Week 7 HW: Genetic Circuits Part 2Assignment Part 1: Intracellular Artificial Neural Networks (IANNs)What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions?
IANNs are ideal for the continuous transcriptomic-driven change observed in cells that are constantly moving and communicating in their intracellular environment – through analog computations. In contrast, much of the early synbio genetic circuit engineering was digital, with discrete logic gate switch programming or perhaps even through gene knock out (present versus absent) if such a connection would be permitted.
Describe a useful application for an IANN; include a detailed description of input/output behavior, as well as any limitations an IANN might face to achieve your goal.
Perhaps it’s still a bit conceptual at this time, but the redox lesion-boundary in my personal HTGAA project may be a useful application for framing an intracellular artificial neural network (IANN). My goal would be to create a perceptron-esque intracellular circuit used to classify tissue sample sites by integrating continuous biochemical inputs into a weighted threshold graded output that identifies lesion boundaries in host animals. This data will be dependent on a time-series with continuous expression and variation that is spatially distributed across host goats. The input layer in this model might not even include a mite or mite eggs as is traditionally used but instead models inputs like ROS, hypoxia, inflamatory damage that spreads across hosts in similar phenotypic patterns. The weights for this model would be the expected promoter strength, affinity, repression, similarity. Activation would be evaluated using threshold nonlinear gene response (it’s biology afterall) and the output would be fluorescent markeing (highlighting) of lesion bounardy cellular expression.
Below is a diagram depicting an intracellular single-layer perceptron where the X1 input is DNA encoding for the Csy4 endoribonuclease and the X2 input is DNA encoding for a fluorescent protein output whose mRNA is regulated by Csy4. Tx: transcription; Tl: translation.
Example Circuit Draw a diagram for an intracellular multilayer perceptron where layer 1 outputs an endoribonuclease that regulates a fluorescent protein output in layer 2.
With a minimal understanding of this week’s lab, I am now going to try to adapt the Cortassa lab’s (2004) computational mitochondrial oscillator on the reactive oxygen species model to the diagram challenge for the intracellular multilayer perceptron challenge. Why? I am endlessly fascinated by both. The Cortassa model hypotheses target spontaneous metabolic oscillations in heart cells, and is built using sound biology and stone-cold math. I also respect that Cortassa was modelling the mitochondria in 2003; we didn’t know anything about mitochondria back then, compared to what we know now. Another great feature of the Cortassa model is the scalability across time. Often, when we discuss the interstellar nature of scale in biology, we focus on space, but time is just as perplexing when it comes to the bioenergetics of metabolic redox reactions, as Cortassa et al (2004) describe, with timescales from milliseconds to hours. Lastly, the Cortassa model even considers Fluorescent probes, so as a side note, if this lab is looking for a PostDoc.
Cortassa Lab Dynamic Mitochondrial Oscillator Model
The dynamics are nonlinear
Each modelled mitochondrion is assumed to possess an inner membrane anion channel abbreviated as IMAC.
An IMAC is activated
An IMAC is modulated by MG2+ and PH
An IMAC is inhibited by amphiphilic molecules.
The model has two state changes:
There is the relaxation mitochondrial oscillator state with slow and fast spaces. Over the slow space ROS builds up in the mitochondrial matrix
There is the stable mitochondrial oscillatory state
Source:
Cortassa S, Aon MA, Winslow RL, O’Rourke B. A mitochondrial oscillator dependent on reactive oxygen species. Biophys J. 2004 Sep;87(3):2060-73. doi: 10.1529/biophysj.104.041749. PMID: 15345581; PMCID: PMC1304608.
Cortassa S, Aon MA, Marbán E, Winslow RL, O’Rourke B. An integrated model of cardiac mitochondrial energy metabolism and calcium dynamics. Biophys J. 2003 Apr;84(4):2734-55. doi: 10.1016/S0006-3495(03)75079-6. PMID: 12668482; PMCID: PMC1201507.
