Week 4 HW: Protein Design Part I
Part A. Conceptual Questions
- How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons)
Meat is about 25% protein so 500 grams of meat is about 125 grams of protein. 100 Daltons is equivalent to 100 grams/mol so it is about 1.25 moles amino acids. Multiplying this by avogadro’s number, 6.022x1023, we get about 7.5275x1024 molecules of amino acids for a 500 gram piece of meat.
- Why do humans eat beef but do not become a cow, eat fish but do not become fish?
We digest what we eat, breaking it down into components to use in our cells (such as amino acids to make new proteins).
- Why are there only 20 natural amino acids?
They fufill a range of roles well (polar, charged, neutral, different sizes), they were based on the elements avalible, and the evolutionary outcome was set at some point as everything down the line started using the same basic building blocks.
- Can you make other non-natural amino acids? Design some new amino acids.
Yes, some examples of uses are for tagging proteins or making bio engineering solutions only work in environments where the non-canonical amino-acid is provided.
- Where did amino acids come from before enzymes that make them, and before life started?
They might have come from space on meteorites, or been created through interactions of elements around hydrothermal vents.
- If you make an α-helix using D-amino acids, what handedness (right or left) would you expect?
D-amino acids form a left handed helix (mirror to the normal right handed helix with L-amino acids).
- Why are most molecular helices right-handed?
The right-handed helix is more stable, and there is no steric hindrance between the side chains in that cofirmation.
- Why do β-sheets tend to aggregate?
- What is the driving force for β-sheet aggregation?
Their edges are prone to binding together (due to unsatisfied hydrogen bonds).
- Why do many amyloid diseases form β-sheets?
- Can you use amyloid β-sheets as materials?
Amyloid diseases include Alzheimer’s and Parkinson’s. Protein misfolding causes the structures to form to stablize, making them hard to break apart. Yes, there are bio engineering applications in things like material for bio fabrics.
Part B: Protein Analysis and Visualization
- Briefly describe the protein you selected and why you selected it.
I choose pdb_00002ms2 which is the MS2 coat protein since that is the bacteriophage we are focusing on for the group final project.
- Identify the amino acid sequence of your protein.
- How long is it? What is the most frequent amino acid?
It is only 127 amino acids long and the most frequent amino acid is Alanine.
