Week 2 HW: DNA Read, Write, Edit Life
Part 1: Benchling & In-silico Gel Art
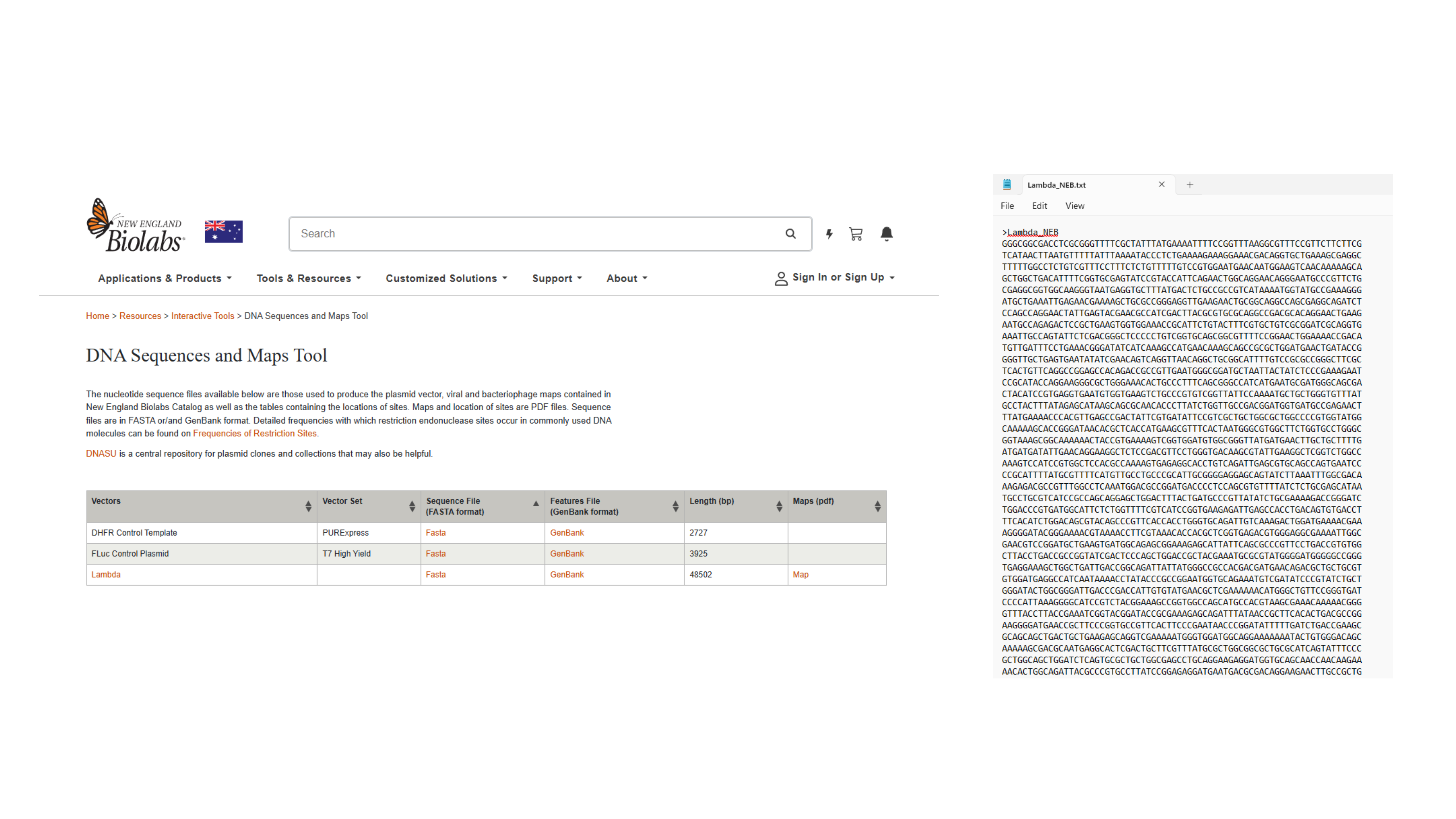

Navigating to Lambda sequence and saving as text. file
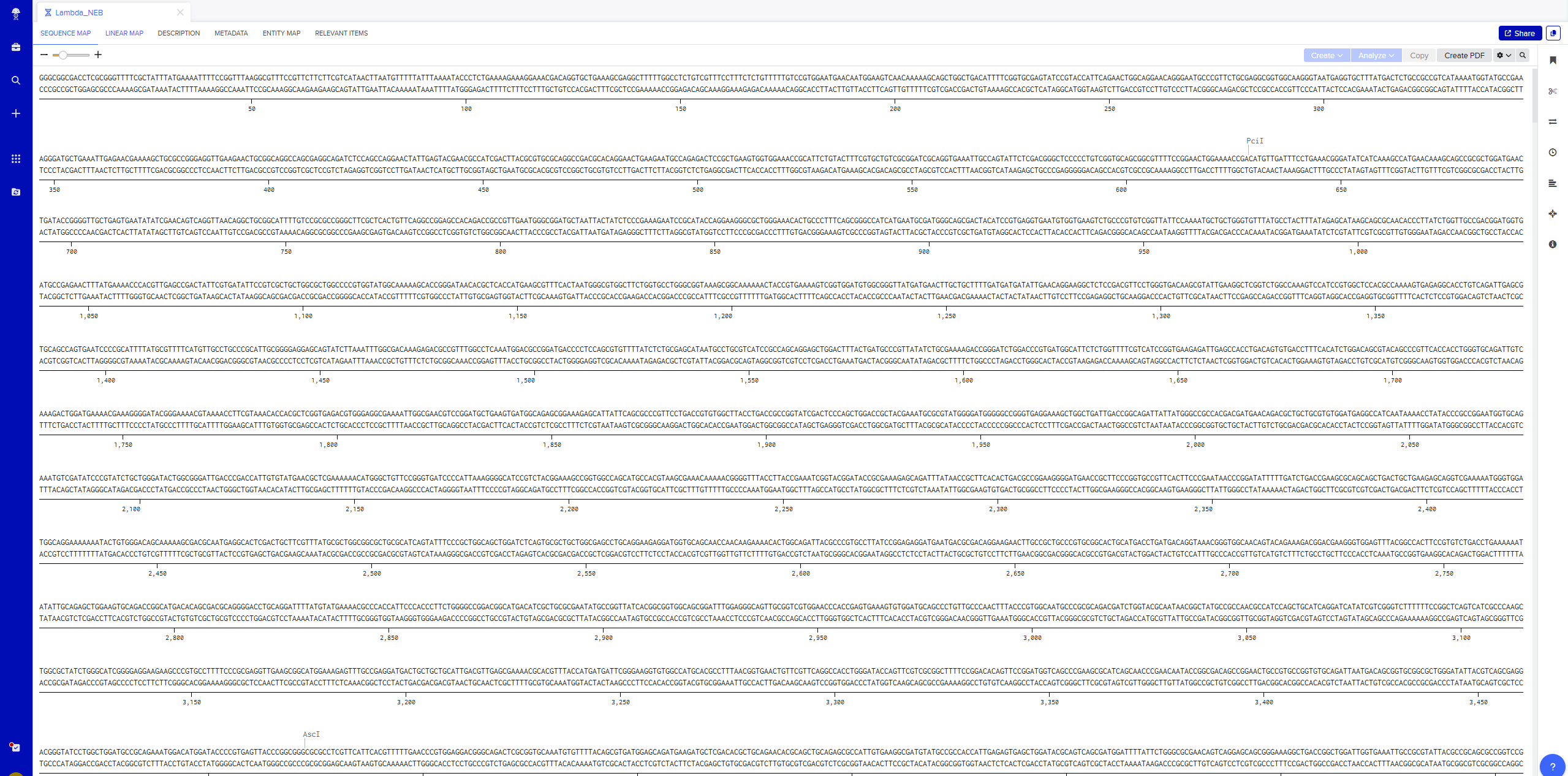


Importing Lambda file by saving FASTA Format of Sequence file
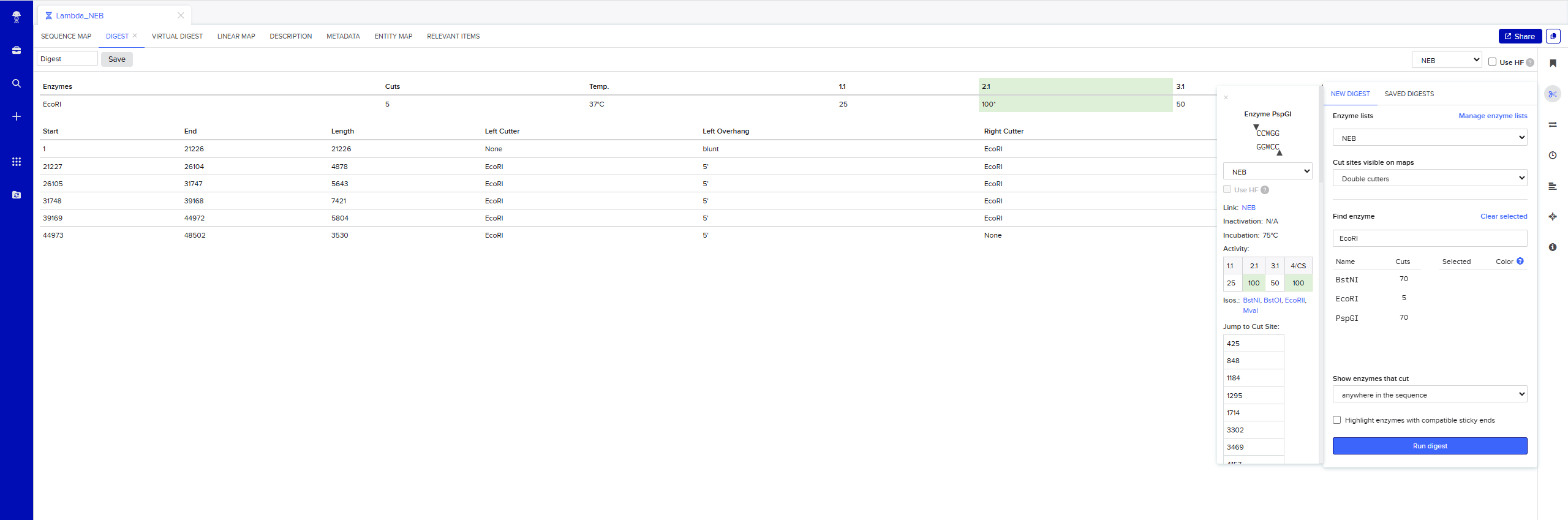