Assignment Part 2: Fungal MaterialsWhat are some examples of existing fungal materials and what are they used for?
What are their advantages and disadvantages over their traditional counterparts?
What might you want to genetically engineer fungi to do and why?
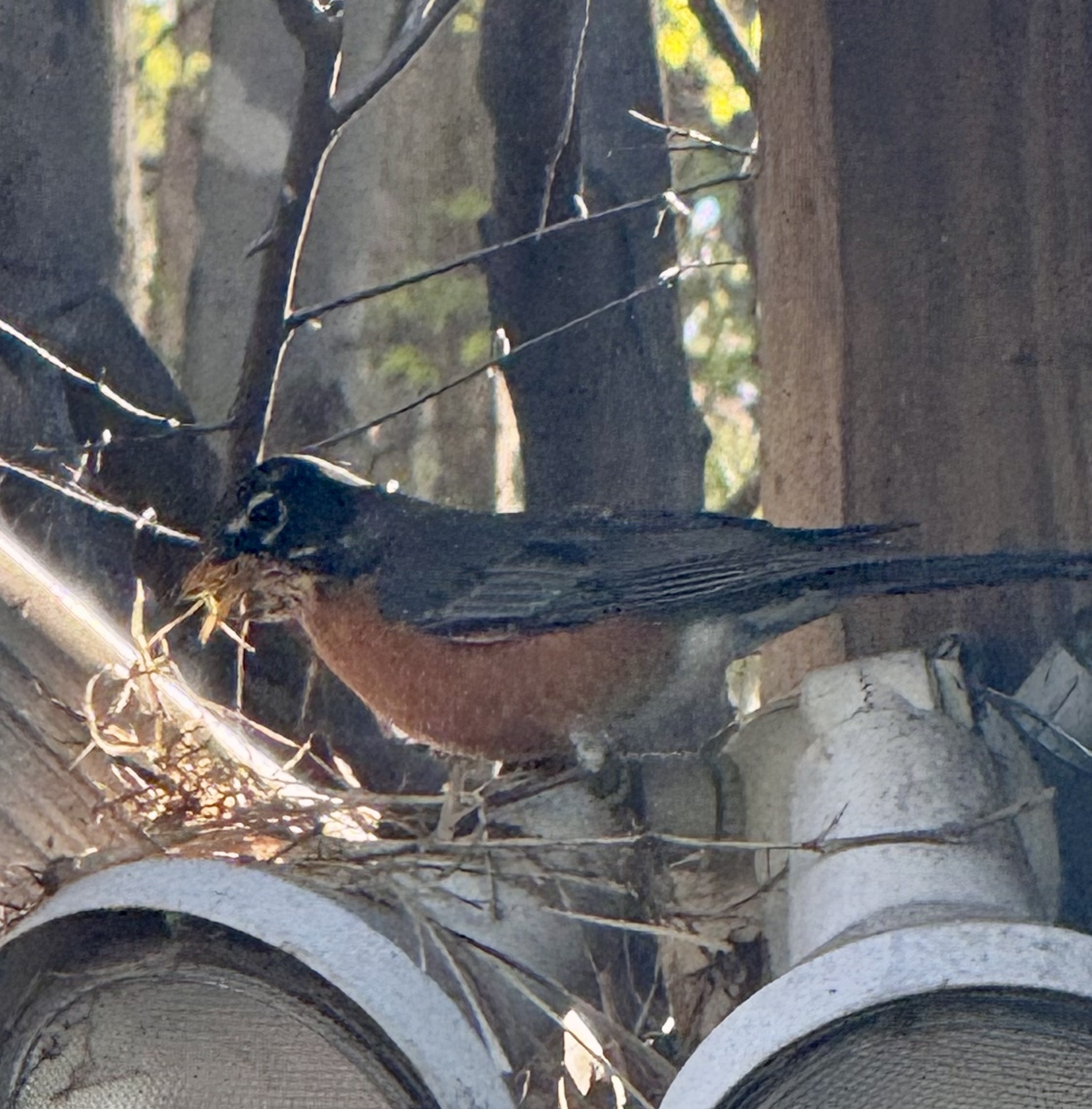
There are so many things that we could genetically engineer with Fungi, so why is it so confounding just to pick one? Ironically, it’s easier to imagine what a bird would genetically engineer with fungi. Shelter is the first demand after all for survival; a bird’s nest could be a noble odyssey for scientific discovery. To start, what do birds build nests with? On this farm, birds use sticks, hay, and straight wool. What do all of these substrates have in common? For the most part, they are straight and bendable, but there are other notable attributes. Hay and Wool for example, are resistant to most environmental antagonists: water, wind, bugs, and fire. I also think they use composites because they are structurally reliable with a predictable function. For example, all three are tough enough and not heavy. A bird can pick up a piece with its beak, fly back to the nest, and drop it where they want it. Does this make it a tool? Anyway, once they drop a piece of stick, wool, or hay they then just have two more steps: tuck in side A and then tuck in side B – done. The problem with all three of these materials is that they exist independently of the task of providing birds with material for nests. Enter a fungi farm for bird nest materials. All we need are yeast plasmids, a PCR machine, a gel electrophoresis device, reagents, electricity, computers, and revolutionary institutions to teach us how. Fortified with our why and how, we commence…
What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
There are several ways to approach this question. We choose bacteria because of their remarkable ability to adapt. While it’s important to acknowledge the bacterium’s collaboration with natural selection, it took over a billion years for the kingdom to develop this trait. Traditionally, we prioritize bacteria in synthetic biology because bacteria occupy the niche first. Why is this significant? Well, it can be summarized in one sentence: the niche is where matter and energy intersect. Fungi eventually