Homework
Weekly homework submissions:
Week 1 HW: Principles and Practices
Class Assignment — DUE BY START OF FEB 10 LECTURE Question 01 First, describe a biological engineering application or tool you want to develop and why. This could be inspired by an idea for your HTGAA class project and/or something for which you are already doing in your research, or something you are just curious about. According to the well-regarded popular science writer Matthew Cobb (2022), since the Asilomar conference in 1975, molecular biologists have been the vanguard in self-regulating when playing God. This means we refrain from conducting our research irresponsibly by deploying unnecessarily hazardous experimental methods. Alas, this also means that some of the most exciting genetic engineering is no longer done. Consider Dr. Oswald Avery’s transforming principle experiment. Blindly take a population of virulent pneumonia bacteria and feed them harmless kin until they lose their aggressive function and magically adapt into weak and indifferent pneumonia. Since Asilomar, this is indeed one kind of experiment that trustworthy principal investigators must abstain from. I get it, and still I contemplate. Wasn’t Avery the best of us, though? Between Schrodinger and Watson, Crick, and Franklin – Dr. Avery intuited DNA into existence with his transforming principle and used it effectively. Surely I didn’t name my oldest son after this man for nothing?
Week 2 HW: DNA Read Write and Edit
Table of contents Software used: Terminal, git, xcode, hugo, benchling, rcdonovan website, twist website. Objective: This week