Subsections of 2026a-ceren-ozgen
Homework
Weekly homework submissions:
Week 1 HW: Principles and Practices
Project concept Insect Movement Memory I want to develop a living material surface that can register insect movement by responding directly to the chemical traces insects naturally leave behind. Instead of using cameras, sensors, or tracking software, the material itself would act as both sensor and display, making insect presence visible through temporary marks that emerge, accumulate, and fade over time.
Part 1: Benchling & In-silico Gel Art 1. Setup I created a Benchling account and opened the Lambda DNA sequence (NC_001416). Initially I had another plasmid open (pBR322_EGFR), but I switched to Lambda DNA after confirming the correct assignment requirements.
- Importing Lambda DNA I opened the Lambda phage genome (NC_001416, ~48.5 kb) inside Benchling. The sequence map displays multiple restriction enzyme sites distributed across the genome.
Subsections of Homework
Week 1 HW: Principles and Practices
Project concept
Insect Movement Memory
I want to develop a living material surface that can register insect movement by responding directly to the chemical traces insects naturally leave behind. Instead of using cameras, sensors, or tracking software, the material itself would act as both sensor and display, making insect presence visible through temporary marks that emerge, accumulate, and fade over time.
Insects communicate and navigate space using chemical signals such as pheromone trails, cuticular hydrocarbons, and contact-based secretions. These signals shape movement, coordination, and collective behavior, yet they remain largely invisible to humans. As a result, insect activity is often overlooked or treated as background noise—especially in built or maintained environments where cleaning, landscaping, and surface replacement routinely erase these traces.
This project explores whether a biofilm or engineered living material can make these otherwise invisible forms of communication perceptible, not by extracting or measuring them digitally, but by responding materially to their presence.
The system is guided by three core principles:
Passive sensing
The material responds directly to insect-deposited chemicals. No external imaging, electronics, or computation are required for the system to function. The surface itself performs the sensing.
Spatial memory
As insects move across the surface, their paths appear as visible patterns. Repeated visits, dense routes, or prolonged presence result in stronger or layered traces. These marks gradually fade, allowing the surface to hold memory temporarily rather than permanently.
Ecological framing
Rather than treating insects as interference or contamination, the system treats them as active contributors to space. By making their traces visible, and erasable, the project highlights how routine maintenance practices remove nonhuman forms of memory from shared environments.
Why Insect Movement Memory
This project aims to make insect communication visible in a way that is non-invasive, non-extractive, and legible to non-experts. It does not attempt to identify, track, or control insects. Instead, it reveals how insects already shape space through everyday movement.
Making these traces visible could support several constructive uses:
Ecological monitoring
Revealing where insects repeatedly travel, linger, or avoid can offer insight into habitat use without tagging or disturbing them.
Environmental indication
Insects respond quickly to changes in moisture, temperature, food availability, and surface conditions. Their movement patterns can act as indirect indicators of environmental change.
Public understanding and education
Seeing insect paths emerge on a surface can help people understand collective insect behavior, spatial coordination, and nonhuman forms of intelligence without requiring scientific background knowledge.
Material and design research
The project explores how living materials can encode time, interaction, and erasure—raising questions about what it means for a surface to remember, forget, and respond.
My motivation is both scientific and artistic. Scientifically, I want to test whether insect chemical traces can be used as inputs for material response without turning living systems into data sources. Artistically, I am interested in creating a surface that reveals how much nonhuman activity exists around us, and how easily it is removed, by simply allowing insects to write their own presence into material form.
Governance/Policy Goals
A. Prevent harm and misuse
A1. Don’t become a surveillance surface. Design and document the system so it cannot be easily used for covert monitoring of people’s spaces. The surface should not function as a “cheap tracker” for mapping occupancy or movement in homes, workplaces, or other sensitive environments.
A2. Don’t enable trapping, baiting, or targeted killing. Avoid configurations that make the material useful as a lure or control device (e.g., additives that attract insects, long-lasting “high-contrast” traces that help locate nests, or deployment guidelines that point toward pest-control use).
B. Biosafety and containment
B1. Keep the biological risk low by default. Use low-risk organisms and materials appropriate for the setting (e.g., well-characterized non-pathogenic strains) and favor formats that limit exposure and handling.
B2. Prevent persistence and unintended spread. Build in “off switches” and end-of-life steps: limited viability outside controlled conditions, clear deactivation methods, and safe disposal protocols. The material should be difficult to keep alive or propagate outside intended use.
C. Ecological responsibility
C1. Don’t meaningfully alter insect behavior. The goal is to read traces, not change them. Avoid deployments or chemistries that attract/repel insects or disrupt local movement patterns at scale.
C2. Treat field work as reversible and conservative. If testing outside controlled spaces is needed, keep it small, time-limited, and removable. Do not release engineered organisms into open environments; use containment-first formats for any public-facing trials.
D. Trust, consent, and framing
D1. Make deployments visible and accountable. If used outside the lab, the surface should be clearly labeled and explained: what it shows, what it does not show (e.g., it does not identify species or track individuals), and how long traces persist.
D2. Avoid “pest policing” narratives. Present the work as ecological interpretation and shared-space awareness, not as a tool for control. Use language and examples that resist fear-based framing and emphasize insects as participants in environmental systems.