- How many protein sequence homologs are there for your protein?
There are 12 blast hits, most of which are other coat/capsid proteins.
- Does your protein belong to any protein family?
It belongs to the Leviviricetes capsid protein family.
- Identify the structure page of your protein in RCSB
- When was the structure solved? Is it a good quality structure?
The structure was released in 1995. The resolution is 2.80 Å which is considered good.
- Are there any other molecules in the solved structure apart from protein?
No, the coat protein is the only solved structure.
- Does your protein belong to any structure classification family?
A few homologs are identified under superfamily RNA bacteriophage capsid protein, domain 2BU1 A.
- Open the structure of your protein in any 3D molecule visualization software:
Visualization in ChimeraX:
Visualization in Pymol:
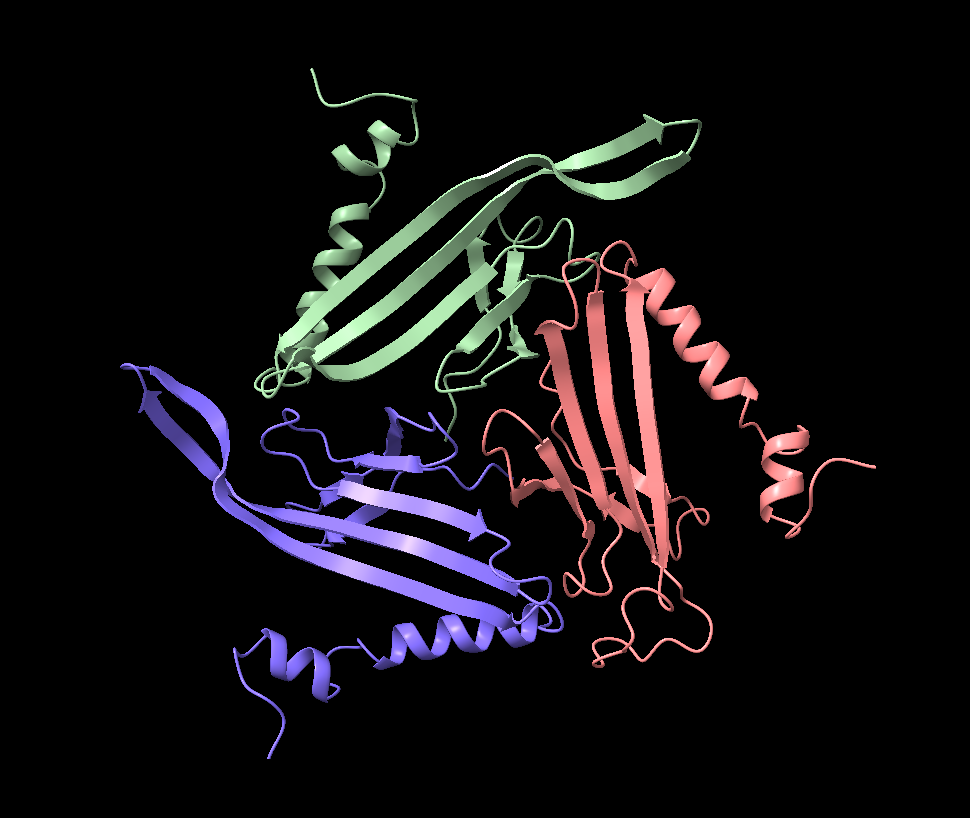
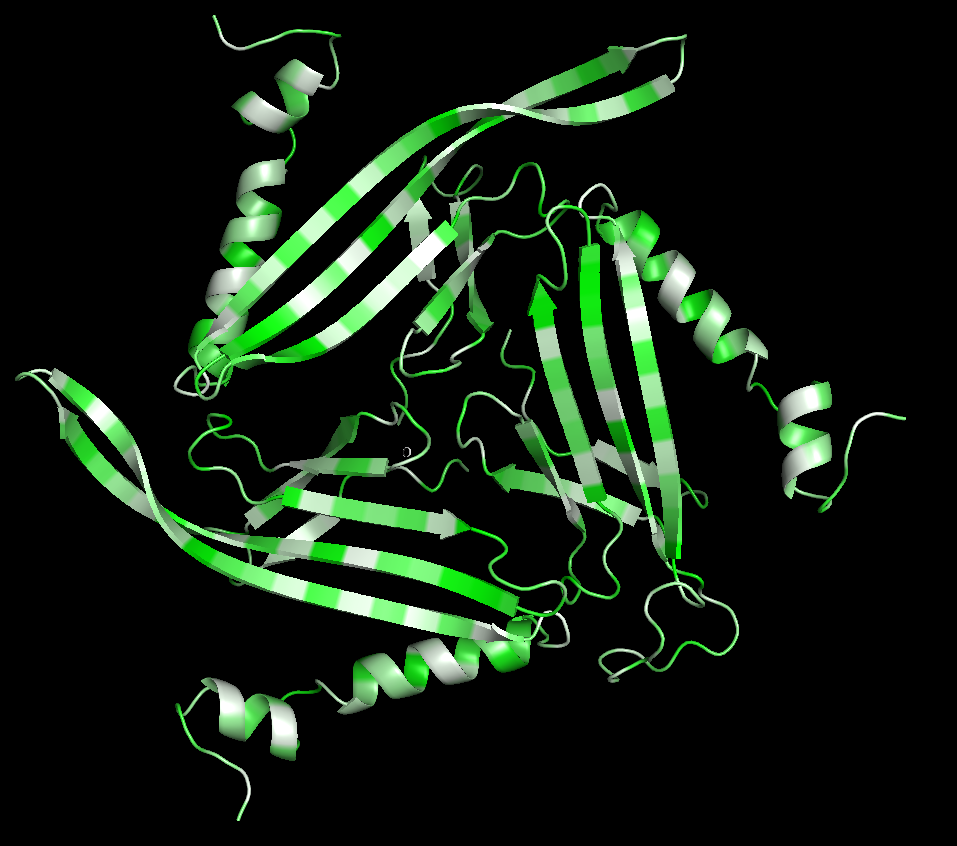
Cartoon
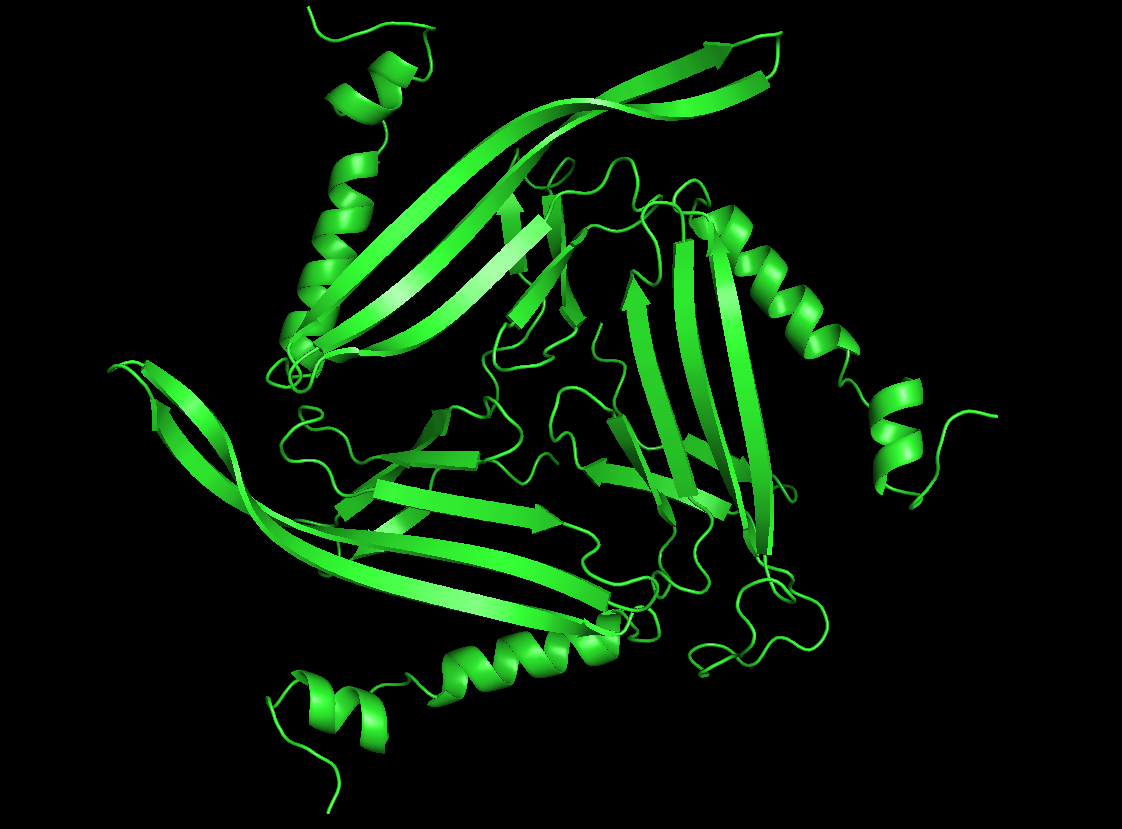

Colored by secondary structure


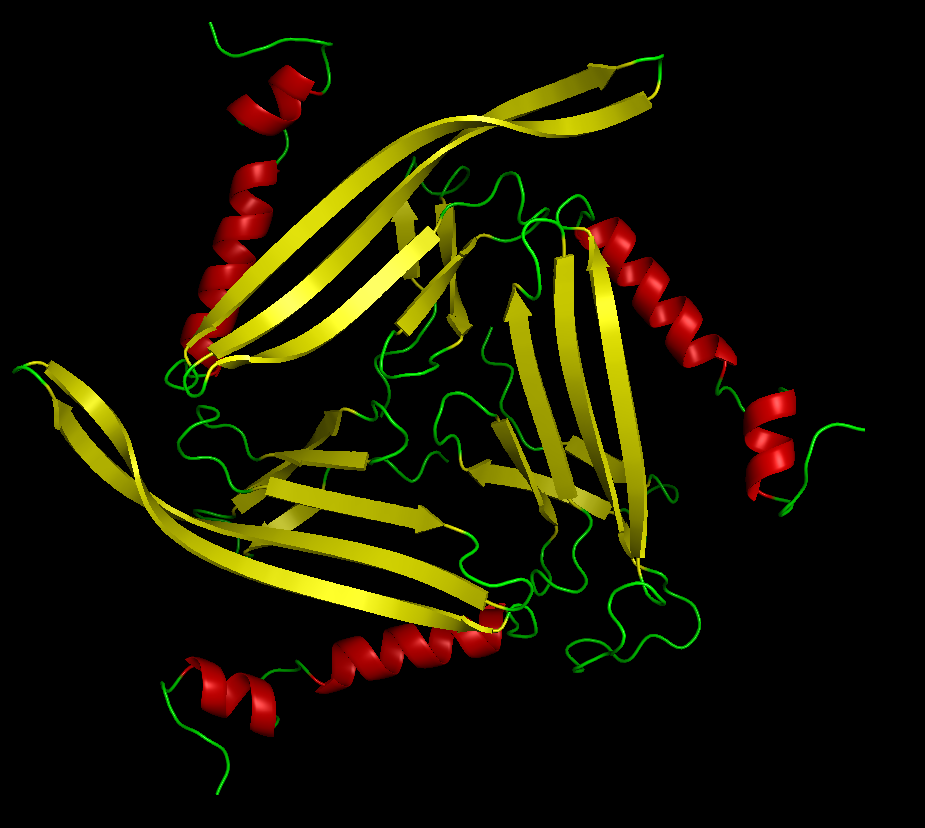
 It has more sheets, with 3 sections with very similar structures of sheets, helices, and loops.
It has more sheets, with 3 sections with very similar structures of sheets, helices, and loops.