explores the read–write–edit toolkit: sequencing and synthesis workflows, restriction digests and gel electrophoresis, and early genome-editing frameworks.
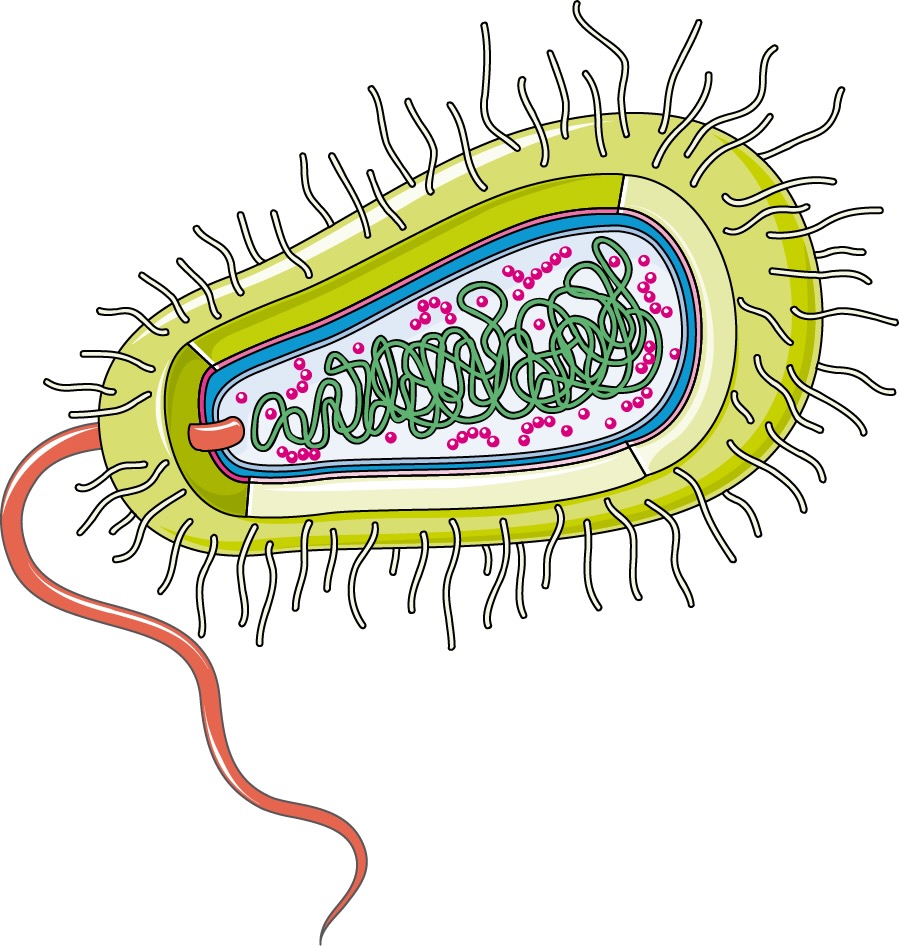
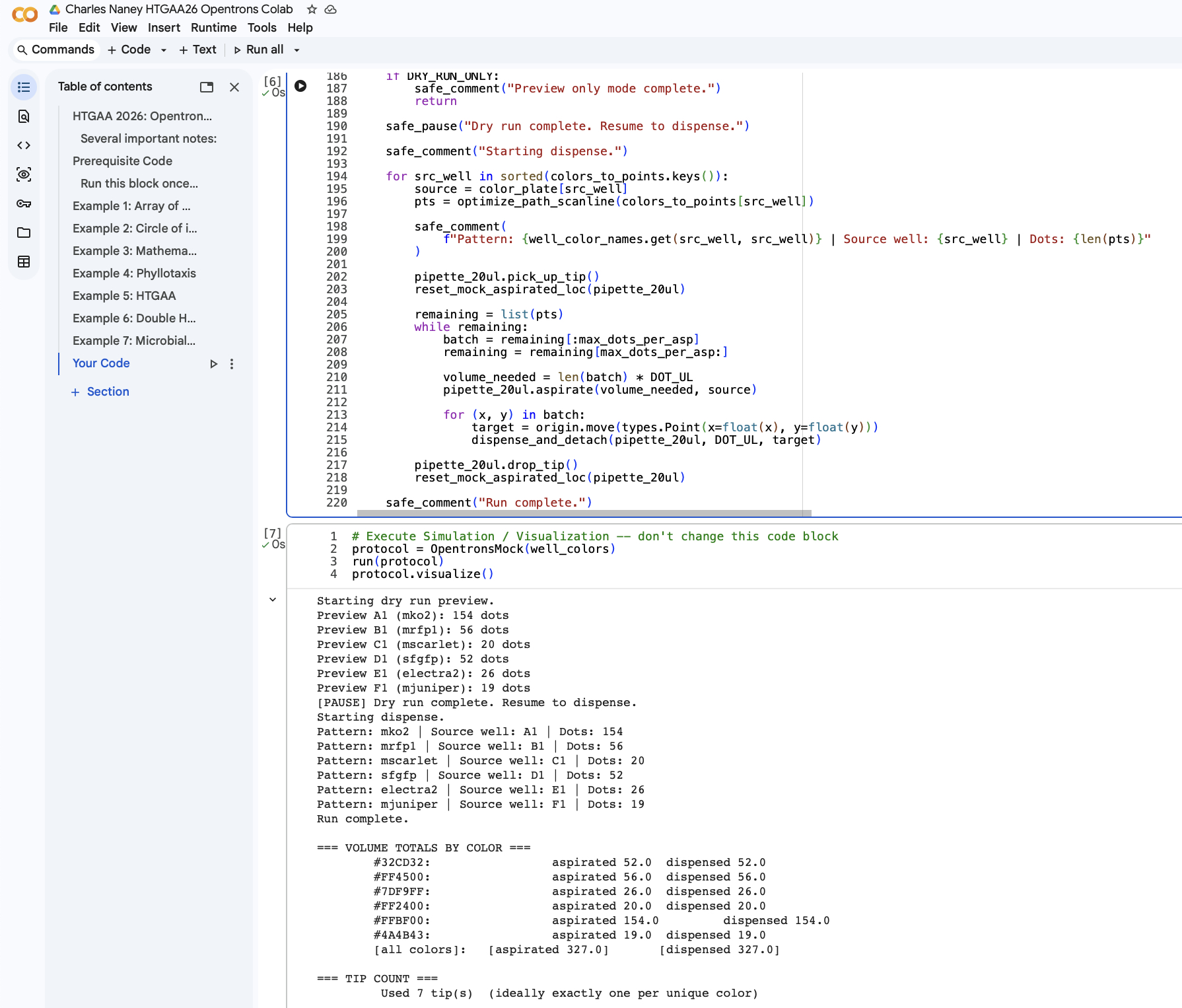
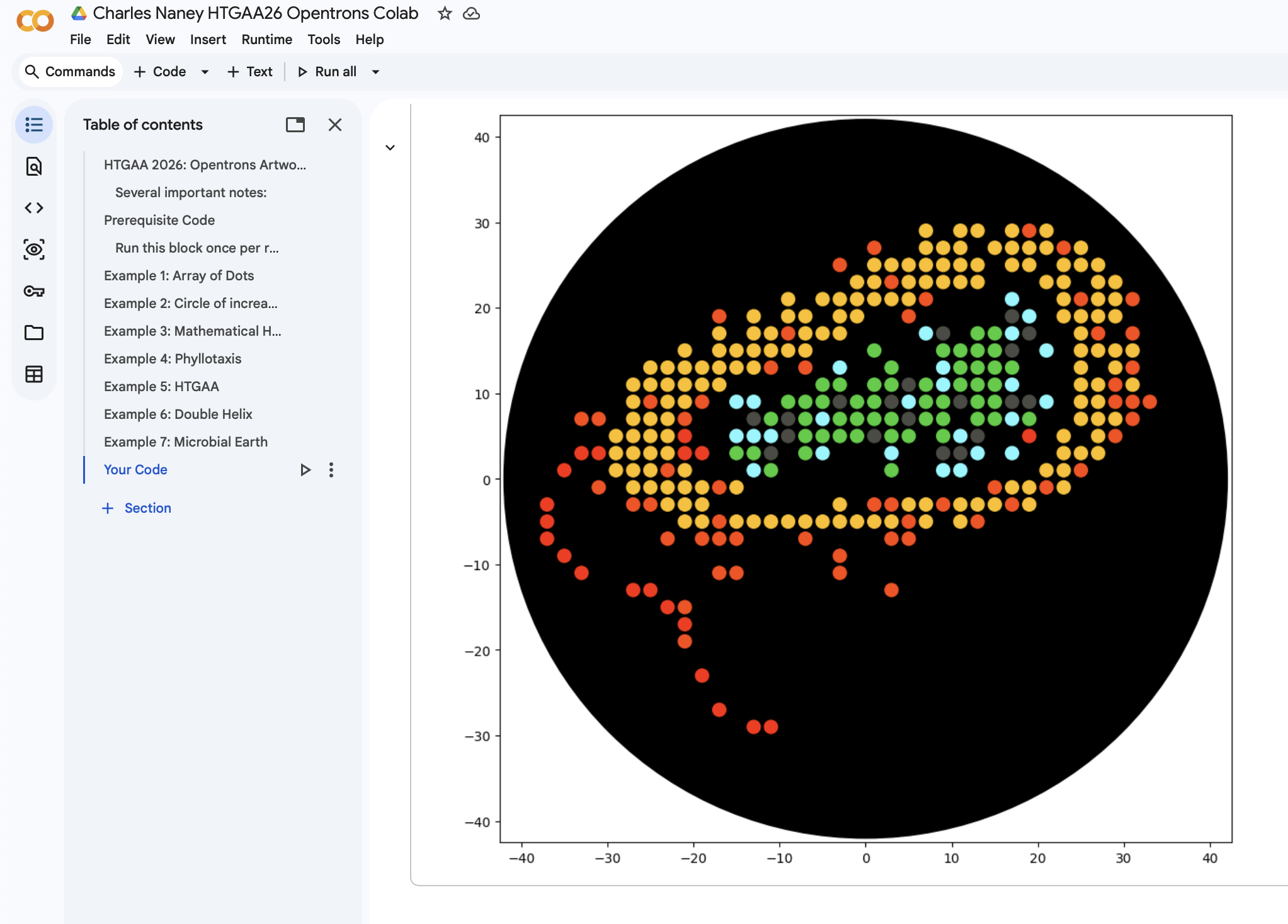
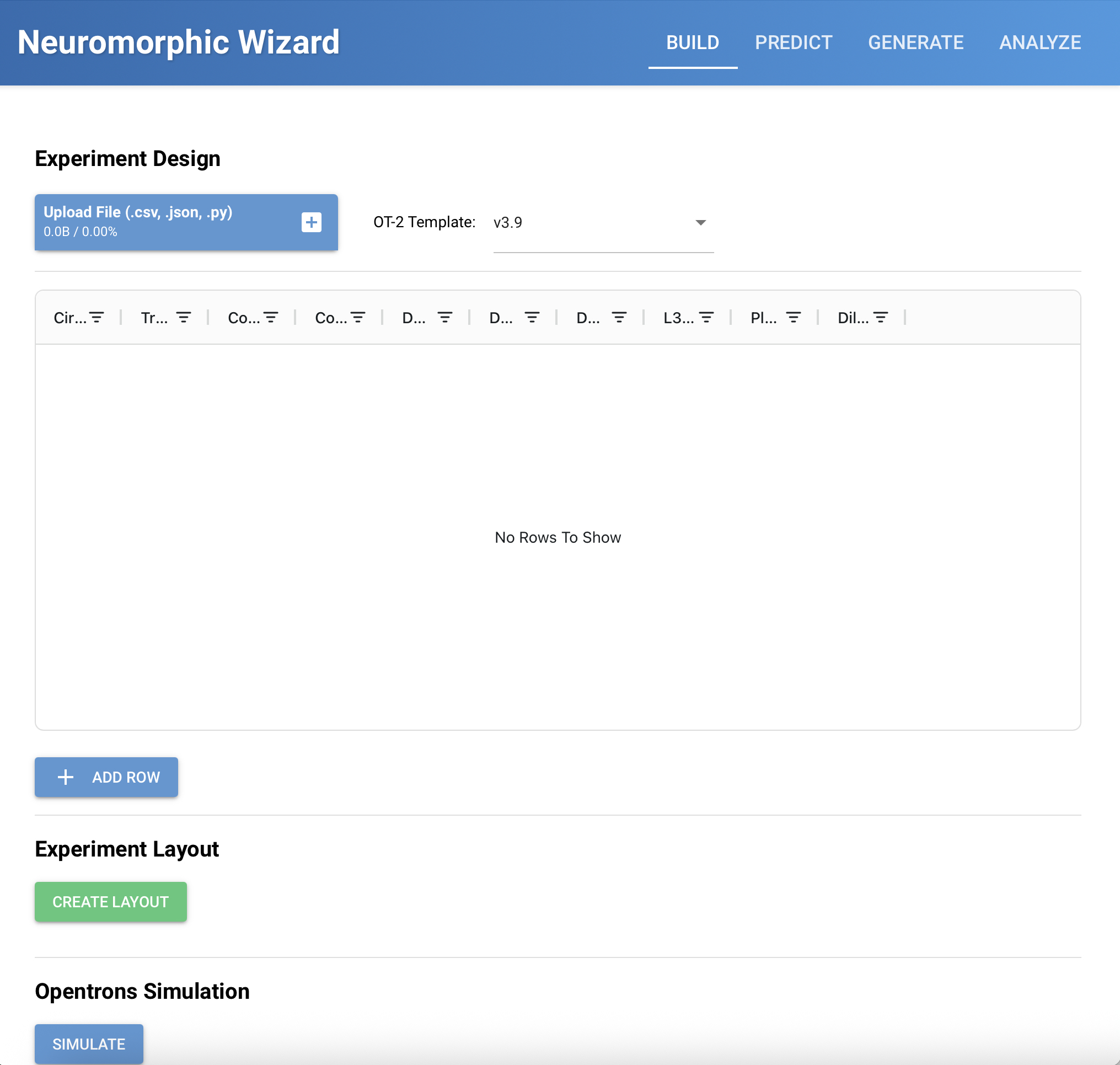
Homework for HTGAA 2026 (Week 03): Lab Automation Table of contents Software used: Terminal, git, Opentrons, rcdonovan website, Google Colab. Objective: This week we get hands-on (or at least code-on) with pipetting robots.