Simulate Restriction Enzyme Digestion with the following Enzymes: EcoRI
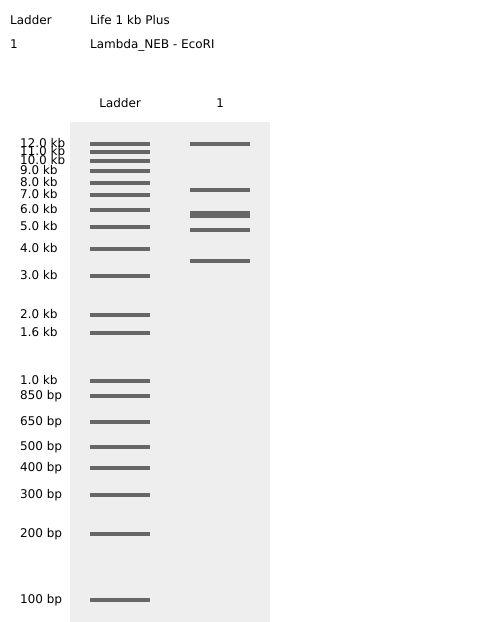

Testing Virtual digest example
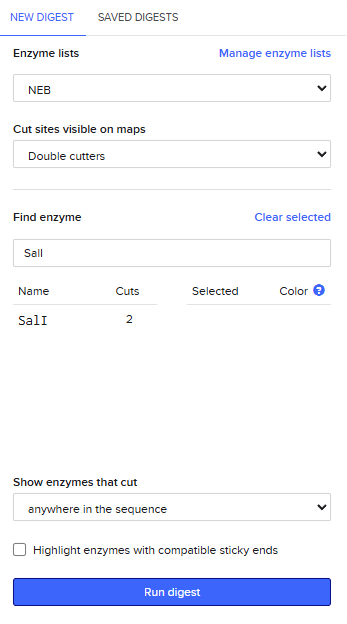

Adding more restrcition enzymes
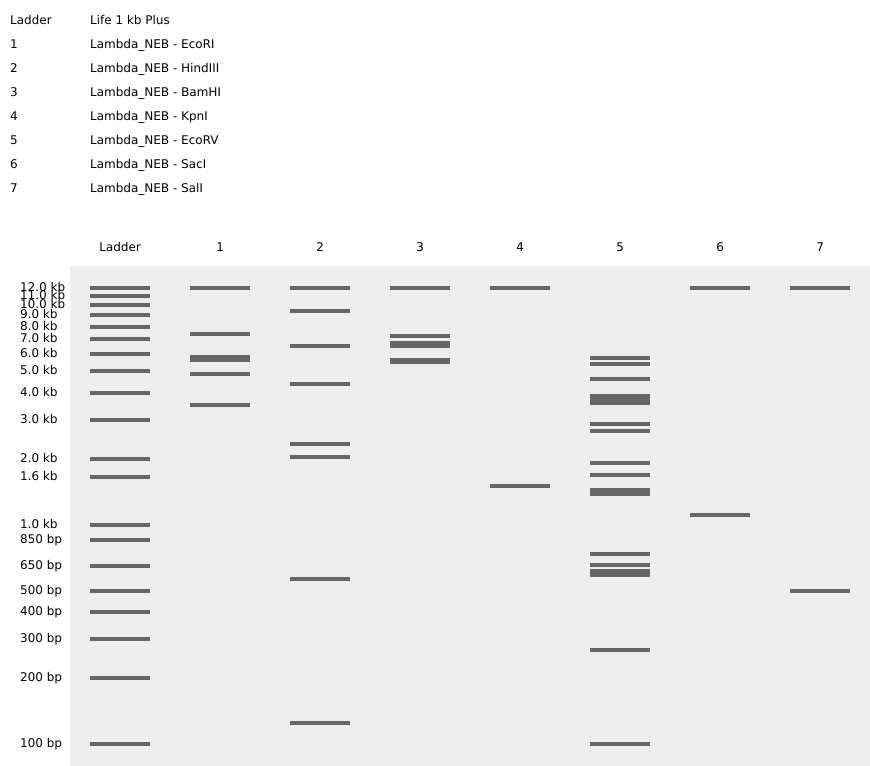


Simulate Restriction Enzyme Digestion with all the Enzymes