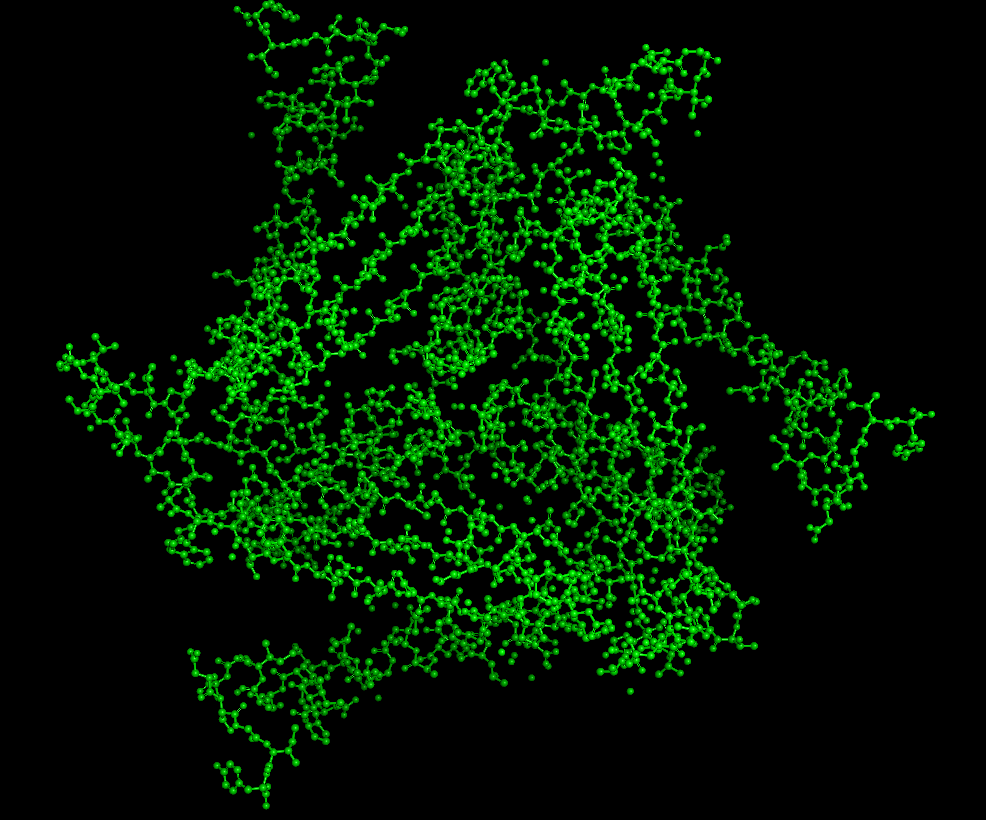
Colored by Residue Type
(all residues)
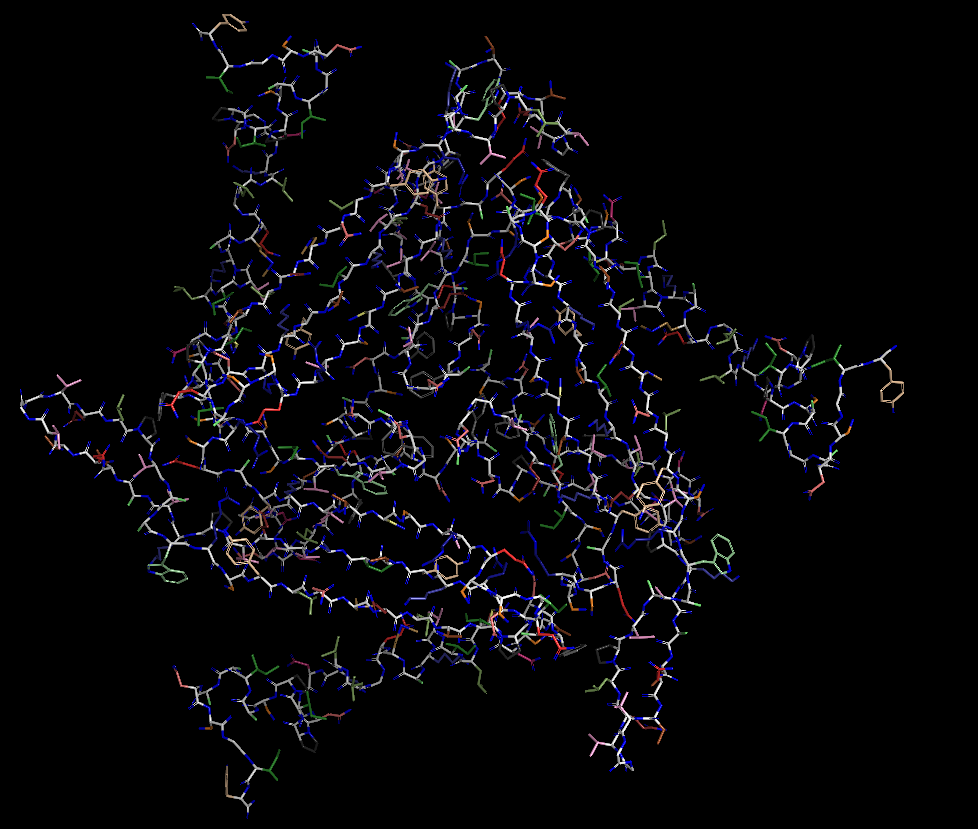
 (residue types, specific)
(residue types, specific)
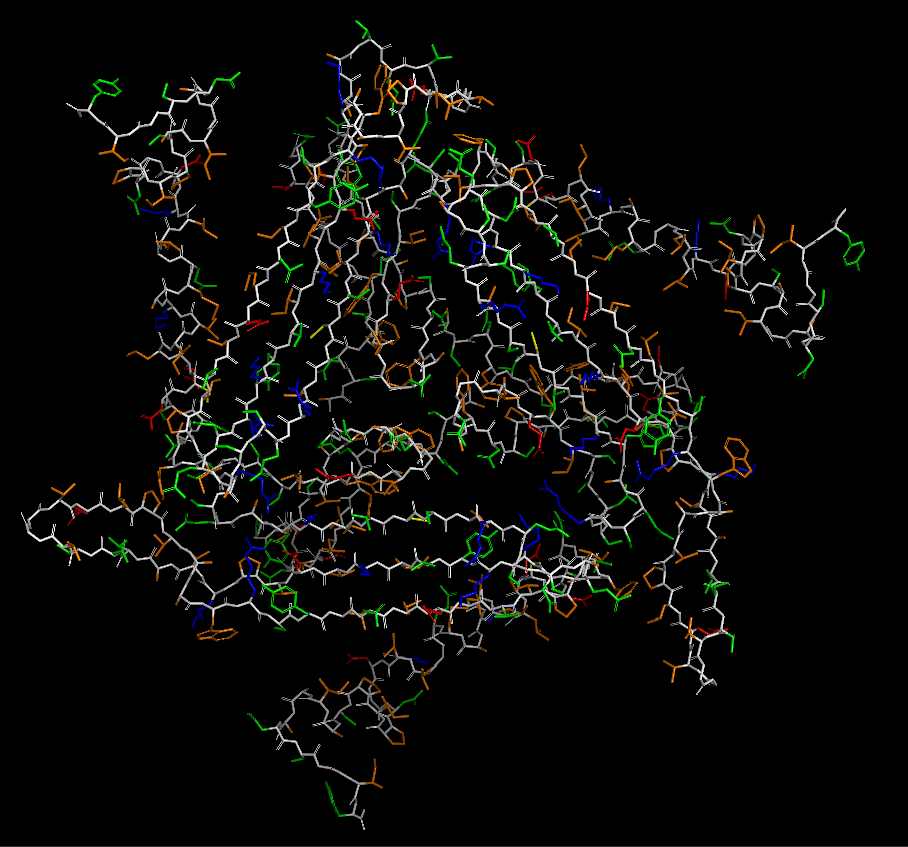