Actions
Action 1: Containment-by-Design for Living Trace Materials
Purpose
Current creative and research uses of living materials often rely on informal judgments such as “non-pathogenic” or “low risk,” without addressing how a material behaves once it leaves the lab bench. For a surface that responds to insect chemical traces, the risk is not only biological spread but unintended persistence or reuse in uncontrolled environments. This action proposes a containment-by-design standard specifically for living trace materials, where non-release is the default condition rather than an exception.
Design
This action would be implemented within the course and host lab environment. Actors: student researcher, course staff, host lab supervisors Require a short Living Trace Material Containment Plan before fabrication or demonstration Limit material choices to BSL-1, non-pathogenic organisms Require immobilized or encapsulated formats (e.g., hydrogels, laminated films, sealed edges) that prevent propagation Define clear end-of-life protocols (dehydration, chemical kill, autoclaving, or bleaching depending on material) Establish a no outdoor deployment default unless explicitly reviewed and approved This mirrors containment approaches used in biomedical coatings and agricultural treatments, where physical form is regulated alongside biological function.
Assumptions
This action assumes that physical immobilization meaningfully reduces the likelihood of environmental persistence and that containment formats still allow sufficient sensitivity to insect-deposited chemicals. It also assumes that clear protocols can be followed consistently in creative research settings.
Risks of Failure and “Success”
Failure could occur if containment is treated as a formality rather than an active design constraint, leading to shortcuts or informal deployments. Conversely, success could make the system appear “safe enough,” potentially discouraging further reflection on ecological context or ethical use. Maintaining containment as a design commitment rather than a checkbox is therefore essential.
Action 2: Material-Limited Resolution to Prevent Surveillance or Control Use
Purpose
A surface that visualizes movement patterns risks being repurposed as a tracking or control tool, even if it is designed for insects. This action proposes embedding ethical limits directly into the material’s expressive capacity, so it cannot easily be used for surveillance, behavioral manipulation, or targeted pest control.
Design
Ethical constraints are implemented through material behavior rather than policy alone. Actors: student researcher, advisors, material collaborators Design the surface to produce low-resolution, diffuse traces rather than sharp, directional paths Use short, predictable fading windows (hours to days) to prevent long-term accumulation Exclude any attractants, repellents, or lures from the system Avoid encoding species-level identification or individual differentiation These design choices ensure the system shows presence and density rather than actionable movement maps.
Assumptions
This action assumes that reducing resolution meaningfully limits misuse and that ethical intent can be expressed through material affordances. It also assumes viewers will interpret traces as indicative rather than authoritative representations.
Risks of Failure and “Success”
Failure may occur if the surface becomes too subtle to read or if users still interpret patterns as precise maps. At the same time, success could unintentionally inspire attempts to “improve” resolution, pushing the system toward more extractive uses. Clear framing and documentation are therefore necessary alongside material limits.
Action 3: Responsible Documentation and Partial Disclosure
Purpose
Open sharing is a core value of art–science research, but some technical details, such as recipes for long-lasting, field, deployable living surfaces, could enable misuse or premature deployment. This action proposes a responsible documentation strategy that balances transparency with restraint.
Design
Documentation is divided into two tiers. Actors: student researcher, course instructors, host lab Public documentation: concept, ecological framing, ethics, failures, and non-deployable prototypes Restricted documentation (course/lab context): detailed material formulations, tuning parameters, and persistence optimization Include a short misuse reflection in public-facing materials describing plausible misuse pathways and why they are discouraged This approach draws from responsible disclosure practices in biology and AI, without claiming to eliminate risk.
Assumptions
This action assumes that partial disclosure meaningfully reduces harm while still enabling critique and learning. It also assumes that those seeking misuse would require detailed implementation knowledge, which may not always be true.
Risks of Failure and “Success”
Failure could occur if information spreads regardless of intent, making gating largely symbolic. Success, however, could create access inequities, where only well-connected groups obtain deeper technical knowledge. These risks make transparency about why details are gated as important as the gating itself.
Action 4: Consent and Visibility for Public-Facing Deployments
Purpose
Even when focused on insects, a surface that reveals movement patterns can feel surveillance-like if deployed covertly. This action establishes consent and visibility as conditions for any use outside controlled lab or studio contexts.
Design
Actors: researchers, galleries, institutions, site hosts Require clear signage explaining what the surface does and does not record (e.g., no human identification, no tracking) Limit deployments to short, documented time windows Prohibit use in private or intimate spaces (homes, workplaces, classrooms) without institutional review Document that insects are not baited, trapped, or harmed
Assumptions
This action assumes that visibility and consent meaningfully reduce covert misuse and that institutions are willing to support ethical deployment requirements.
Risks of Failure and “Success”
Failure could occur if signage is ignored or consent is bypassed. Success may make deployment more administratively complex, potentially limiting grassroots or community use. However, this friction reinforces the project’s stance that interpretation should not come at the cost of trust.
| Does the option: | Option 1 | Option 2 | Option 3 |
|---|---|---|---|
| Enhance Biosecurity | |||
| • By preventing incidents | 1 | 2 | 2 |
| • By helping respond | 2 | 2 | 2 |
| Foster Lab Safety | |||
| • By preventing incident | 1 | 3 | 3 |
| • By helping respond | 1 | 3 | 3 |
| Protect the environment | |||
| • By preventing incidents | 1 | 2 | 1-2 |
| • By helping respond | 2 | 2 | 2 |
| Other considerations | |||
| • Minimizing costs and burdens to stakeholders | 2 | 2 | 3 |
| • Feasibility? | 1 | 2 | 2 |
| • Not impede research | 2 | 2 | 3 |
| • Promote constructive applications | 1 | 2 | 2 |
Prioritization and trade-offs
Based on the scoring and the nature of the project, I prioritize Option 1: Containment-by-Design for Living Trace Materials, with Option 3: Consent and Visibility for Public-Facing Deployments as a necessary secondary measure.
