Week 4 HW: Protein Design Part I
Homework: Protein Design I
Assignment
Objective:
Learn basic concepts: amino acid structure, 3D protein visualization, and the variety of ML-based design tools. Brainstorm as a group how to apply these tools to engineer a better bacteriophage (setting the stage for the final project).
Key Links:
HTGAA Protein Engineering Tools, HTGAA Protein Engineering Feedback
Part A. Conceptual Questions
Answer any of the following questions by Shuguang Zhang:
1. How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons)
2. Why humans eat beef but do not become a cow, eat fish but do not become fish?
Although the saying goes we are what we eat, our genomes disagree. The genes preserve function against infinite intrinsic and extrinsic stresses or other organisms’ genes. Vertically inherited animal genomes from ancestors, parents, and offspring are protected in the germ line but there also exists the functional genes that create proteins in specialized tissue mosaics that also selfishly persist as long as constituent cells survive – all cells are selfish like that. These specialized cells are sustained, though by new macromolecules which is likely why the human eat the cow in the first place. Here though Natural Selection has again preserved the reproductive programs of each species genome – starting with the taste buds and olfactory centers of the human brain. A carnivore has a digestive tract that can disassemble and denature raw muscle protein very efficiently, largely because they have different enzymes. Are lions susceptible to prion diseases? That question will make more sense in a moment, but a quick search of the internet indicates they are highly resistant (particularily to Chronic Wasting Disease). Again the enzymes in their gut protect them against 97-98% of CWDs they are exposed to. The risk to humans is different, but then so is the type of prion disease. Human also have digestive tracts though that denature the proteins and lipids of the cow before it even enters the digestive tract. Unfortunately humans also have large scale industrial agriculture and rampid inequality in access to cuts and quality of cow meat. This means that economics largely determines what parts of cows humans will be exposed and how concentrated the cows on the feed lots will be along with the condition of slaughter floor. It’s hard to know today because enzymes outside of niches are fragile things, but once humans started cooking their cow meat before shredding and emulsification in their digestive systems maybe they had more of the lion’s enzymes. The loss of those enzymes is no more felt than in the epidemiological triangle between prion in cow and the prion in man, here not gene-to-gene but protein-to-protein interaction between these two animals and the zoonotic pathogen adds a new layer of complexity.
3. Why there are only 20 natural amino acids? > This is such a great question. Likely it was because Earth is a special planet, at least the most exceptional planet we have any knowledge about, thus far, for forming these 20 amino acids. Foundational to this question is what I love most about the study of Earth life. Evolution is change in matter in response to energetic landscapes in constant motion. Everything in biology is nested in this evolutionary onion through which cosmic evolution begets planetary evolution all the way down to the sub-atomic particles of atoms and back up to cells-within-cells. All this evolutionary change, as my astrobiologist friends tell me, started with the creation of our Universe 13.8 billion years ago. Therefore, as I learned from Dr. Graves to Dr. Lane, single-celled organisms evolved into multicellular organisms through Natural Selection and biochemical pathways that can be traced back from the 20 natural amino acids and other macromolecules that then formed single-celled organisms and multicelled organisms. We can trace origin of individual atoms in those micromolecules to macromolecules, one element at a time or in families. Throughout all evolutionary nesting, from the Cosmos to the Microcosmos, there is the unifying story of energy flowing and matter cycling. This is the root cause of the first tradeoff in Natural Selection – the struggle for existence of living organisms constrained by dynamic environmental conditions. Life, or the 20 natural amino acids, thus formed from elemental particles flowing from energy sink to energy sink over the vast horizon of evolutionary time in one long continuous chain reaction. If anyone doubts this, break every natural amino acid down to elemental atoms, starting with Hydrogen the most abundant element in the Universe.
6. Can you make other non-natural amino acids? Design some new amino acids. We sure can but we have to be heretical and leave the canonical 2o natural amino acids behind. The process then becomes a matter of what your trajectory will be through the process. Are you going to design new amino acids within cells or externally and then from the bottom up or top down?
7. Where did amino acids come from before enzymes that make them, and before life started. > Earth was formed from the same cosmic evolutionary process that formed the other planets and their moons over 4 billion years ago. Cells are an administrative variable today. What is less understood is the abiotic chemistry of the primordial oceans, could the hydrothermal vents Infact we know that orginally, when the Earth was first formed, it didn’t even have a moon. Then there was movement of Jupiter and Mars was forever changed and a giant piece of rock hit Earth which led to the formation of our Moon. Again, all this was over 4 billion years ago, and there were no amino acids or enzymes at this time. Now, I am going to throw a bunch of science at you, but just remember when I do – the Earth and Moon are a twin study. The simple answer is that when Amino Acids are synthesized in a lab, outside of a cell, they form a racemic mixture of both L-amino acids and D-amino acids. However, when they are synthesized within cells between genes and proteins for specific functions, they are almost exclusively L-amino acids. This is because cells use enzymes to speed up biological synthesis, and this contributes to the preponderance of L-amino acids. Samples of the moon’s surface from the Apollo missions contained glycine, alanine, glutamic acid, aspartic acid, serine, and threoine. This indicates that there was life generating amino acids on the moon, but the process stopped. Coincidentally, that over 4 billion years ago when the Earth was forming, a huge rock collided with it, leading to the formation of the orbiting Moon body. This allows us to deduce that some simple R-group amino acids left Earth and entered the vacuum of space where they essentially were frozen in time. Meanwhile, more natural L-amino acids continued to evolve on Earth. Where did they evolve? In bacterial and archaeal cells, the central dogma of molecular biology tells us that genes cannot transcribe and translate proteins without enzymes. Now knowing that ontogeny recapitulates phylogeny the question now becomes, who is the index bacterium, Earth life’s original synthesizer?