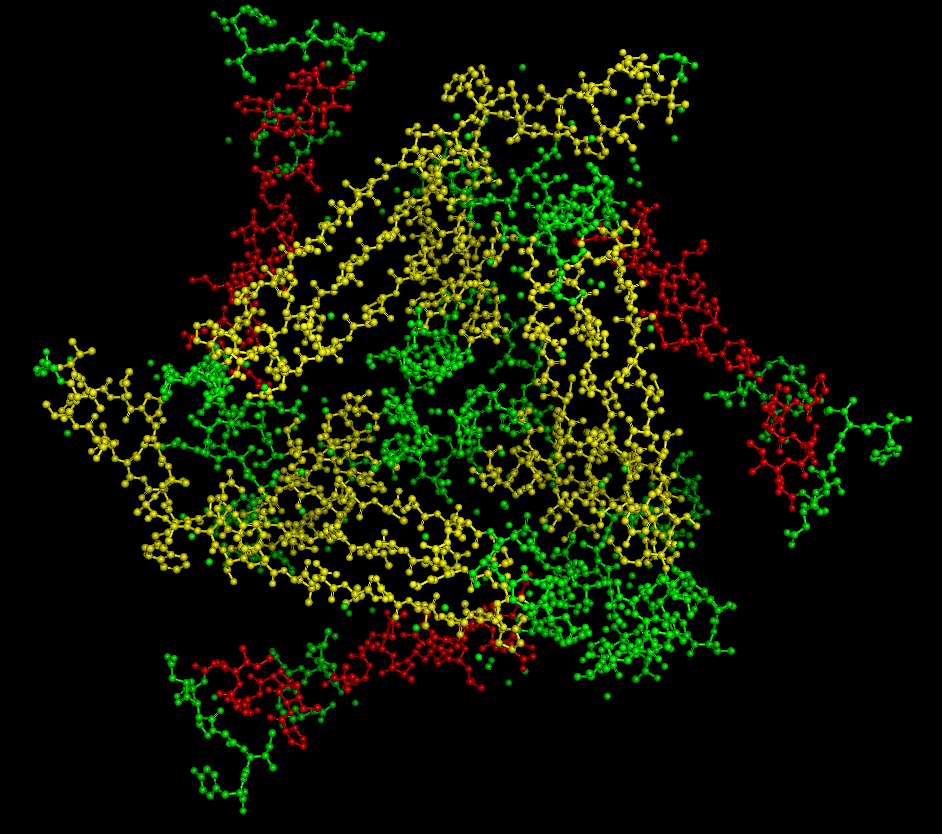
 (residue types, hydrophobic/hydrophilic)
(residue types, hydrophobic/hydrophilic)
 There are mostly polar/nonpolar residues. The hydrophobic and hydrophilic residues are evenly distributed throughout the protein.