Option 1 addresses risk at the most fundamental level: the material’s physical and biological behavior. Because Insect Movement Memory relies on living or semi-living materials that respond to environmental input, the main concern is not malicious use but accidental persistence, spread, or reuse outside controlled settings. Embedding containment into the material format, through immobilization, encapsulation, and clear end-of-life protocols—reduces these risks early and effectively, without depending on later enforcement or user discretion. This approach is also well suited to a course-based research context, where lab practices and instructor oversight are already in place.
Option 3 becomes important once the work moves beyond the lab into exhibitions or public demonstrations. Even though the system targets insects, visible movement traces could be misread as surveillance if deployed without explanation. Requiring visibility, signage, and consent helps prevent misinterpretation and covert use, and reinforces the project’s ecological framing.
The trade-off is reduced speed and flexibility. Containment and consent requirements can slow experimentation and complicate public display, and they assume consistent compliance by researchers and institutions. However, lighter governance at early stages would risk misuse or ecological harm that would be far more difficult to correct later.
This prioritization is aimed primarily at research labs and course instructors, and secondarily at galleries or public venues hosting work with living materials.
Thought Process
This project grows out of my ongoing interest in how nonhuman beings already write, sense, and organize space, and how design often ignores or erases those processes. Across my work—movement-based workshops, multispecies design experiments, and projects like Insect Techno, I keep returning to the same question: what changes when we stop treating nonhuman life as background and instead let it shape form, rhythm, and meaning?
Insect Movement Memory emerged from noticing a mismatch. Insects rely heavily on chemical traces to navigate and communicate, yet these traces are systematically removed through cleaning, maintenance, and landscaping. What remains visible are human priorities: smoothness, sterility, and control. I became interested in whether a material could sit quietly within this system and register what is already happening, without adding sensors, data pipelines, or behavioral manipulation.
My previous work with movement and performance, especially in collective, embodied contexts, has shaped how I think about this material. In events like Insect Techno, the goal was not to represent insects accurately, but to shift perception: to invite humans to feel attunement, rhythm, and coordination beyond the human scale. Similarly, this project is less about measuring insects and more about making their presence felt—through traces that appear, accumulate, and disappear.
I am deliberately resisting extractive logics common in sensing and monitoring systems. I do not want to identify species, optimize paths, or turn insect behavior into data streams. Instead, I am interested in material responsiveness as interpretation: a surface that reacts, remembers briefly, and then forgets. The fading of traces is as important as their appearance, reflecting how nonhuman memory is fragile and easily overwritten in shared environments.
Thinking through governance and ethics became part of the design itself. The more I developed the idea, the clearer it became that the risks were not dramatic misuse scenarios, but subtle ones: accidental persistence, misinterpretation as surveillance, or reuse in pest-control contexts. This pushed me toward containment-by-design, low-resolution material responses, and clear framing—choices that align with my broader commitment to care-oriented, more-than-human design.
Overall, this project is a way for me to bring together my interests in embodiment, multispecies relations, living materials, and speculative yet grounded design. It treats insects not as subjects to be managed, but as co-authors whose movements already shape space—whether we notice them or not.
AI Statement
I used AI primarily as a language and structure aid while writing this assignment. My project idea, framing, and ethical concerns—especially around insect chemical communication, passive sensing, containment, and non-extractive design—were already formed through my prior work in multispecies design, movement-based research, and projects like Insect Techno. AI helped me refine wording, tighten arguments, and check clarity across sections, particularly in the governance and prioritization parts of the assignment.
AI was not used to generate biological protocols, engineered organisms, material recipes, or experimental methods. All technical assumptions, design constraints, and ethical positions reflect my own judgment and responsibility. I treated AI as a tool for articulating ideas rather than producing them, using it to make my thinking legible to an interdisciplinary audience without delegating scientific or ethical decision-making.
Week 2 HW: DNA r/w/e
Part 1: Benchling & In-silico Gel Art
1. Setup
I created a Benchling account and opened the Lambda DNA sequence (NC_001416). Initially I had another plasmid open (pBR322_EGFR), but I switched to Lambda DNA after confirming the correct assignment requirements.
2. Importing Lambda DNA
I opened the Lambda phage genome (NC_001416, ~48.5 kb) inside Benchling. The sequence map displays multiple restriction enzyme sites distributed across the genome.
3. Restriction Digest Exploration
I simulated restriction digests using the following enzymes:
EcoRI HindIII BamHI KpnI EcoRV SacI SalI
Initially I was confused and didn’t really understand what I was “filtering.” Benchling scans the DNA, finds specific enzyme patterns, simulates cuts, and gives fragment sizes, but at first I kept thinking: then what? I wasn’t familiar with the terminology or the real-world application, so it felt abstract.