8. If you make an alpha-helix using D-amino acids, what handedness (right or left) would you expect?
The 20 primary amino acids are all L-amino acids, as are most protein building blocks of cells. Alpha-helices here will be the B-DNA, favoring right-handedness. Thus by the power of deduction that leaves D-amino acids, the exceptions, to the way of left-handedness. Now, the cool science here is that B-DNA favors right-handedness on Earth, but because of complementary strand synthesis, the pattern of right-handedness carries over to the A-form RNA during transcription, but not exclusively! This means that our converse hypothesis was correct, and there is another handedness in RNA, also known as Z-RNA. Furthermore, there are specific conditions that generate the Z-RNA form.
9. Can you discover additional helices in proteins?
Yes, there are primary, secondary, tertiary, and quaternary forms in proteins based on the structural chemistry of the bonding molecules. The ⍺-helix is the most ordinary secondary protein structure but in biology there are already exceptions. Examples of these exceptional helices include the 310 and pi helices.
10. Why most molecular helices are right-handed?
Most life on Earth is evolutionary rooted in B-DNA helices with a right-handed confirmation due to origin in saltwater oceans, passed on to self-replicating cells synthesized from macromolecules shaped by complementarities in form dominated by non-covalent weak interactions. source,
11. Why do beta-sheets tend to aggregate?
Two ways to answer this question: Hysteresis and Natural Selection. This is a why question after all, and that’s what evolution shines light on, the why behind biological structure. If you dig to the bottom of the fossil pile or record you find energetic attractor conditions for the clumping of polypeptides. Let’s start with the canonical amino acid ingredients of beta-sheet secondary structures. Every amino acid from the natural 20 is capable of contributing to beta-sheets but some are very improbable to be represented, including: Proline, Glycine, Asparatic acid, Glutamic acid, and Lysine. Conversely, some frequent amino acid suspects and properties favor beta-sheet formation. For example, the alternating chain pattern of hydrophobicity and polarity. Therefore, high probability beta-sheet producers include: Valine, Isoleucine, Phenylalanine, Serine, and Tyrosine.
12. What is the driving force for b-sheet aggregation?
Before we look at the science my speculative answer is structural stability. In nature we see the tensil strength of silk
13. Why many amyloid diseases form b-sheet?
14. Can you use amyloid b-sheets as materials?
15. Design a b-sheet motif that forms a well-ordered structure.
Part B: Protein Analysis and Visualization
In this part of the homework, you will be using online resources and 3D visualization software to answer questions about proteins.
I summon mTOR for my protein (including 4JSV, 4JSN, 4DRH, 4DRI, 3ML9, 5WBH, 5GPG, 5H64, 5FLC, 6BCX, 6BCU, 6SB0, 6SB2, 6ZWM, 7PE7, 7PEC, 8ERA, 9ED4, 9ED6, 9ED7, 9ED8).