Create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks
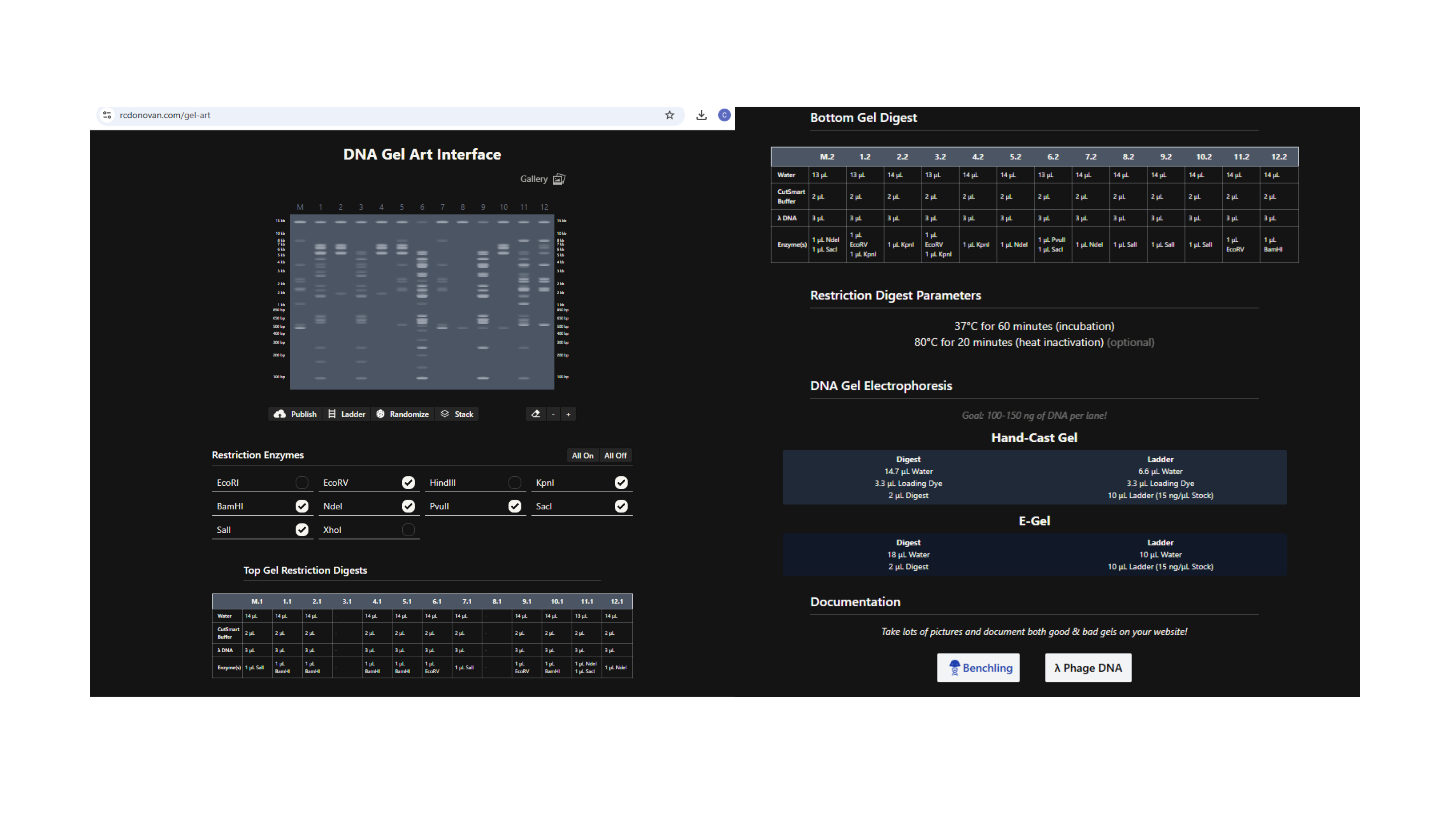

Using https://rcdonovan.com/gel-art to create randomised Gel Art
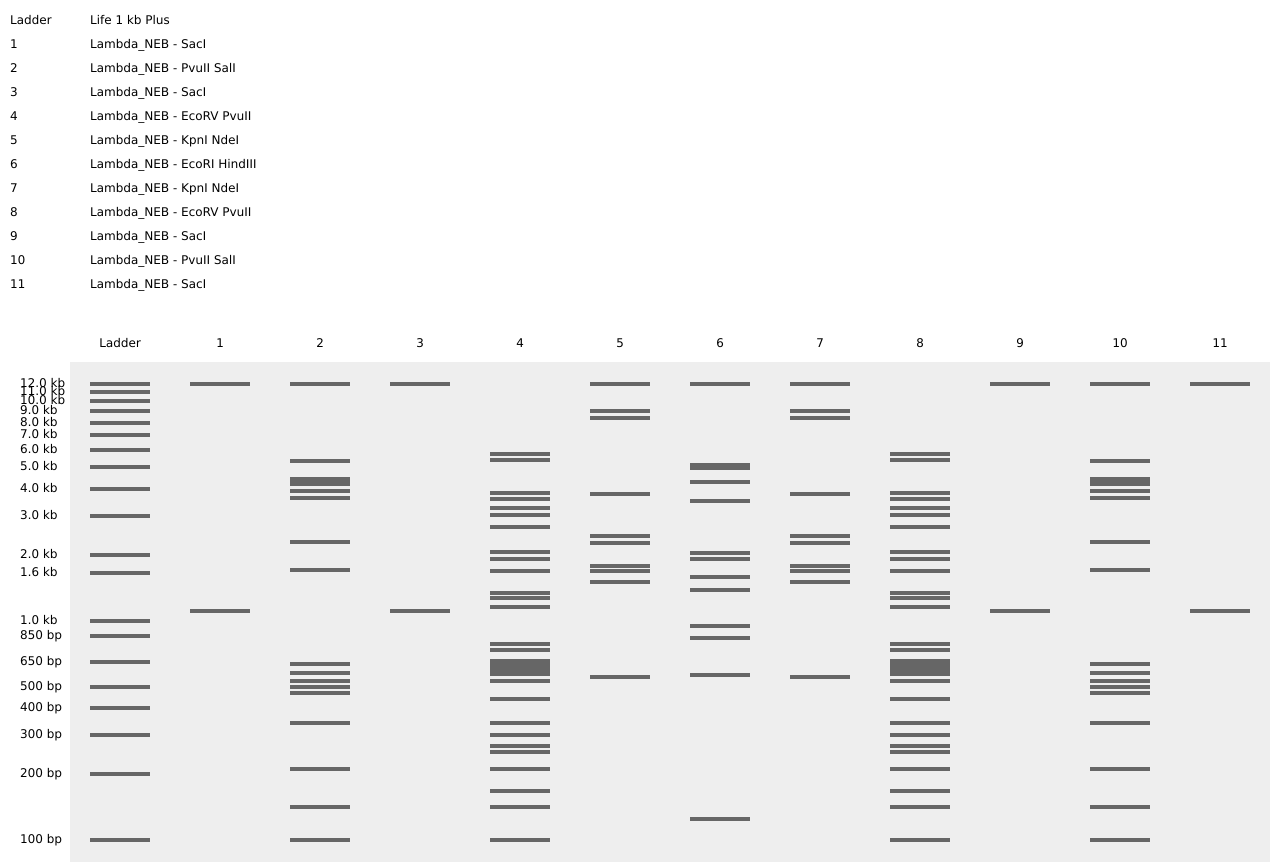
 Fireworks and Flower Gel Art Shape
Fireworks and Flower Gel Art Shape
Part 3: DNA Design Challenge 3.1 Choose your Protein