To understand better, I asked ChatGPT and watched several explanatory videos. Even then I struggled a bit, but gradually I realized that each enzyme works like a specific filter or rule. By testing different enzymes, I could start to see the pattern differences they create in the DNA fragments.
This helped me understand that I was learning how different enzyme “filters” generate different visual rhythms, so I could begin deciding which enzymes to use intentionally for my visual composition.
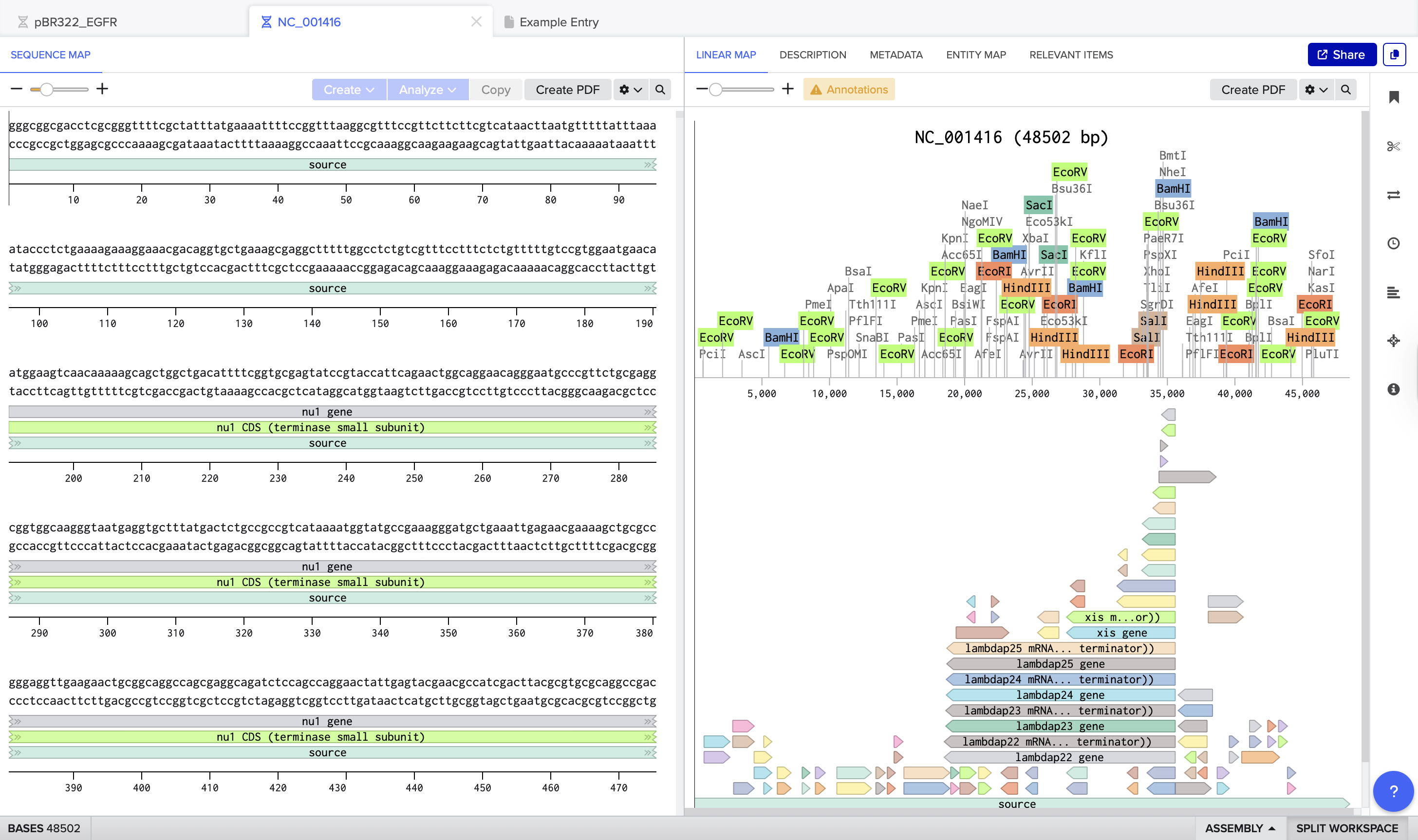

After running the digest, I could see how each selected enzyme produced a different number of cuts and fragment sizes. For example, EcoRV generated many cuts(21), creating a dense distribution of smaller fragments, while EcoRI(5) and HindIII(6) produced fewer cuts.