occupy the niche also and when they do, they provide structure and in a less specific way. Perhaps their universality is because fungi possess the genes of plants and animals. Scale, is another reason Fungi. Bacteria leaned millions of years ago to the illusion of invisibility to multicellular ocular adaptations, which was advantageous to their race for niches against multicellular competitors. Not to downplay humans too much, we figured this out too, and we can even watch bacteria at their little spinning wheels and transform them from the inside out. However, be it tangential or not, I still wonder where the evolutionary studies are following the synbio bacteria prospectively as they reintegrate with nature. Therefore, in summary, fungi over bacteria because they replicate so slowly for microbiota, which for example better adjusts to human chronological understanding of space and time.
Installation Guide and Walk-through of the Neuromorphic Wizard Software.
Open link for the installation file: https://drive.google.com/drive/folders/10_gEzYV2J5hVOdKt6sNBeSH8cEMMGX8O
Click the three dots.
Click Download.
Move the folder to somewhere you can access it from Terminal with cd command.
If the downloaded file is zipped, then unzip it.
In terminal run cd into NeuromorphicWizard and ls contents
In terminal run conda create -n neuro_wiz python==3.10
In terminal run conda activate neuro_wiz
In terminal run pip install -r requirements.txt
In terminal run python main.py
Assignment Part 3: First DNA Twist OrderReview the Individual Final Project documentation guidelines.Submit this Google Form with your draft Aim 1, final project summary, HTGAA industry council selections, and shared folder for DNA designs. DUE MARCH 20 FOR MIT/HARVARD/WELLESLEY STUDENTS
Review Part 3: DNA Design Challenge of the week 2 homework. Design at least 1 insert sequence and place it into the Benchling/Kernel/Other folder you shared in the Google Form above. Document the backbone vector it will be synthesized in on your website. Reading & ResourcesThe perceptron, the basis of artificial neural networks: https://www.geeksforgeeks.org/deep-learning/what-is-perceptron-the-simplest-artificial-neural-network/
Many examples of artificial neural networks made using biomolecules: https://doi.org/10.1016/j.biosystems.2024.105164
|