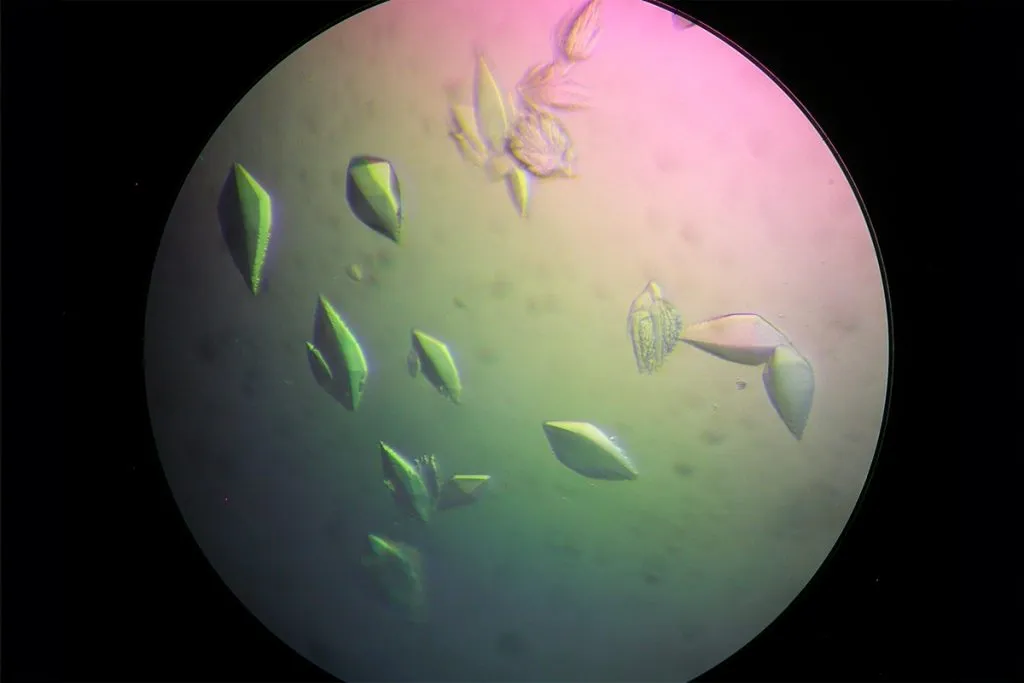

Figure 1
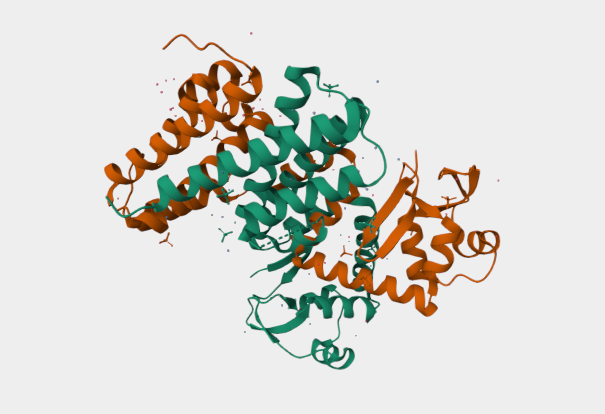

Figure 2
My chosen protein is DdrC Protein seen in figure 1 and 2 above, because of its ability to respond to DNA damage by putting a stop to the damage and alerting the cell to begin repair process. For my final project I would like to research into material that is able to self-repair and contain the deterioation. Hence I wanted to research about how DdrC from Deinococcus radioduran is able to stabilise DNA breaks by binding to both single and double stranded leisions. Something that I thought was quite interesting through the readings of this protein is its structural asymmetry allowing it precise lesion recognition (“The Unique Mechanism of DdrC in Enhancing DNA Stability,” 2024). It bears two asymmetric DNA binding sites located on either side of the dimer and can modulate the topology and level of compaction of circular DNA (Gueguen et al., 2022).