Week 4 HW: Protein Design Part I
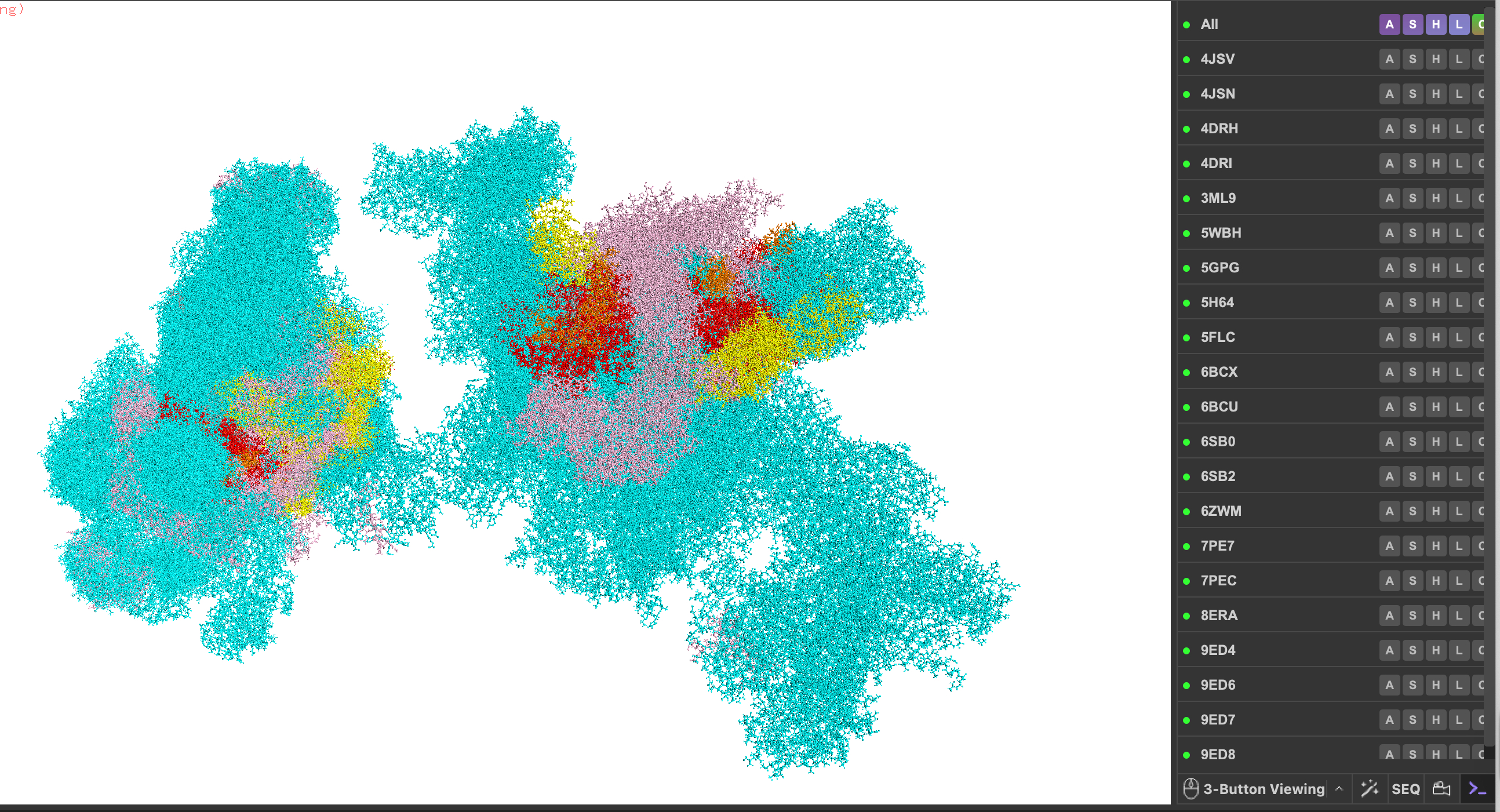
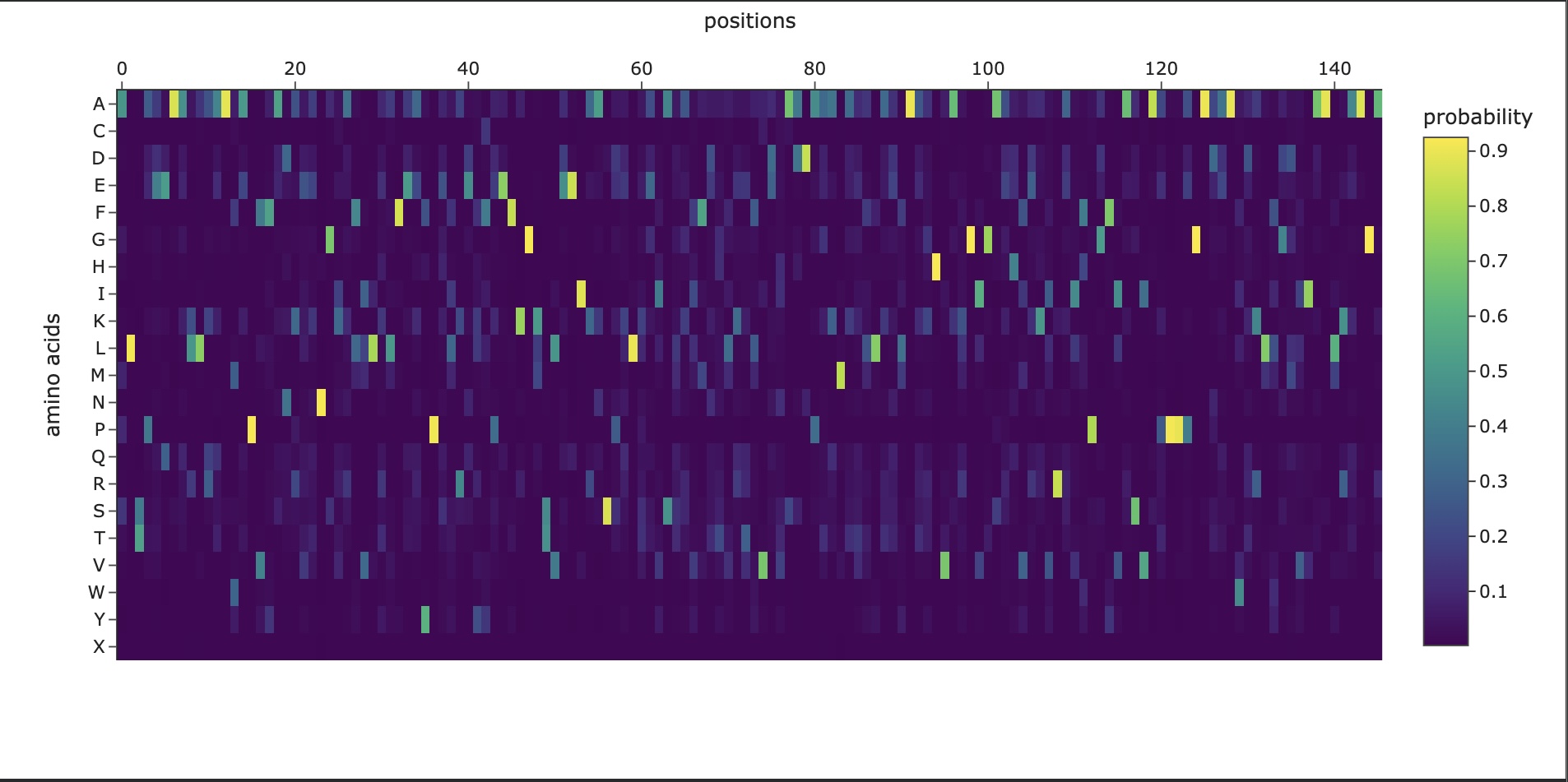
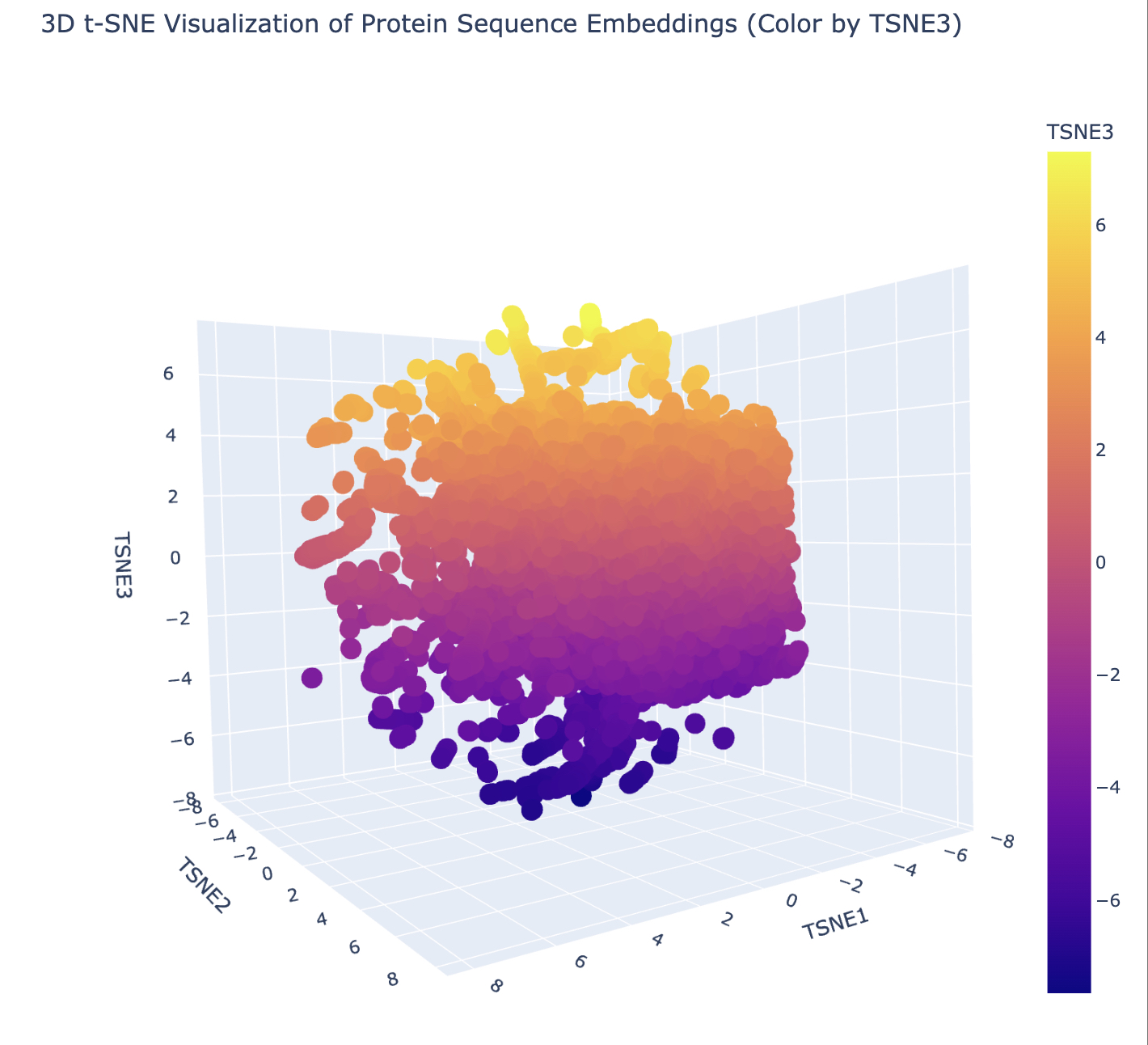
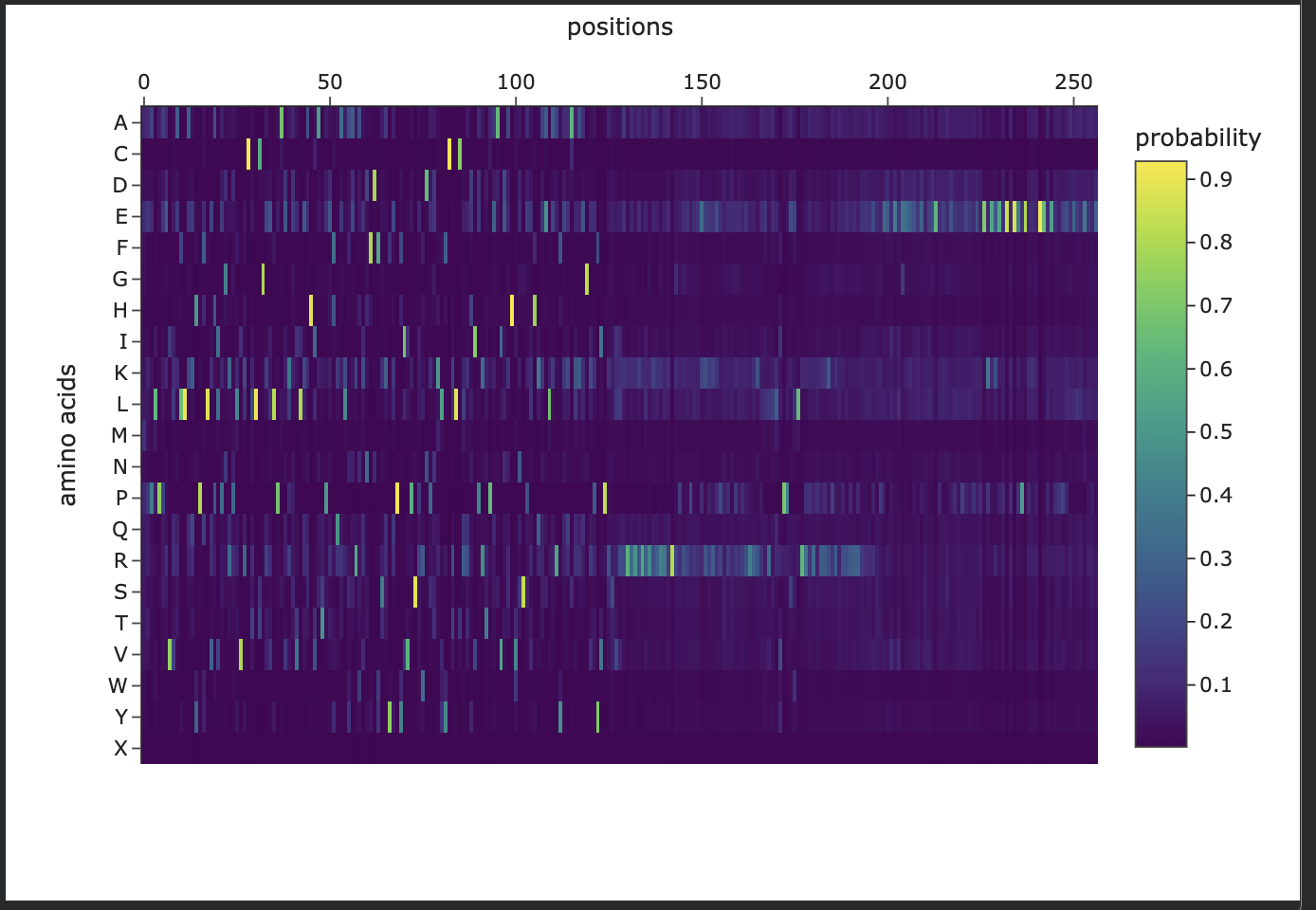
Homework: Protein Design I Assignment Objective: Learn basic concepts: amino acid structure, 3D protein visualization, and the variety of ML-based design tools. Brainstorm as a group how to apply these tools to engineer a better bacteriophage (setting the stage for the final project).
Week 5 HW: Protein Design Part ii
[] Homework — DUE BY START OF MAR 10 LECTURE Part A: SOD1 Binder Peptide Design (From Pranam) Superoxide dismutase 1 (SOD1) is a cytosolic antioxidant enzyme that converts superoxide radicals into hydrogen peroxide and oxygen. In its native state, it forms a stable