Week 3 HW: Lab Automation
Python Script for Opentrons Artwork
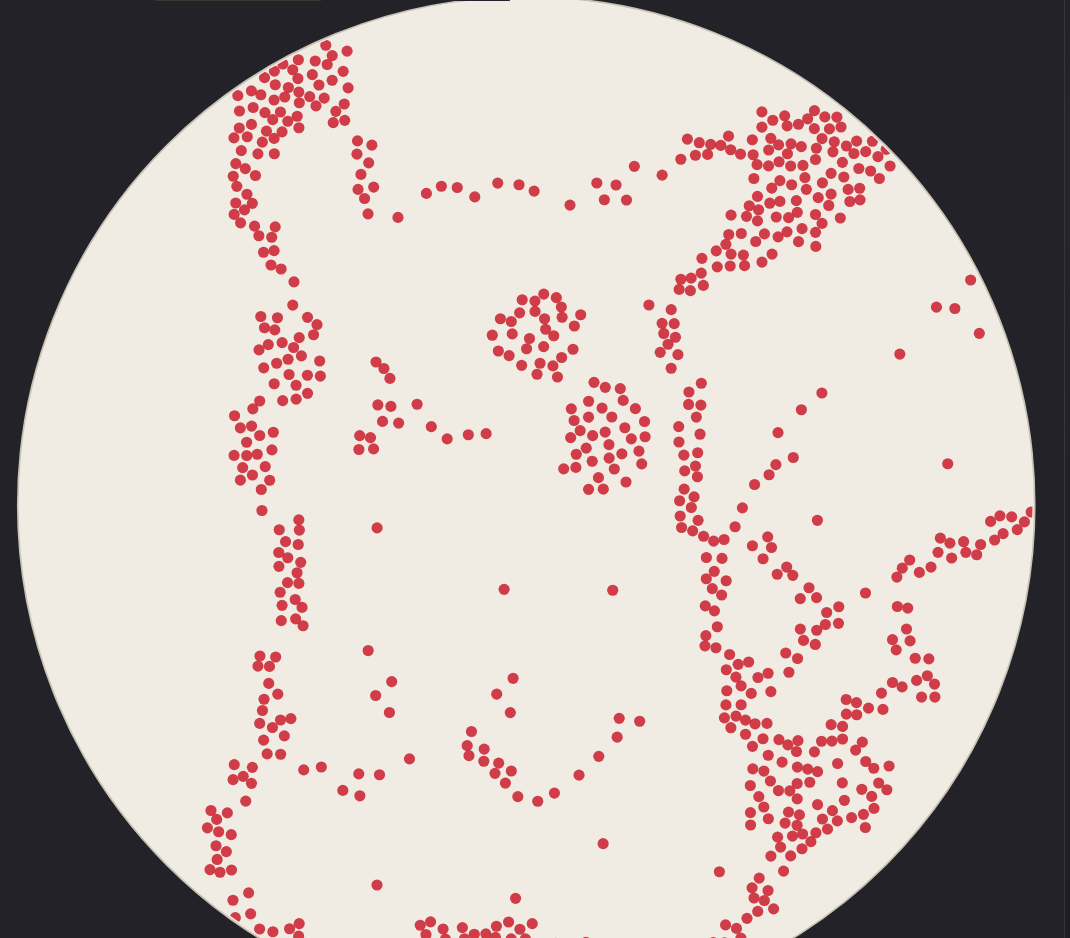
- Designs
Using the GUI at opentrons-art.rcdonovan.com
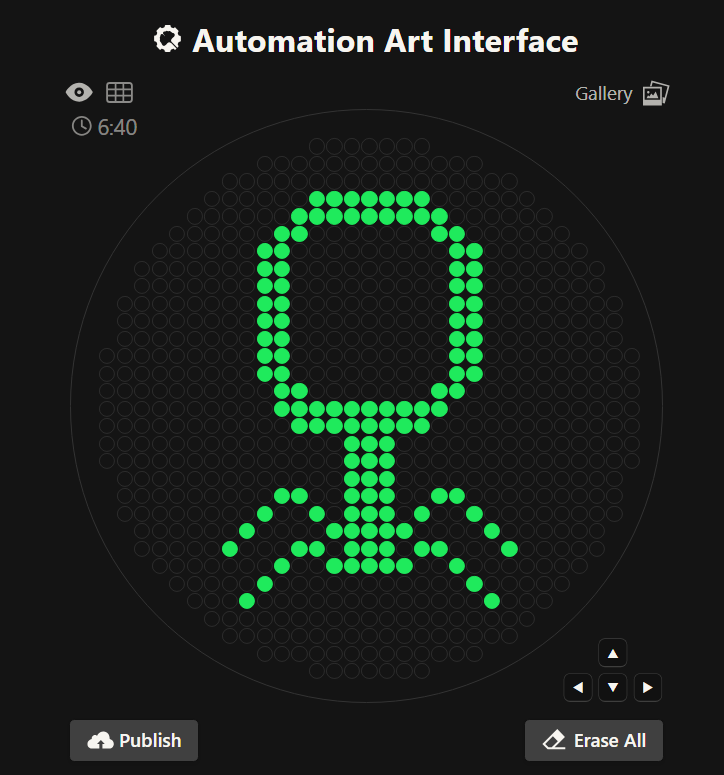

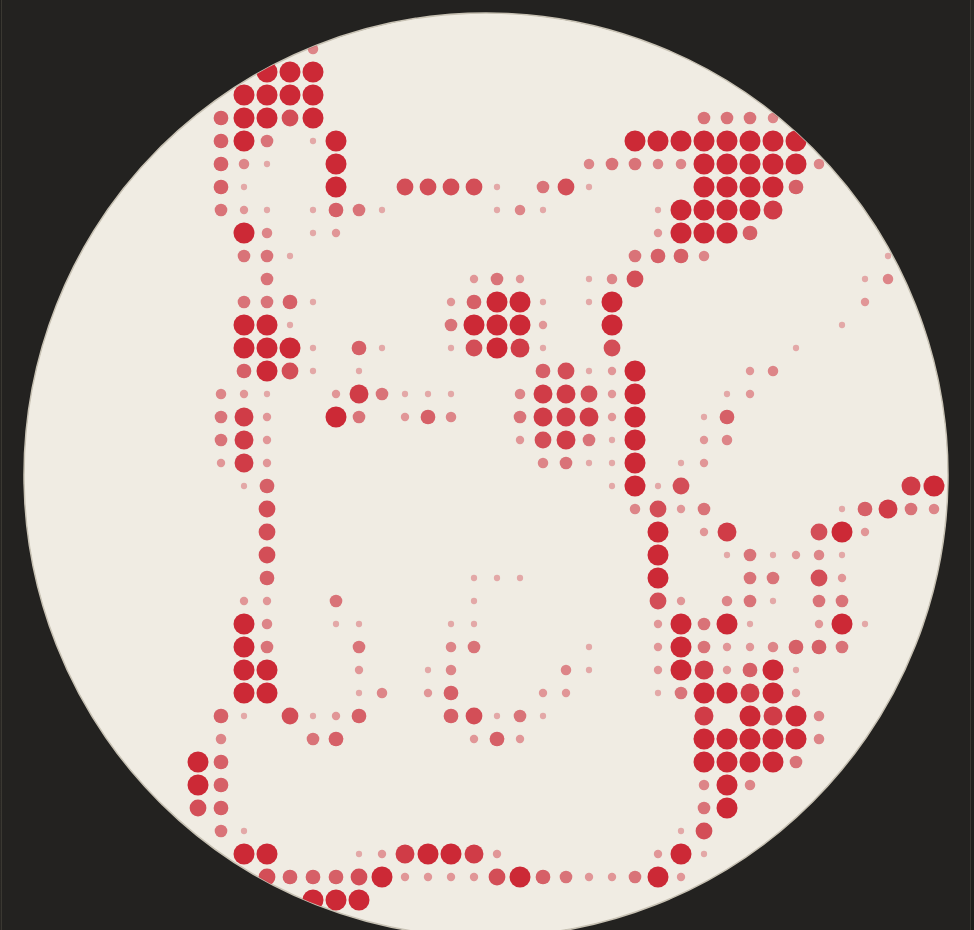
Using custom design tools (trying out stippling, different dot sizes)
- Python Files
What the different dot design looks like after programming so far (still need to scale a bit)
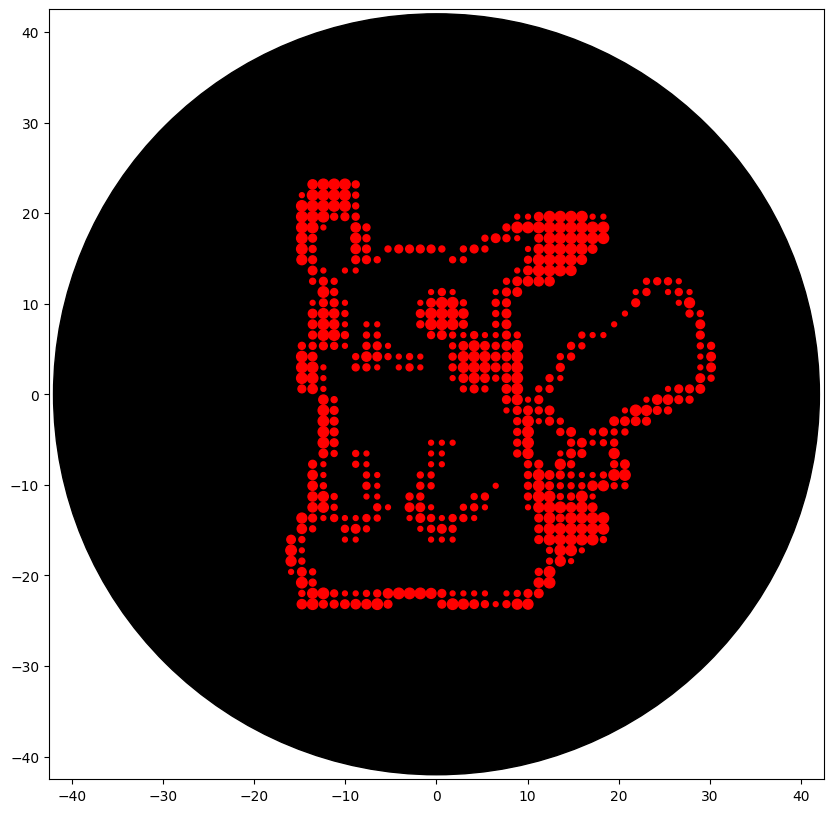

- AI Usage documentation
I utilized Claude Opus 4.6 to help program a web app to assign values (light-dark as 0-8) to pixels of an image so that it could be represented in different dot sizes or clustering on a petri dish. I also used the model to help scale my programming and cordinates to the proper plate size. Claude models work well for general programming related tasks, but sometimes struggle with nailing down details (e.g. it worked out the logic for pixel calculations very well, but struggled to fix the display attributes and relative sizing)
Post-Lab Questions
- Paper Discription
Paper found here: High-Throughput Bacteriophage Testing with Potency Determination: Validation of an Automated Pipetting and Phage Drop-Off Method
The paper by Dufour et al. presents and evaluates using liquid handling robots for high-throughput phage susceptibility testing. This is important since as bacteriophage are used more for phage therapy and other applications, single phage and cocktail testing is used to determine the best phage for the particular target, but can be very time consuming to test manually. The method with the liquid handling robot was found to have lower variance and very similar mean results to the manual assays.