Protein Sequence
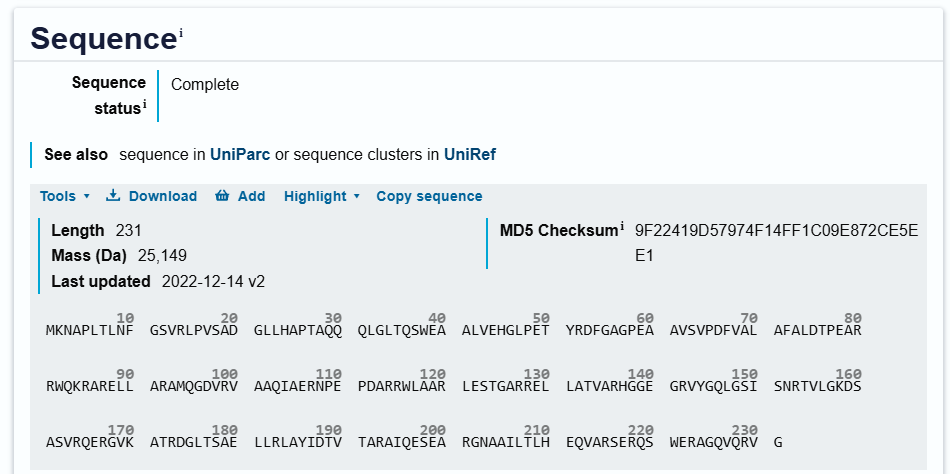

sp|Q9RYE6|DDRC_DEIRA DNA damage response protein C OS=Deinococcus radiodurans (strain ATCC 13939 / DSM 20539 / JCM 16871 / CCUG 27074 / LMG 4051 / NBRC 15346 / NCIMB 9279 / VKM B-1422 / R1) OX=243230 GN=ddrC PE=1 SV=2 MKNAPLTLNFGSVRLPVSADGLLHAPTAQQQLGLTQSWEAALVEHGLPETYRDFGAGPEA AVSVPDFVALAFALDTPEARRWQKRARELLARAMQGDVRVAAQIAERNPEPDARRWLAAR LESTGARRELLATVARHGGEGRVYGQLGSISNRTVLGKDSASVRQERGVKATRDGLTSAE LLRLAYIDTVTARAIQESEARGNAAILTLHEQVARSERQSWERAGQVQRVG
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence
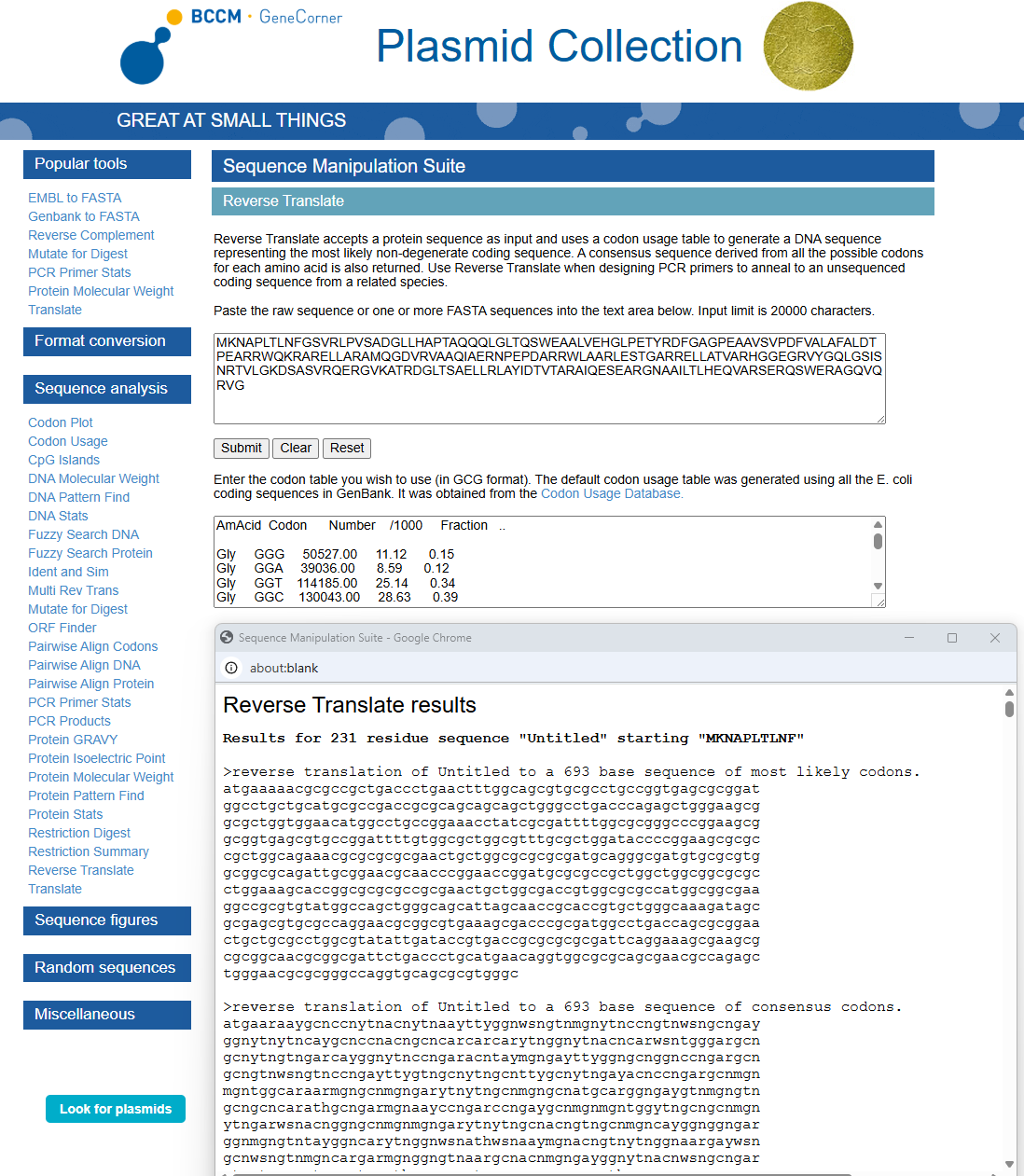

Using https://www.genecorner.ugent.be/rev_trans.html to create reverse translation of Protein DdrC