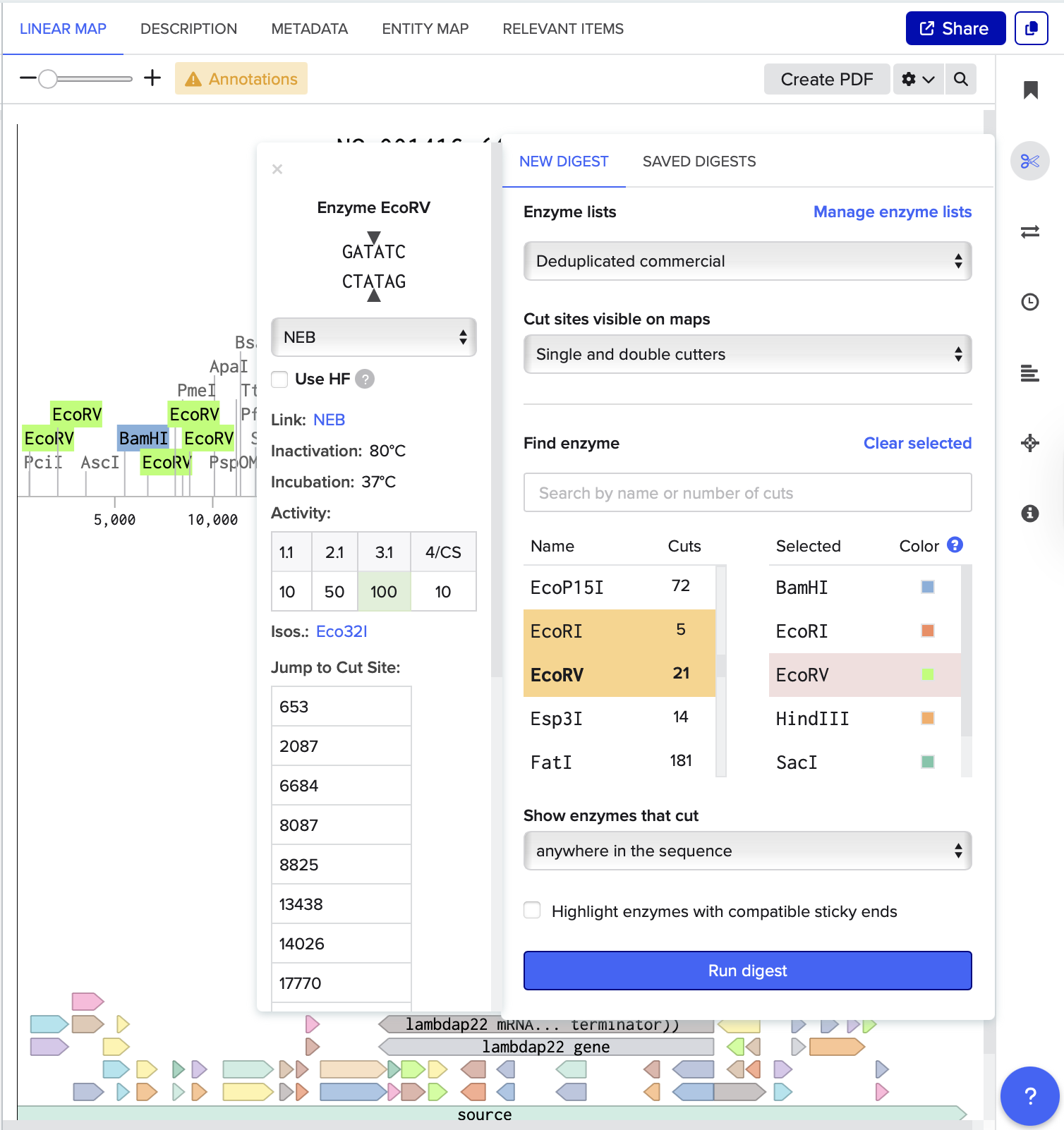

I also noticed that some enzymes, such as KpnI and SacI, did not appear prominently in the results or produced very few cuts in this sequence.
4. Gel Art Design Strategy
At this stage I stopped thinking about the enzymes only as a biology tool and started looking at them as pattern generators. Each enzyme produced different fragment distributions, so I explored how switching enzymes changed the visual rhythm of the gel.
The interface doesn’t allow precise editing of each lane. I couldn’t manually design bands. I had to cycle through combinations using the arrows and observe what happened. Because of that, my strategy became more about selecting which enzymes to keep active and then browsing through outcomes until I found something that felt balanced.
5. Iterations & Failed Attempts
Before this final version, I tried to create clearer shapes and more intentional patterns. I turned on many enzymes at once hoping to get complex results, but most of the outputs looked messy or repetitive.
One frustration was the lack of control. I expected to design the image directly, but the system works more like a generator. Many early attempts felt too basic or visually similar even when I changed enzymes.
After several tries, I stopped chasing a specific image and focused more on choosing a composition that felt visually stable.