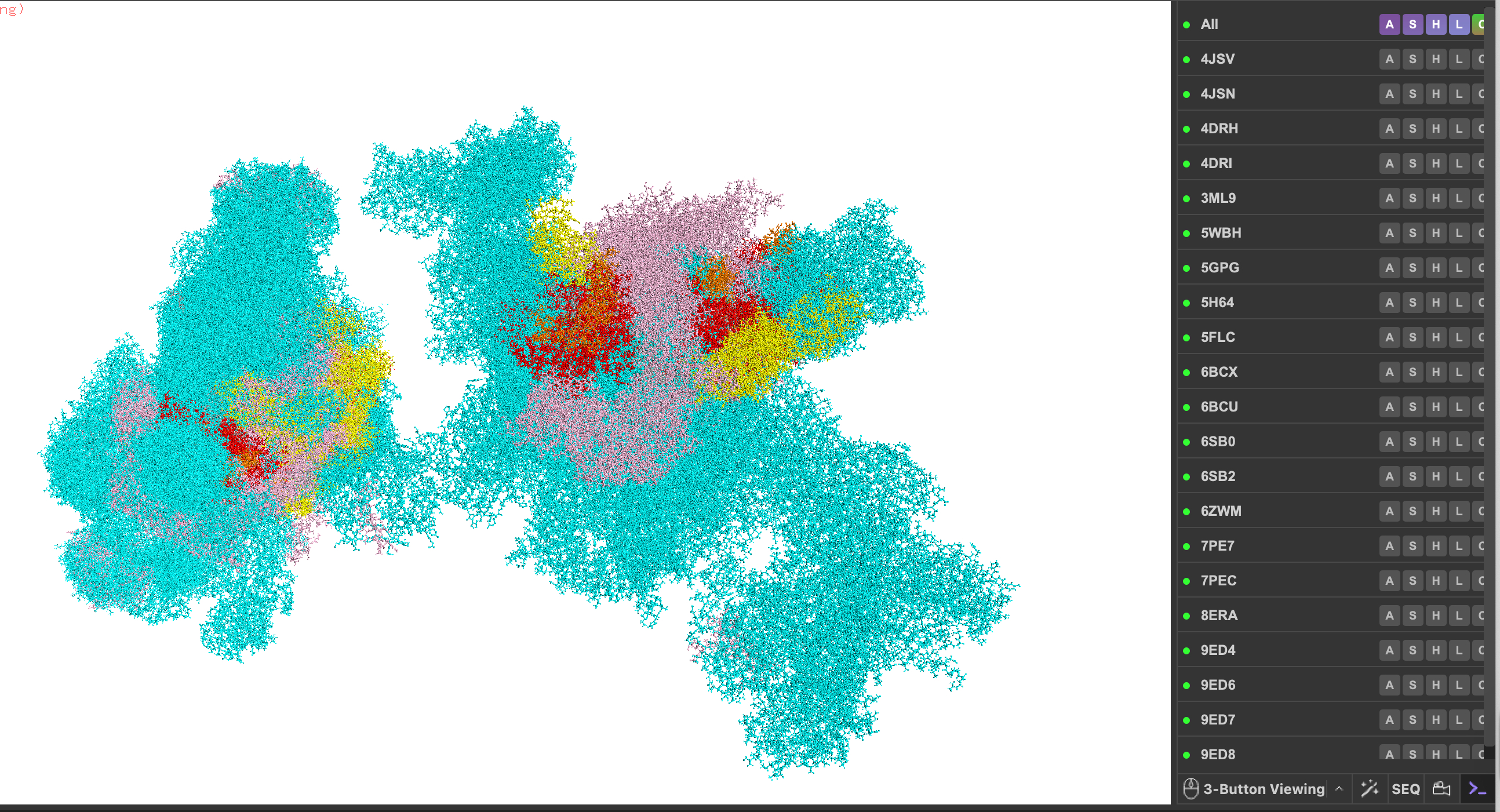


Pick any protein (from any organism) of your interest that has a 3D structure and answer the following questions.
- Briefly describe the protein you selected and why you selected it. The protein I am now selecting now that I better understand the exercise and tooling is the mighty mTOR TOR mechanistic target of Rapamycin protein kinase. I pick this protein because just trying to write the name is challenging. In add there is conserved and then there is mTOR conserved, it’s as common as Cytochrome C. Also I like the puzzle of navigating all of the codes just to plug it into the codebase – the combinations are amazing as my son’s Pokomon characters would say. Lastly, it has a lot of wiring for considering circuitry as a quintessential regulator of Natural Selection programs for growth, maintenance, and metabolism under stress.
- Identify the amino acid sequence of your protein.