There are mostly polar/nonpolar residues. The hydrophobic and hydrophilic residues are evenly distributed throughout the protein.
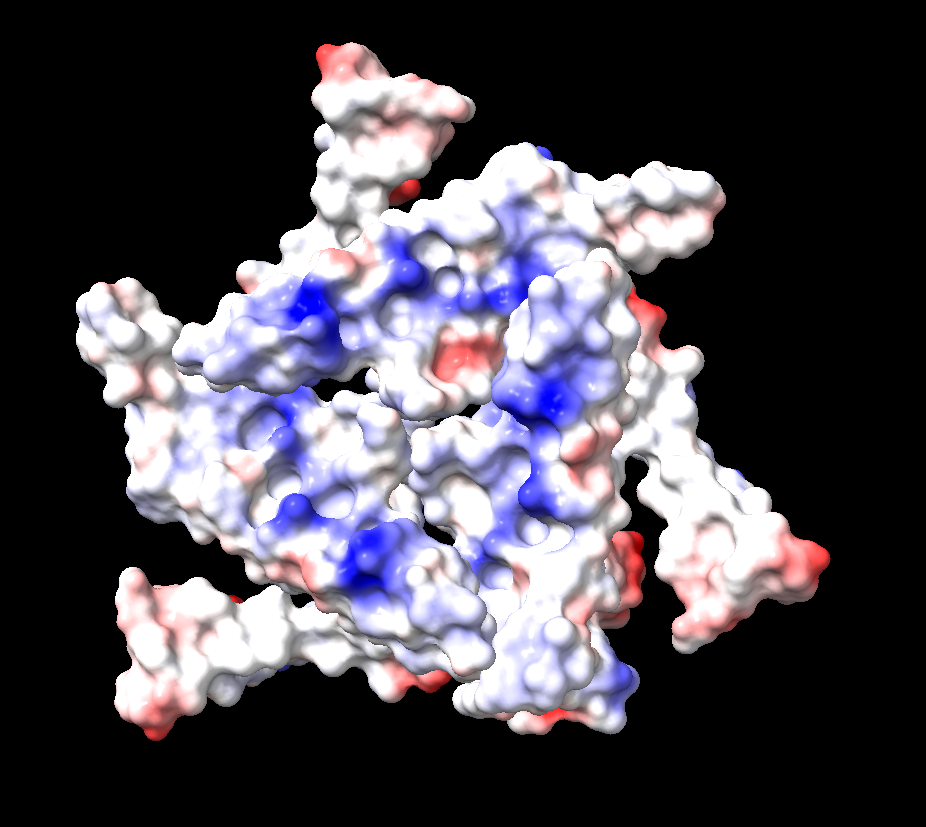
Surface
