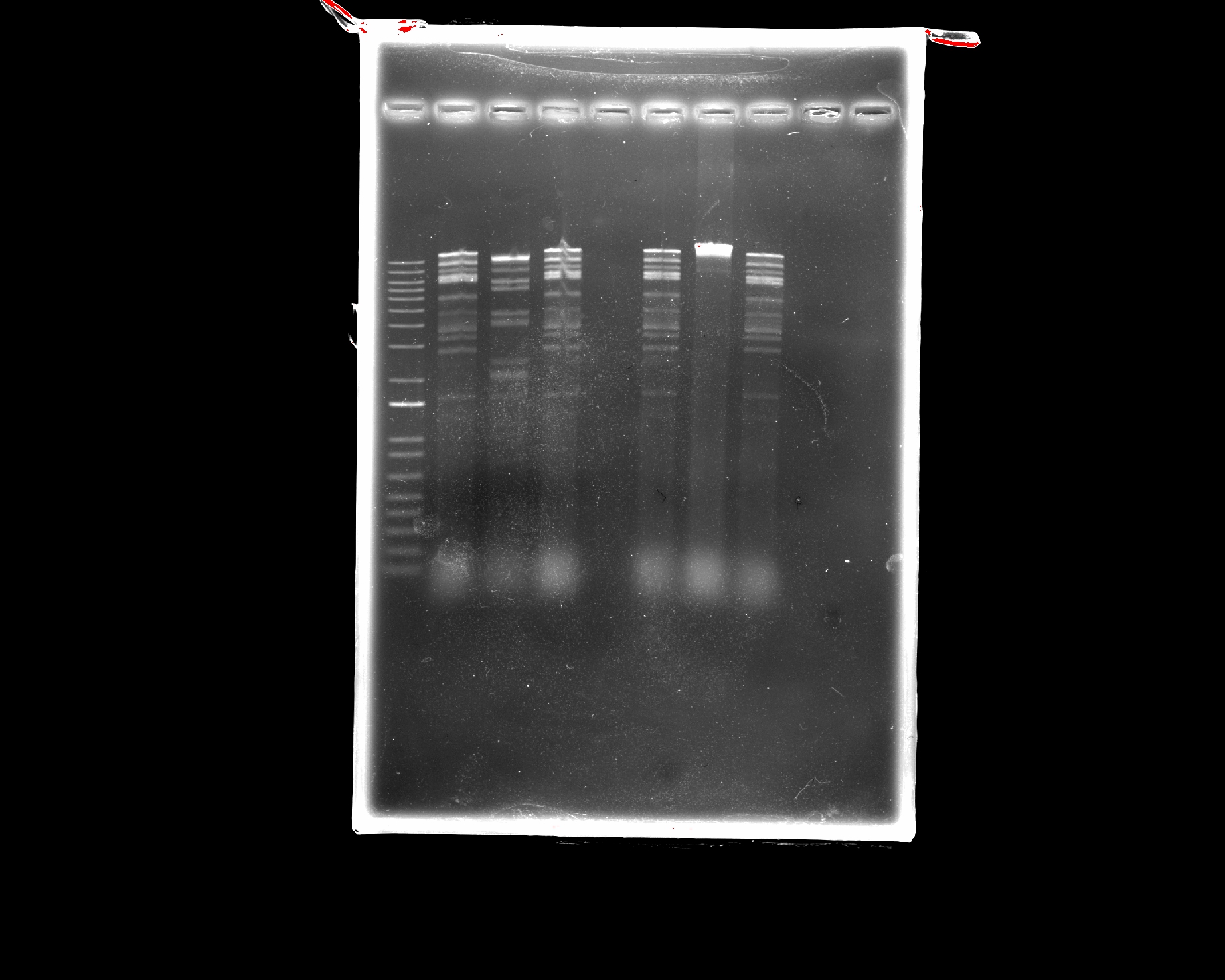

Part 3: DNA Design Challenge
3.1. Choose your protein.
I chose Antirepressor protein ant from Salmonella phage P22 since my lab is interested in antirepressors in bacteriophage and how they induce phage.