6. Final Gel Composition
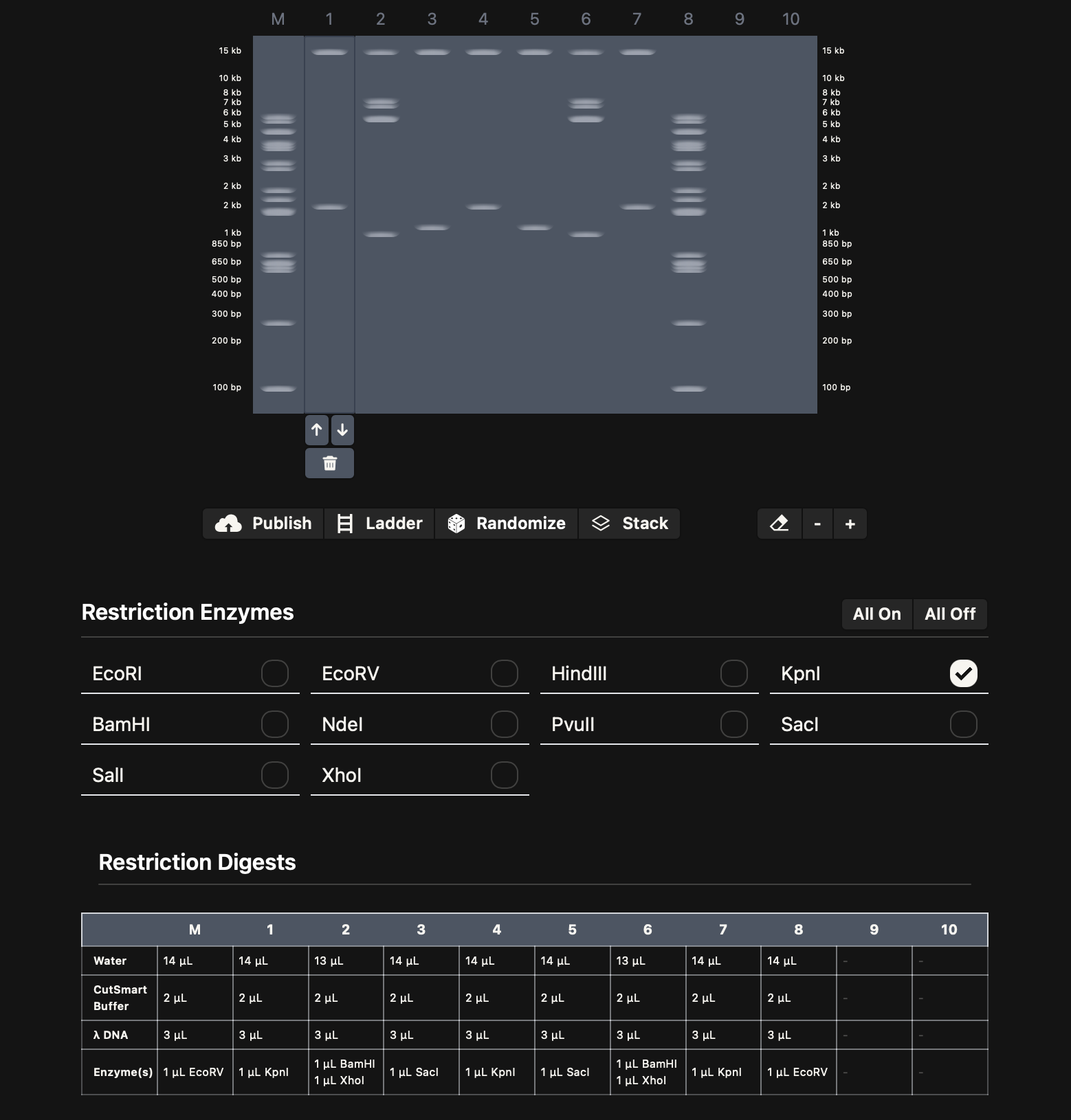

The final gel is simpler than what I first imagined, but it feels more readable. Instead of forcing a highly detailed figurative pattern, I aimed for a subtle, animal-like smiling face emerging from the band distribution.
Some lanes are denser. I imagined them as hair (EcoRV-heavy), while others remain more minimal (EcoRI / HindIII), creating lighter spacing that suggests the smile or mouth. I also used combined enzyme digests in lanes 2 and 6 because those areas form both the eyes and part of the mouth, helping balance the composition without making it too crowded.
7. Reflection
At the beginning I didn’t really understand what I was controlling or what “cutting or filtering” meant. The terminology and interface felt confusing, and I kept asking myself, “okay… but then what?”
Over time I realized that the control comes from choosing which enzymes to use, not from drawing the image directly. The system generates patterns automatically, so my role was more about testing, observing, and selecting what worked visually.
Honestly, I still think the interface limits how expressive the results can be. Many outputs felt repetitive, and it was harder than I expected to shape a strong visual composition. But the process helped me see how biological tools can be used in a more creative, exploratory way instead of only as technical steps.
Part 3: DNA Design Challenge
1. Choose your protein
For this assignment, I selected Green Fluorescent Protein (GFP). GFP is a protein originally isolated from the jellyfish Aequorea victoria. It emits green fluorescence when exposed to blue or ultraviolet light. I chose this protein because it is widely used in biotechnology as a visual reporter, allowing researchers to observe gene expression and cellular processes in real time. As a designer interested in perception and visualization, GFP represents a powerful example of how biological systems can translate invisible molecular activity into visible signals.
Protein sequence (sfGFP):
Protein sequence source: UniProtKB – P42212 (Green Fluorescent Protein, Aequorea victoria) https://www.uniprot.org/uniprotkb/P42212
2. Reverse Translate: Protein → DNA
At the beginning, I struggled to understand what “reverse translation” actually meant. I was confused about how one protein sequence could correspond to many possible DNA sequences, because I initially imagined it as a chemical state rather than a coding system. What helped me was realizing that DNA works more like a code than an ingredient. The sequence is read in groups of three letters, and different three-letter combinations can still produce the same amino acid. This means that reverse translation does not recover the original DNA, but instead generates one possible nucleotide sequence that could produce the protein.
To complete this step, I copied the GFP amino acid sequence from UniProt and used an online reverse translation tool. I learned that I do not need to manually convert each amino acid; instead, the tool assigns codons based on the genetic code. Through this process, I began to understand that the DNA sequence functions more like an instruction format than a visual structure, and that multiple DNA variations can lead to the same final protein.