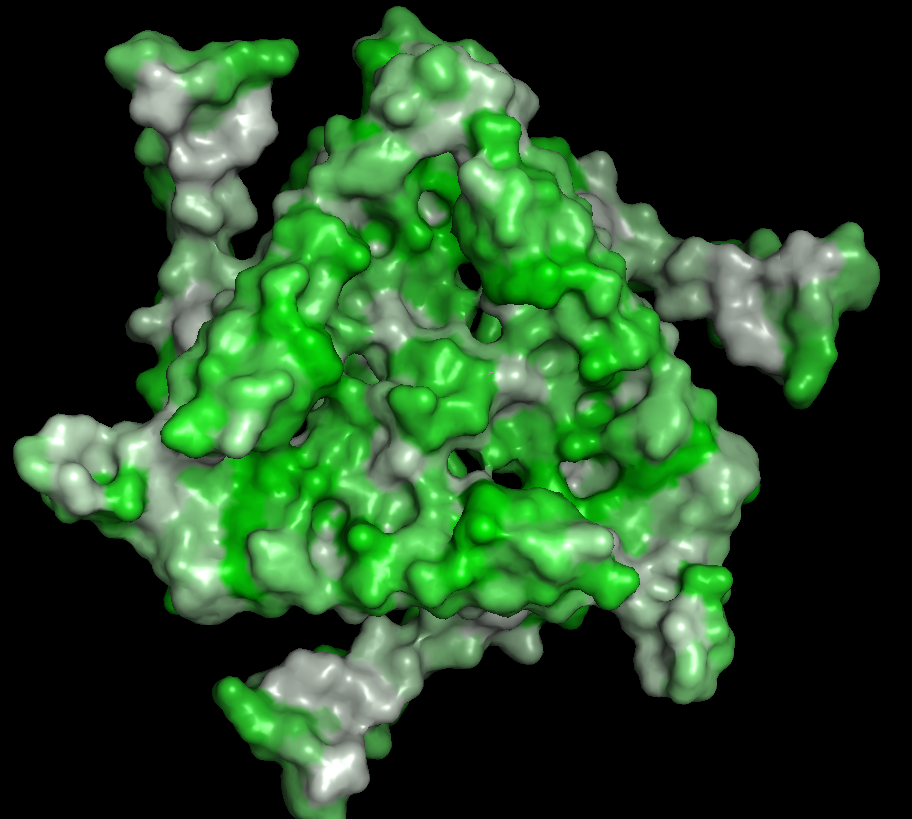

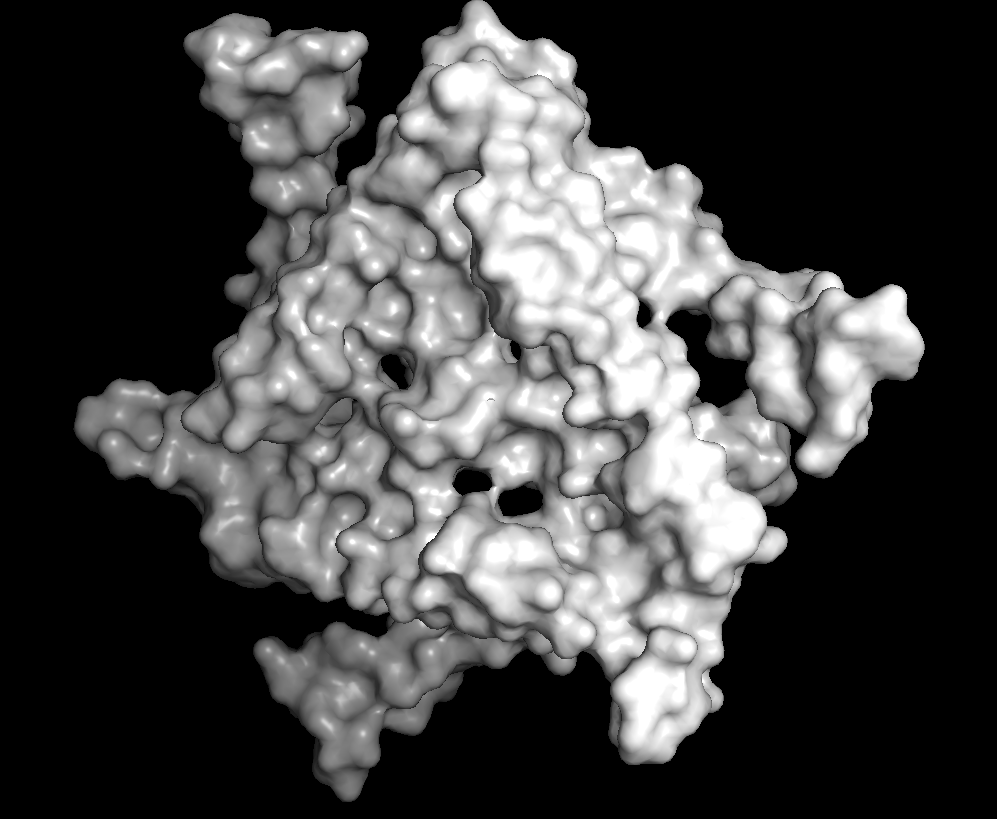
 There are many holes and structural pockets for things to bind.
There are many holes and structural pockets for things to bind.