>sp|P03037|RANT_BPP22 Antirepressor protein ant OS=Salmonella phage P22 OX=10754 GN=ant PE=1 SV=1 MNSIAILEAVNTSYVPFNGQHVLTAMVAGVAYVAMKPVVDNIGLSWSSQVQKLLKMKDKFNYVDIDMVAGDMKKRLMGCIPLKKLNGWLFSINPEKVRADIRDKLIKYQEECFTVLYDYWTKGKAENPRKKTSVDERTPLRDAVNMLVSKKHLMYPEAYAMIHQRFNVESIEELEASQIPLAVEYIHRVVLEGEFIGKQEKKTNDLSAKEANSLVWLWDYANRSQALFRELYPAMRQIQSNYSGKCYDYGHEFSYIIGIARDVLINHTRDVDINEPDGPTNLSAWMRLKDKELPPSLHRY
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
Ant protein DNA sequence with Codon-Optimization
ATGATGAATTCTATTGCTATTTTAGAAGCTGTTAATACTTCTTATGTTCCTTTTAATGGTCAACATGTTTTAACTGCTATGGTTGCTGGTGTTGCTTATGTTGCTATGAAACCTGTTGTTGATAATATTGGTTTATCTTGGTCTTCTCAAGTTCAAAAATTATTAAAAATGAAAGATAAATTTAATTATGTTGATATTGATATGGTTGCTGGTGATATGAAAAAACGTTTAATGGGTTGTATTCCTTTAAAAAAATTAAATGGTTGGTTATTTTCTATTAATCCTGAAAAAGTTCGTGCTGATATTCGTGATAAATTAATTAAATATCAAGAAGAATGTTTTACTGTTTTATATGATTATTGGACTAAAGGTAAAGCTGAAAATCCTCGTAAAAAAACTTCTGTTGATGAACGTACTCCTTTACGTGATGCTGTTAATATGTTAGTTTCTAAAAAACATTTAATGTATCCTGAAGCTTATGCTATGATTCATCAACGTTTTAATGTTGAATCTATTGAAGAATTAGAAGCTTCTCAAATTCCTTTAGCTGTTGAATATATTCATCGTGTTGTTTTAGAAGGTGAATTTATTGGTAAACAAGAAAAAAAAACTAATGATTTATCTGCTAAAGAAGCTAATTCTTTAGTTTGGTTATGGGATTATGCTAATCGTTCTCAAGCTTTATTTCGTGAATTATATCCTGCTATGCGTCAAATTCAATCTAATTATTCTGGTAAATGTTATGATTATGGTCATGAATTTTCTTATATTATTGGTATTGCTCGTGATGTTTTAATTAATCATACTCGTGATGTTGATATTAATGAACCTGATGGTCCTACTAATTTATCTGCTTGGATGCGTTTAAAAGATAAAGAATTACCTCCTTCTTTACATCGTTATTAA
3.3. Codon optimization.
Different organisms have different bias to certain tRNA which makes them quicker as translating certain codons. Therefore to make the most proteins the quickest, you need to optimize for the organism that will be producing the protein (is being engineered to make it). I have optimized for Salmonella typhimurium (LT2) since it is the only option I found in benchling from the Salmonella genus (which the phage infects).
Ant protein DNA sequence with Codon-Optimization
ATGATGAACAGCATCGCGATTTTGGAGGCCGTGAATACGTCGTATGTCCCGTTTAATGGCCAGCATGTCCTGACCGCAATGGTAGCGGGCGTCGCATATGTGGCGATGAAGCCGGTTGTTGATAATATCGGTCTTAGTTGGTCGTCCCAGGTCCAAAAACTTCTGAAAATGAAAGATAAGTTCAACTATGTTGATATCGACATGGTTGCGGGCGATATGAAAAAAAGACTGATGGGCTGCATTCCGCTGAAGAAATTGAACGGGTGGCTCTTCTCCATAAATCCCGAAAAAGTACGAGCGGATATTCGTGACAAGCTGATCAAATATCAGGAAGAGTGCTTTACAGTACTTTACGACTATTGGACGAAAGGAAAAGCCGAGAACCCGCGTAAAAAAACGTCTGTGGACGAACGGACCCCGTTACGCGATGCGGTTAACATGCTCGTGAGCAAAAAACACCTGATGTACCCGGAAGCTTATGCTATGATCCATCAGCGCTTTAACGTGGAATCAATCGAGGAACTGGAAGCCTCGCAAATTCCATTAGCCGTCGAATACATTCACCGCGTGGTGCTCGAAGGTGAGTTTATTGGCAAACAGGAAAAGAAGACCAATGATTTGTCCGCAAAAGAGGCCAACAGCCTGGTGTGGCTATGGGACTACGCCAATCGCAGCCAGGCTCTGTTTCGTGAACTGTACCCGGCGATGCGTCAGATTCAAAGCAATTATAGCGGAAAATGTTATGATTATGGCCATGAATTCTCTTATATCATTGGGATCGCGCGTGACGTATTAATTAATCATACCCGCGATGTCGATATAAACGAACCTGACGGTCCAACTAACCTGAGTGCGTGGATGCGGCTGAAAGATAAAGAGCTGCCTCCCTCACTGCACCGCTACTAA