To complete this step, I copied the GFP amino acid sequence from UniProt and used a bioinformatics reverse translation tool. I learned that I do not need to manually convert each amino acid; instead, the tool assigns codons based on the genetic code.
Example reverse-translated DNA sequence:
3. Codon Optimization
Different organisms prefer different codons for the same amino acid because they have different amounts of transfer RNA (tRNA). If a gene uses rare codons for a host organism, protein expression becomes inefficient.
For the codon optimization step, I created an account on the Twist Bioscience platform and uploaded the GFP amino acid sequence. I selected Escherichia coli as the codon usage table because it is a common organism used for protein expression and was also discussed during recitation.
When several pop-up warnings appeared, such as messages about missing stop codons and vector assignment. I was confused about whether I needed to order the gene or select cloning options, since my goal was only to generate an optimized sequence for the assignment. I asked ChatGPT to help me interpret these warnings, and I learned that the platform assumes users want to synthesize DNA, while I only needed the optimized output sequence.
After selecting the codon usage table, the platform automatically generated a new nucleotide sequence that keeps the same protein but changes the DNA letters to better match how E. coli prefers to read codons.
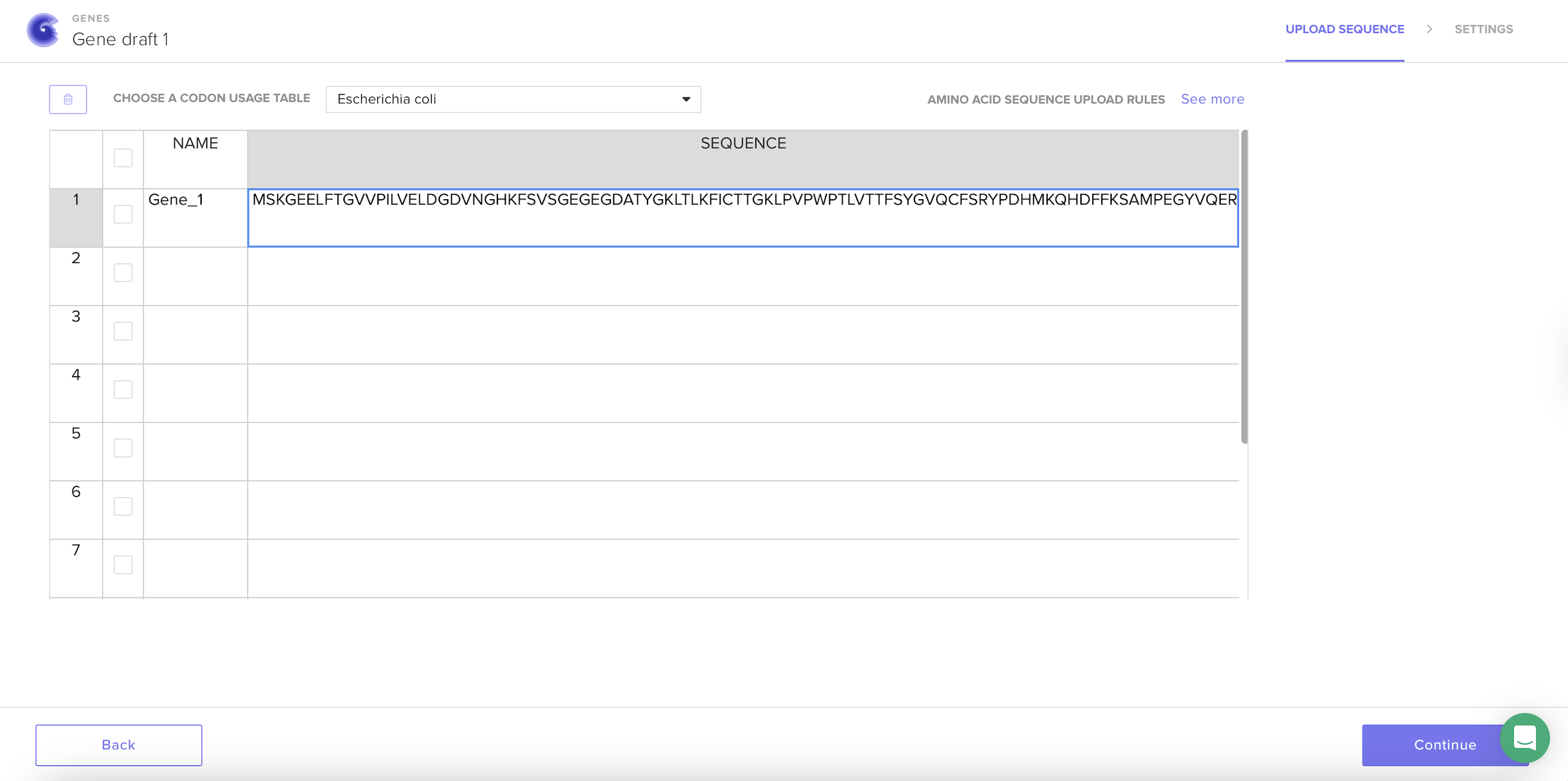

I understand that codon optimization improves; translation efficiency, protein yield and mRNA stability.
I optimized the GFP sequence for E. coli because it grows quickly. It is commonly used for protein production and plasmid expression systems are well established.