| PDB_A | PDB_B | Protein AA Sequence |
|---|---|---|
| 4JSV | 4JSV_2 Target of rapamycin complex subunit LST8 | MNTSPGTVGSDPVILATAGYDHTVRFWQAHSGICTRTVQHQDSQVNALEVTPDRSMIAAAGYQHIRMYDLNSNNPNPIISYDGVNKNIASVGFHEDGRWMYTGGEDCTARIWDLRSRNLQCQRIFQVNAPINCVCLHPNQAELIVGDQSGAIHIWDLKTDHNEQLIPEPEVSITSAHIDPDASYMAAVNSTGNCYVWNLTGGIGDEVTQLIPKTKIPAHTRYALQCRFSPDSTLLATCSADQTCKIWRTSNFSLMTELSIKSGNPGESSRGWMWGCAFSGDSQYIVTASSDNLARLWCVETGEIKREYGGHQKAVVCLAFNDSVLG |
| 4JSV | 4JSV_1 Serine/threonine-protein kinase mTOR | DDNGIVLLGERAAKCRAYAKALHYKELEFQKGPTPAILESLISINNKLQQPEAAAGVLEYAMKHFGELEIQATWYEKLHEWEDALVAYDKKMDTNKDDPELMLGRMRCLEALGEWGQLHQQCCEKWTLVNDETQAKMARMAAAAAWGLGQWDSMEEYTCMIPRDTHDGAFYRAVLALHQDLFSLAQQCIDKARDLLDAELTAMAGESYSRAYGAMVSCHMLSELEEVIQYKLVPERREIIRQIWWERLQGCQRIVEDWQKILMVRSLVVSPHEDMRTWLKYASLCGKSGRLALAHKTLVLLLGVDPSRQLDHPLPTVHPQVTYAYMKNMWKSARKIDAFQHMQHFVQTMQQQAQHAIATEDQQHKQELHKLMARCFLKLGEWQLNLQGINESTIPKVLQYYSAATEHDRSWYKAWHAWAVMNFEAVLHYKHQNQARDEKKKLRHASGANITNATTAATTAATATTTASTEGSNSESEAESTENSPTPSPLQKKVTEDLSKTLLMYTVPAVQGFFRSISLSRGNNLQDTLRVLTLWFDYGHWPDVNEALVEGVKAIQIDTWLQVIPQLIARIDTPRPLVGRLIHQLLTDIGRYHPQALIYPLTVASKSTTTARHNAANKILKNMCEHSNTLVQQAMMVSEELIRVAILWHEMWHEGLEEASRLYFGERNVKGMFEVLEPLHAMMERGPQTLKETSFNQAYGRDLMEAQEWCRKYMKSGNVKDLTQAWDLYYHVFRRISKQLPQLTSLELQYVSPKLLMCRDLELAVPGTYDPNQPIIRIQSIAPSLQVITSKQRPRKLTLMGSNGHEFVFLLKGHEDLRQDERVMQLFGLVNTLLANDPTSLRKNLSIQRYAVIPLSTNSGLIGWVPHCDTLHALIRDYREKKKILLNIEHRIMLRMAPDYDHLTLMQKVEVFEHAVNNTAGDDLAKLLWLKSPSSEVWFDRRTNYTRSLAVMSMVGYILGLGDRHPSNLMLDRLSGKILHIDFGDCFEVAMTREKFPEKIPFRLTRMLTNAMEVTGLDGNYRITCHTVMEVLREHKDSVMAVLEAFVYDPLLNWRLMDTNTKGNKRSRTRTDSYSAGQSVEILDGVELGEPAHKKTGTTVPESIHSFIGDGLVKPEALNKKAIQIINRVRDKLTGRDFSHDDTLDVPTQVELLIKQATSHENLCQCYIGWCPFW |
| 4JSN | ||
| 4DRH | ||
| 4DRI | ||
| 3ML9 | ||
| 5WBH | ||
| 5GPG | ||
| 5H64 | ||
| 5FLC | ||
| 6BCX | ||
| 6BCU | ||
| 6SB0 | ||
| 6SB2 | ||
| 6ZWM | ||
| 7PE7 | ||
| 7PEC | ||
| 8ERA | ||
| 9ED4 | ||
| 9ED6 | ||
| 9ED7 | ||
| 9ED8 |
How long is it? What is the most frequent amino acid? You can use this notebook to count most frequent amino acid - https://colab.research.google.com/drive/1vlAU_Y84lb04e4Nnaf1axU8nQA6_QBP1?usp=sharing
The length of the Surfeit locus protein 2 protein is: 257 amino acids. The most common amino acid is: L, which appears 27 times. All Amino Acid Frequencies: L: 27 (10.51%) K: 25 (9.73%) E: 24 (9.34%) R: 24 (9.34%) S: 20 (7.78%) P: 18 (7.00%) D: 15 (5.84%) Q: 15 (5.84%) T: 14 (5.45%) H: 13 (5.06%) F: 12 (4.67%) G: 10 (3.89%) A: 8 (3.11%) Y: 8 (3.11%) V: 7 (2.72%) N: 7 (2.72%) C: 5 (1.95%) M: 2 (0.78%) I: 2 (0.78%) W: 1 (0.39%)
How many protein sequence homologs are there for your protein? Hint: Use the pBLAST tool to search for homologs and ClustalOmega to align and visualize them. Tutorial Here
Does your protein belong to any protein family?
Identify the structure page of your protein in RCSB
When was the structure solved? Is it a good quality structure? Good quality structure is the one with good resolution. Smaller the better (Resolution: 2.70 Å)
Are there any other molecules in the solved structure apart from protein?