3.4. You have a sequence! Now what?
The DNA can be transcribed and translated into the protein by inserting it into a plasmid which is put into the host (bacteria) through methods like chemical transformation or electroporation (which interupt the stability of the cell membrane), and then the host starts expressing it like it would for one of its own proteins.
Part 4: Prepare a Twist DNA Synthesis Order
Part 5: DNA Read/Write/Edit
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why?
I am interested in sequencing bacteriophage and environmental samples. Bacteriophage are becoming a possibility for drug resistant bacteria treatments, and they can be a vector in bio engineering. Their prevalence in the environment and interactions with bacteria also make it interesting to study metagenomic samples with and without phage.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?\
In the past, we have used illumina (NGS) and plasmidsaurus (nanopore) for phage. Plasmidsaurus is quicker and less expensive but has had variable results. Metagenomics also uses NGS and sometimes nanopore (can be more portable for lab work).
- NGS is second-gen, nanopore is third-gen.
- Input is the library which is fragmented extracted DNA which has gone through end repair, a-tailing, and adapter ligation, also possibly pcr if the target sequence needs to be amplified.
- Illumina works by isothermically amplifying the fragments, fragment strands binding to oligos on one end, polymerase creating the reverse strands of the fragments, and then bridge amplification repeating over and over for all the fragments. The reverse strands are discarded and then the forward strands are sequenced through fluorescently tagged nucleotides being added one by one, with the signals being read for each added base and associated with the length.
- The output of illumina is millions or more of short reads. The output for plasmidsaurus is long reads or whole plasmid.
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why?
I think that DNA encoded sensors are interesting. With a background in computing, I can see a lot of places where similar principals would apply and biological sensors could be useful (e.g. water testing, health testing, soil testing, working with hardware to give timelines, etc.).
(ii) What technology or technologies would you use to perform this DNA synthesis and why?\
Enzymatic DNA Synthesis would be good to use since it is more sustainable (and better for longer single strands), but it would be likely easier to start with phosphoramidite method synthesis which is industry standard and more cost-effective (for now).
- The essential steps of the phosphoramidite method for oligonucleotide synthesis are
- Deprotection (acid removes protecting group to expose hydroxyl group)
- Coupling (nucleoside phosphoramidite is activated and added to hydroxyl group)
- Capping (unreacted hydroxyl groups are “capped” to prevent further nucleotide addition)
- Oxidation/Sulfurization (phosphite triester linkage is converted into stable pentavalent phosphotriester bond)
- Limitations of the phosphoramidite method include length (only 100-200bp possible) and having way less sustainability (toxic organic waste is generated).
5.3 DNA Edit
(i) What DNA would you want to edit and why?
I think more can be done for flora, especially in the accessibility of parts (such as seeds) with edits. I think that in human medicine, preventatives and sensors are interesting. My aunt was diagnosed with stage 4 liver cancer in December with no prior warning. It would be great to have better ways to detect and understand silent conditions.
(ii) What technology or technologies would you use to perform these DNA edits and why?\
CRISPR Cas12a can be useful for biosensors, and bacterium mediated transfer and engineered phage (through plasmid transformation) are of interest for targeting plants.
- CRISPR Cas12a processes its own precursor (unlike cas9) into crRNA which combines with the enzyme to form ribonucleoprotein (RNP) complexes. The complex recognizes and binds to a specific Protospacer Adjacent Motif (PAM) site, then breaks both strands (forms sticky ends) and starts breaking down ssDNA close by.
- Preperation includes selecting the target sequence downstream of a PAM and designing the crRNA.
- Limitations include requiring a specific PAM and lower efficiency than cas9 in some mammalian cells