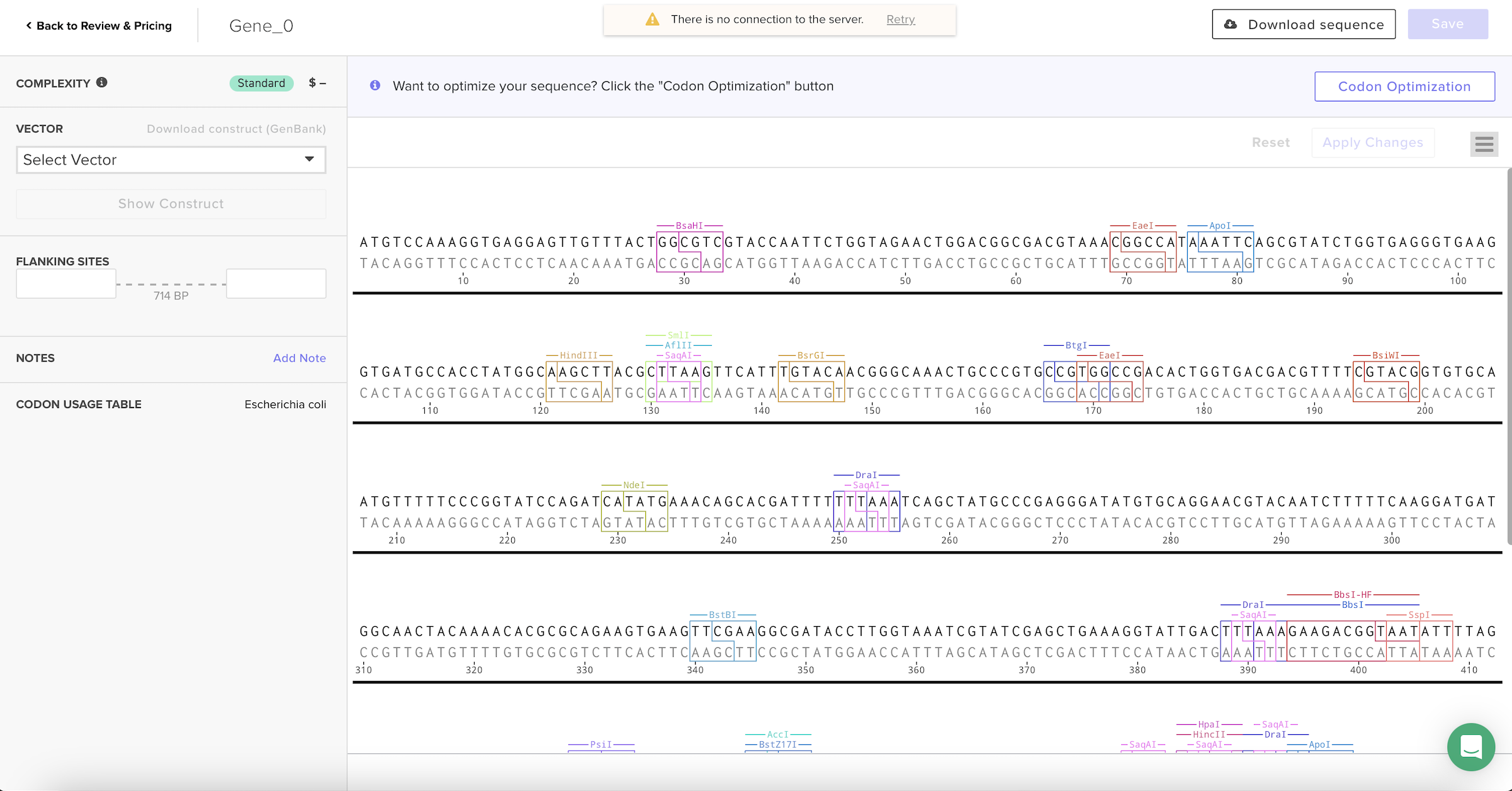

4. You have a sequence! Now what?
Once the codon-optimized DNA sequence is obtained, several technologies can produce the GFP protein.
Cell-dependent method (using bacteria) The optimized GFP gene is inserted into a plasmid vector containing: promoter ribosome binding site antibiotic resistance gene
The plasmid is transformed into E. coli.
Inside the bacteria: DNA is transcribed into mRNA. Ribosomes translate mRNA into GFP protein.
The bacteria glow green under UV light, indicating successful expression.
Cell-free method Instead of living cells, a cell-free expression system uses extracted transcription and translation machinery. Mixing the DNA template with enzymes, ribosomes, and energy sources allows GFP production in a test tube. This approach is faster and avoids cellular complexity.
5. How one gene produces multiple proteins
Labs
Lab writeups:
Subsections of Labs
Week 1 Lab: Pipetting
Projects
Final projects:
Subsections of Projects
Individual Final Project
Group Final Project
Ceren — HTGAA Spring 2026


About me
Hi, I’m Ceren. I’m curious by nature and enjoy learning through doing and exploring new ways of working. I work at the intersection of design, embodiment, and more-than-human worlds, with a background in industrial design and urban planning. I’m a kinesthetic learner drawn to experience design, movement-based practices, and material experimentation, and I’m excited to learn how to grow and work with living systems through hands-on biological making.