homodimer and binds copper and zinc. Mechanis Mutations in SOD1 cause familial Amyotrophic Lateral Sclerosis (ALS). Among them, the A4V mutation (Alanine → Valine at residue 4) leads to one of the most aggressive forms of the disease. The mutation subtly destabilizes the N-terminus, perturbs folding energetics, and promotes toxic aggregation.
Week 6 HW: Genetic Circuits Part i
[]Homework — DUE BY START OF MAR 17 LECTURE Week 6 HW: Genetic Circuits Part 01 Assignment: DNA Assembly Protocol and Study Questions What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose? A proprietary gold standard heat-stable DNA polymerase alternative to Taq reagent synthesized and sold by Thermo Fisher Scientific. Unlike Taq which was isolated from thermophilic bacteria, Phusion emulates an archaea-based enzyme that evolved in the hydrothermal vents from extremeophile species. They function as DNA polymerases essentially in a form biomimickry with minimal replication error. The purpose of Phusion is to amplify target DNA sequences in the PCR protocol. Phusion PCR is more expensive but worth the investment to increase the accuracy of the run.
Week 7 HW: Genetic Circuits Part 2
[]Homework — DUE BY START OF MAR 31 LECTURE at 2PM ET Week 7 HW: Genetic Circuits Part 2 Assignment Part 1: Intracellular Artificial Neural Networks (IANNs) What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions? IANNs are ideal for the continuous transcriptomic-driven change observed in cells that are constantly moving and communicating in their intracellular environment – through analog computations. In contrast, much of the early synbio genetic circuit engineering was digital, with discrete logic gate switch programming or perhaps even through gene knock out (present versus absent) if such a connection would be permitted.
Week 9 HW: Cell Free Systems Assignment Part 1: ABC? EFG Reading & Resources
Week 10 HW: Imaging and Measurement
Week 10 HW: Imaging and Measurement Assignment Part 1: ABC? EFG Reading & Resources
![DNA-based digital data storage technology. Source: Archives in DNA: Workshop Exploring Implications of an Emerging Bio-Digital Technology through Design Fiction - Scientific Figure on ResearchGate. Available from: https://www.researchgate.net/figure/DNA-based-digital-data-storage-technology_fig1_353128454 [accessed 11 Feb 2025]](image4.png)