Part C. Using ML-Based Protein Design Tools
C1. Protein Language Modeling
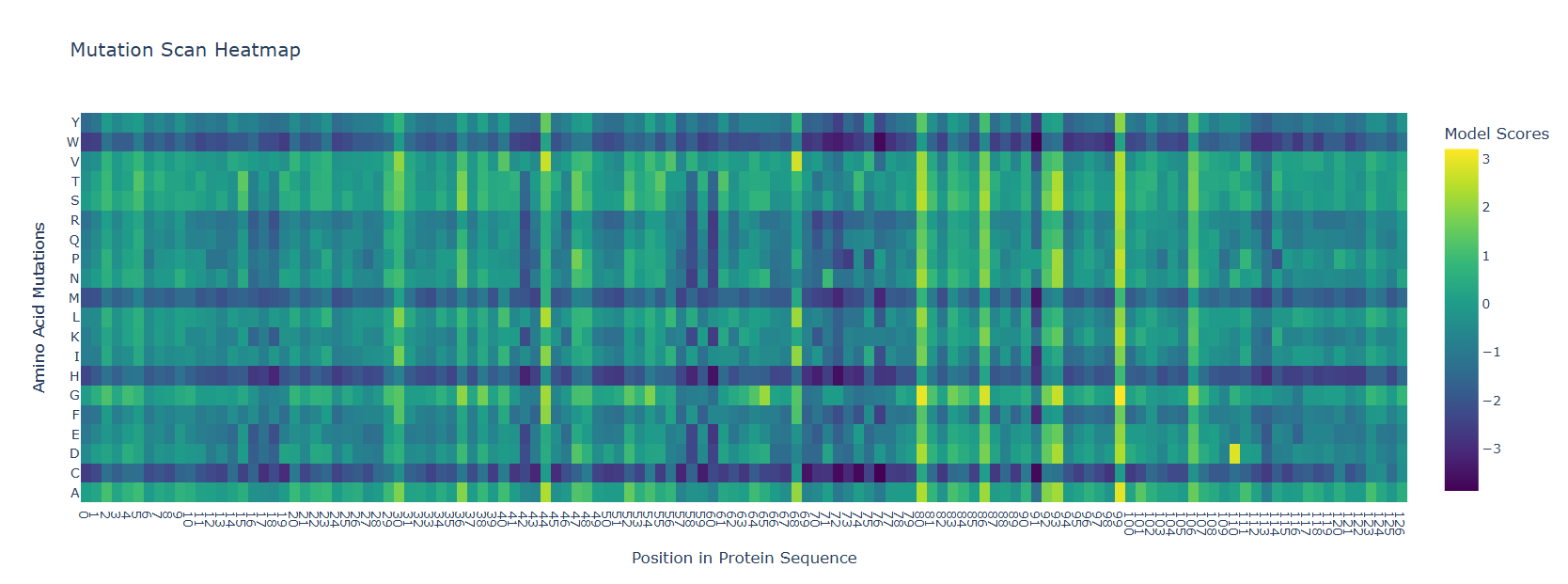
- Use ESM2 to generate an unsupervised deep mutational scan of your protein based on language model likelihoods.
- Can you explain any particular pattern? (choose a residue and a mutation that stands out)
- (Bonus) Find sequences for which we have experimental scans, and compare the prediction of the language model to experiment.
Latent Space Analysis
- Use the provided sequence dataset to embed proteins in reduced dimensionality.
- Analyze the different formed neighborhoods: do they approximate similar proteins?
- Place your protein in the resulting map and explain its position and similarity to its neighbors.
C2. Protein Folding
Folding a protein
- Fold your protein with ESMFold. Do the predicted coordinates match your original structure?
- Try changing the sequence, first try some mutations, then large segments. Is your protein structure resilient to mutations?
C3. Protein Generation
Inverse-Folding a protein: Let’s now use the backbone of your chosen PDB to propose sequence candidates via ProteinMPNN
- Analyze the predicted sequence probabilities and compare the predicted sequence vs the original one.
- Input this sequence into ESMFold and compare the predicted structure to your original.
Part D. Group Brainstorm on Bacteriophage Engineering
- Choose one or two main goals from the list that you think you can address computationally (e.g., “We’ll try to stabilize the lysis protein,” or “We’ll attempt to disrupt its interaction with E. coli DnaJ.”).
- Write a 1-page proposal (bullet points or short paragraphs) describing:
- Which tools/approaches from recitation you propose using (e.g., “Use Protein Language Models to do in silico mutagenesis, then AlphaFold-Multimer to check complexes.”).
- Why do you think those tools might help solve your chosen sub-problem?
- Name one or two potential pitfalls (e.g., “We lack enough training data on phage–bacteria interactions.”).
- Include a schematic of your pipeline.
- This resource may be useful: HTGAA Protein Engineering Tools