- Final Project Lab Automation
For my final independent project I would like to use the Opentron T2 to run a large range of phage susceptibility tests, similar to my choosen paper. For my more computational project ideas this could be used to validate phage-host range results or possibly be done in the future with the edited phage designs, and for my phage sensor idea this would be done with the transformed bacteria to gather the luminance-titer results.
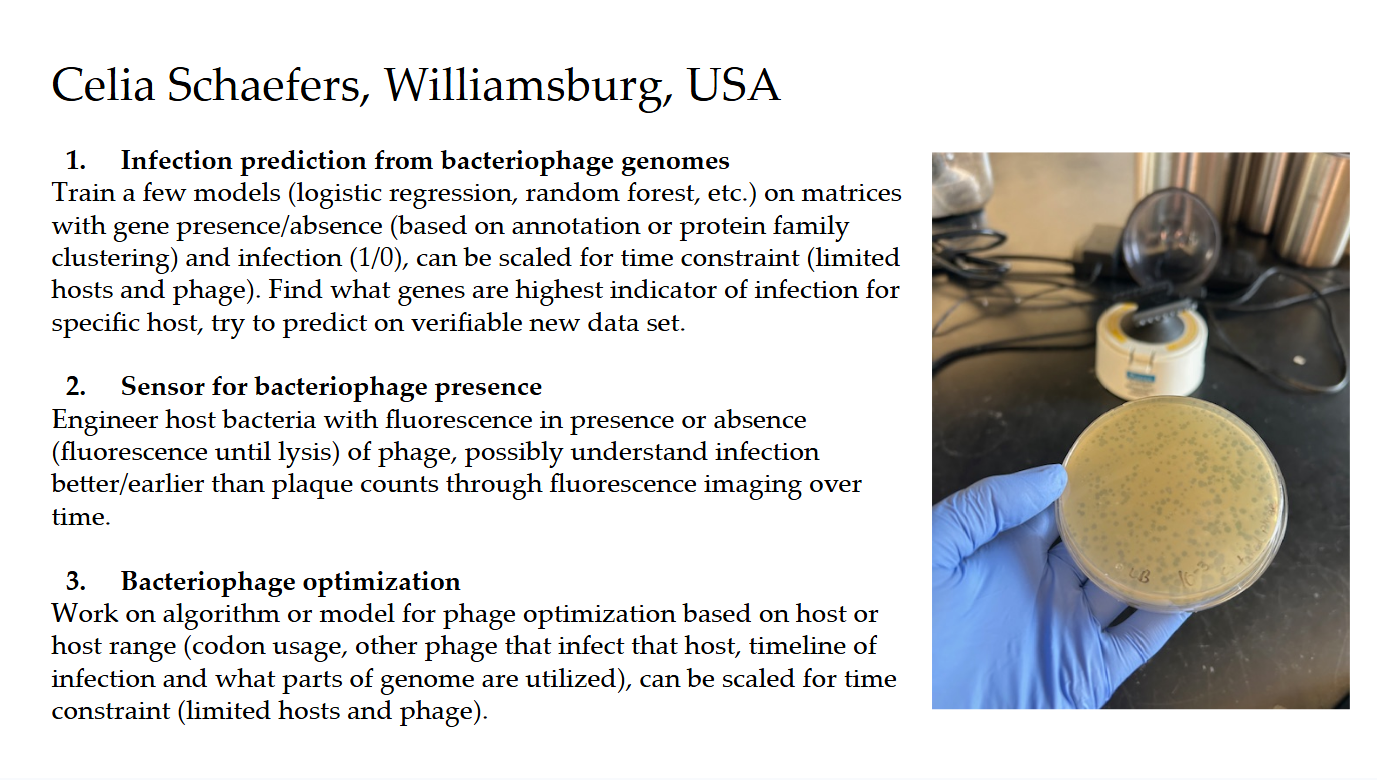
Final Project Ideas