Does your protein belong to any structure classification family? SURF2
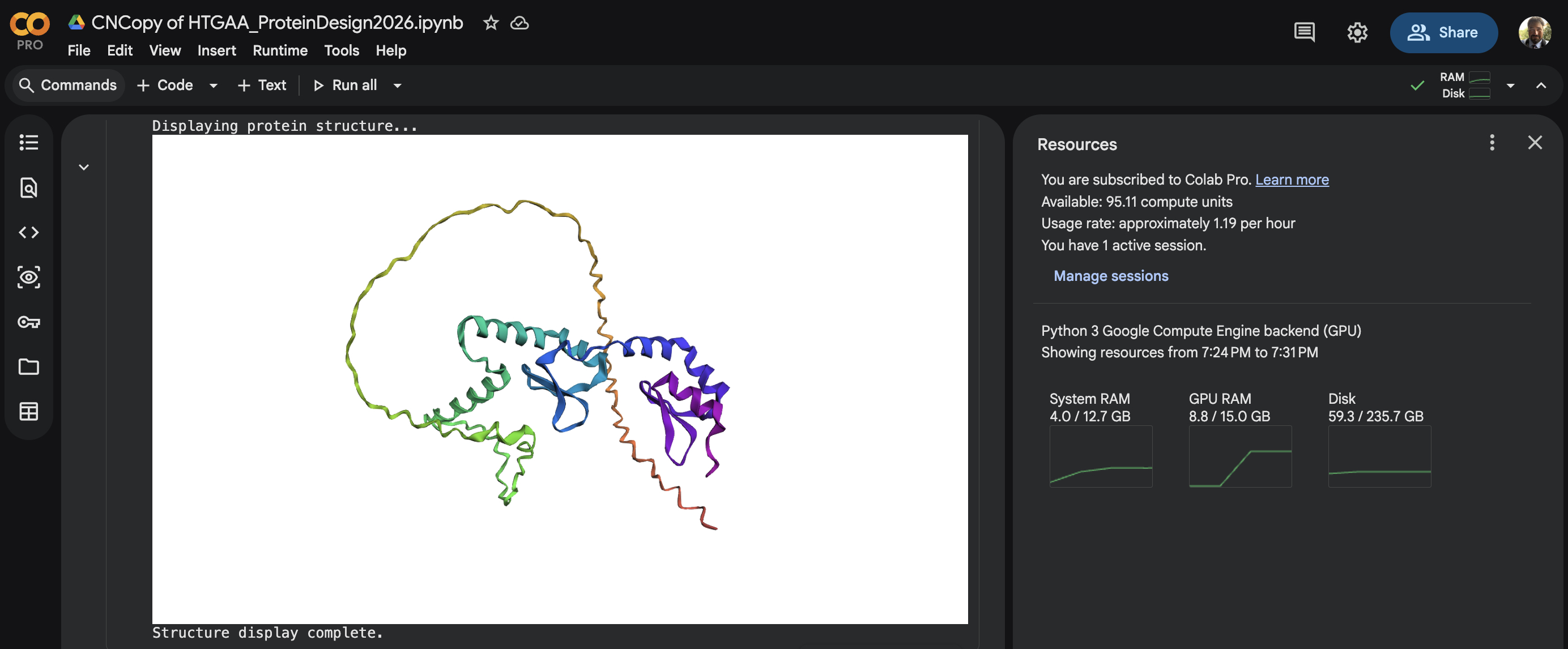
Open the structure of your protein in any 3D molecule visualization software:
- PyMol Tutorial Here (hint: ChatGPT is good at PyMol commands)
- Visualize the protein as “cartoon”, “ribbon” and “ball and stick”.
- Color the protein by secondary structure. Does it have more helices or sheets?
- Color the protein by residue type. What can you tell about the distribution of hydrophobic vs hydrophilic residues?
- Visualize the surface of the protein. Does it have any “holes” (aka binding pockets)?
Part C. Using ML-Based Protein Design Tools
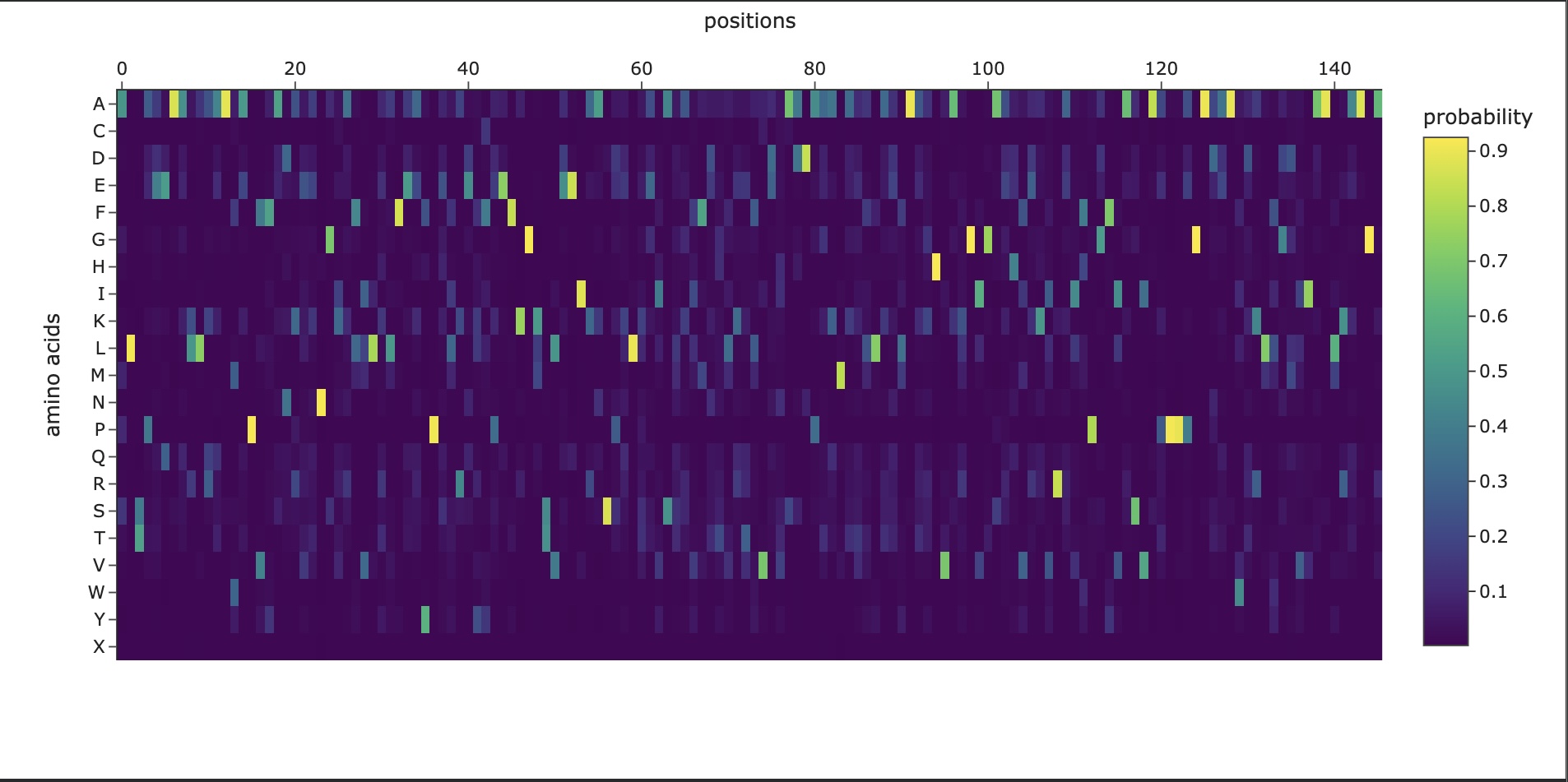
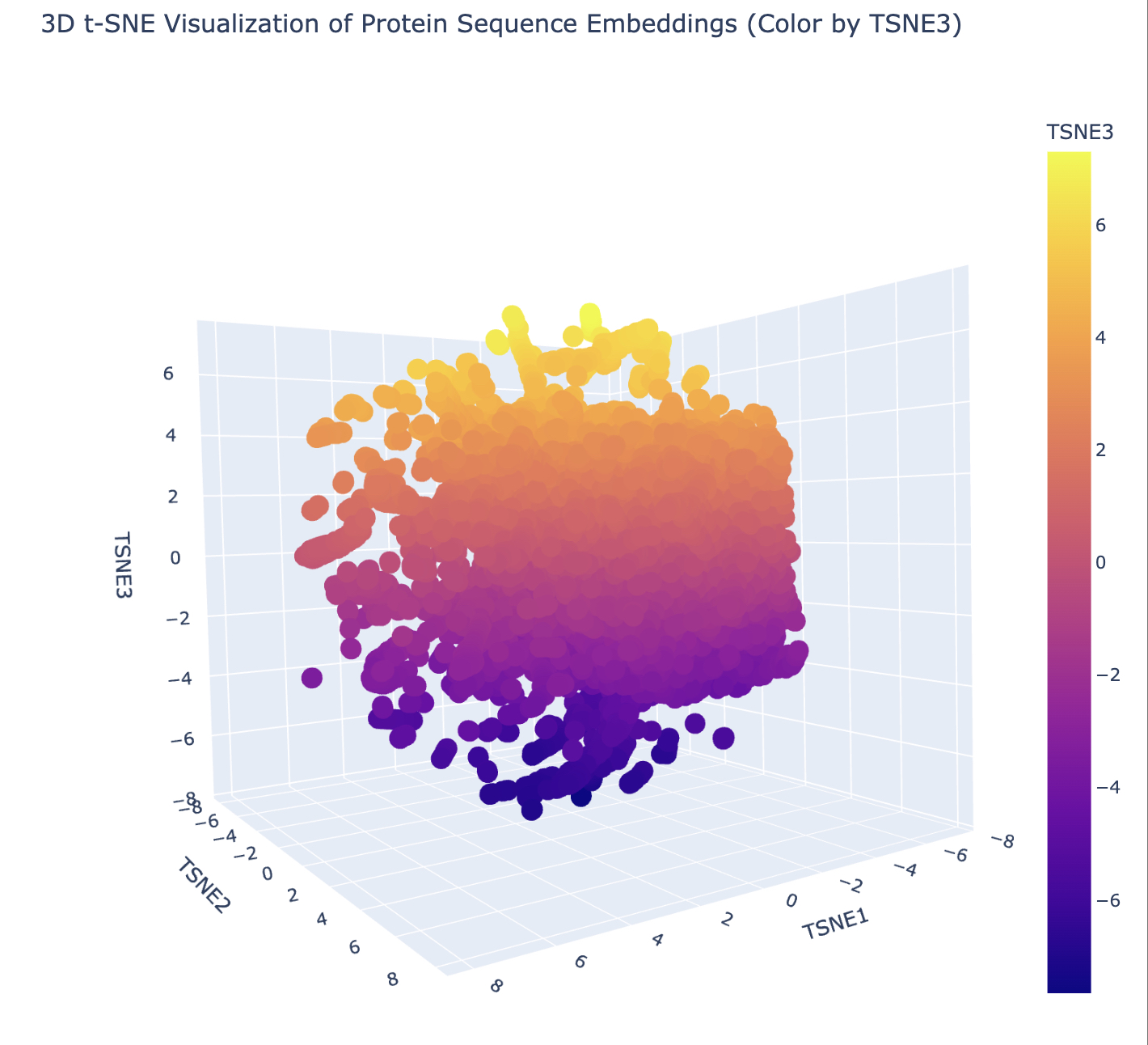
In this section, we will learn about the capabilities of modern protein AI models and test some of them in your chosen protein. Copy the notebook below and set up a colab instance with GPU for this section: HTGAA_ProteinDesign2026.ipynb Choose your favorite protein from the PDB. We will now try multiple things, report each of those results in your homework page: Protein Language Models: Deep Mutational Scans Use ESM2 to generate an unsupervised deep mutational scan of your protein based on language model likelihoods. Can you explain any particular pattern? (choose a residue and a mutation that stands out) (Bonus) Find sequences for which we have experimental scans, and compare the prediction of the language model to experiment Latent Space Analysis Use the provided sequence dataset to embed proteins in reduced dimensionality Analyze the different formed neighborhoods: do they approximate similar proteins? Place your protein in the resulting map and explain its position and similarity to its neighbors Attention Maps Analyze the attention maps of ESM2. Investigate if its layers correlate to the 2D map of residue distances of your protein Protein Folding: Folding a protein Fold your protein with ESMFold. Do the predicted coordinates match your original structure? Try changing the sequence, first try some mutations, then large segments. Is your protein structure resilient to mutations? Protein Generation: Inverse-Folding a protein Let’s now use the backbone of your chosen PDB to propose sequence candidates via ProteinMPNN Analyze the predicted sequence probabilities and compare the predicted sequence vs the original one Input this sequence into ESMFold and compare the predicted structure to your original
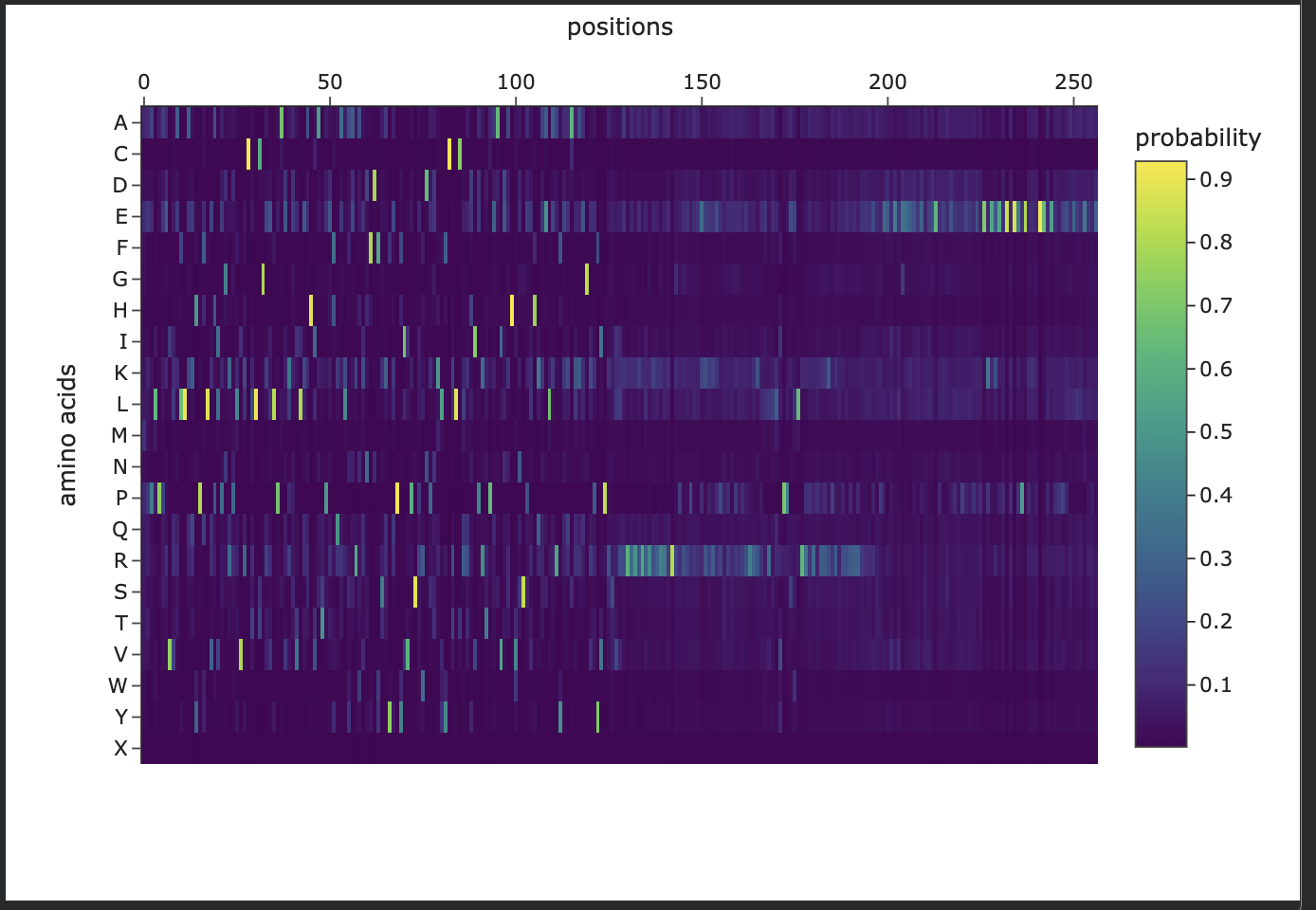


Last step in my copy of HTGAA_ProteinDesign2026.ipynb script
Generating sequences… tmp, score=2.2338, fixed_chains=[], designed_chains=[‘A’], model_name=v_48_020 MDEPPSDVLAFLRQHPSLRLLPNTRKVRCSLTGHELPCRLPELQEYTRGKKYQRLSSSFSNFDYAAFEPHIVPSTKNRHQLFCKLTLRHINKSPEHVLRHTQGRRYQRALHQYEECQKQGVEYVPACLLHKRKKREDQTNSDELPGQRTGFWEPASSDEEDALSDDSMTDLYPPELFTKRELGKPKNDDTPEDFLTDQQDEKPEHSEEKSFREREEARVGHKRGRKLRKKQLTSLTKKFKSYHHKPKNFSSFKQLGR T=0.1, sample=0, score=1.1620, seq_recovery=0.2568 MPPLPPEVVAFLAQHPHLVALPGQPLVRCTLTGEELPAELPVLRAHVATPRHQALAAREKNFDFSKYEPHIVPSRWDPDKLFCRLCLKEIPKTPEAVEAHVNSKEHQEALKEYEEAKKRGKRYIPKRLRKRRRRRRRRRRRRRGRRKRRKRRPPPPRRRPRRKRRRRRRRLVPREWLRRRRRRRRRRRRRRRPRRRRRPPRRVVGAAPEPAVAALAEAPAPPAPPAPPPPPEERPEPPPPPPERREPPPEELEEEEE
New Sequence:MPPLPPEVVAFLAQHPHLVALPGQPLVRCTLTGEELPAELPVLRAHVATPRHQALAAREKNFDFSKYEPHIVPSRWDPDKLFCRLCLKEIPKTPEAVEAHVNSKEHQEALKEYEEAKKRGKRYIPKRLRKRRRRRRRRRRRRRGRRKRRKRRPPPPRRRPRRKRRRRRRRLVPREWLRRRRRRRRRRRRRRRPRRRRRPPRRVVGAAPEPAVAALAEAPAPPAPPAPPPPPEERPEPPPPPPERREPPPEELEEEEE
Part D. Group Brainstorm on Bacteriophage Engineering
Find a group of ~3–4 students Review the Bacteriophage Final Project Goals: Increased stability (easiest) Higher titers (medium) Higher toxicity of lysis protein (hard) Brainstorm Session Choose one or two main goals from the list that you think you can address computationally (e.g., “We’ll try to stabilize the lysis protein,” or “We’ll attempt to disrupt its interaction with E. coli DnaJ.”). Write a 1-page proposal (bullet points or short paragraphs) describing: Which tools/approaches from recitation you propose using (e.g., “Use Protein Language Models to do in silico mutagenesis, then AlphaFold-Multimer to check complexes.”). Why you think those tools might help solve your chosen sub-problem. One or two potential pitfalls (e.g., “We lack enough training data on phage–bacteria interactions.”). Include a schematic of your pipeline This resource may be useful: HTGAA Protein Engineering Tools Individually put your plan on your website page Each group’s short plan for engineering a bacteriophage Schedule time ( HTGAA Protein Engineering Feedback) to get feedback/discuss your ideas, and put the feedback on your website [Optional] Part E. Find a drug for an oncology target