atgaaaaacgcgccgctgaccctgaactttggcagcgtgcgcctgccggtgagcgcggat ggcctgctgcatgcgccgaccgcgcagcagcagctgggcctgacccagagctgggaagcg gcgctggtggaacatggcctgccggaaacctatcgcgattttggcgcgggcccggaagcg gcggtgagcgtgccggattttgtggcgctggcgtttgcgctggataccccggaagcgcgc cgctggcagaaacgcgcgcgcgaactgctggcgcgcgcgatgcagggcgatgtgcgcgtg gcggcgcagattgcggaacgcaacccggaaccggatgcgcgccgctggctggcggcgcgc ctggaaagcaccggcgcgcgccgcgaactgctggcgaccgtggcgcgccatggcggcgaa ggccgcgtgtatggccagctgggcagcattagcaaccgcaccgtgctgggcaaagatagc gcgagcgtgcgccaggaacgcggcgtgaaagcgacccgcgatggcctgaccagcgcggaa ctgctgcgcctggcgtatattgataccgtgaccgcgcgcgcgattcaggaaagcgaagcg cgcggcaacgcggcgattctgaccctgcatgaacaggtggcgcgcagcgaacgccagagc tgggaacgcgcgggccaggtgcagcgcgtgggc
3.3. Codon optimization
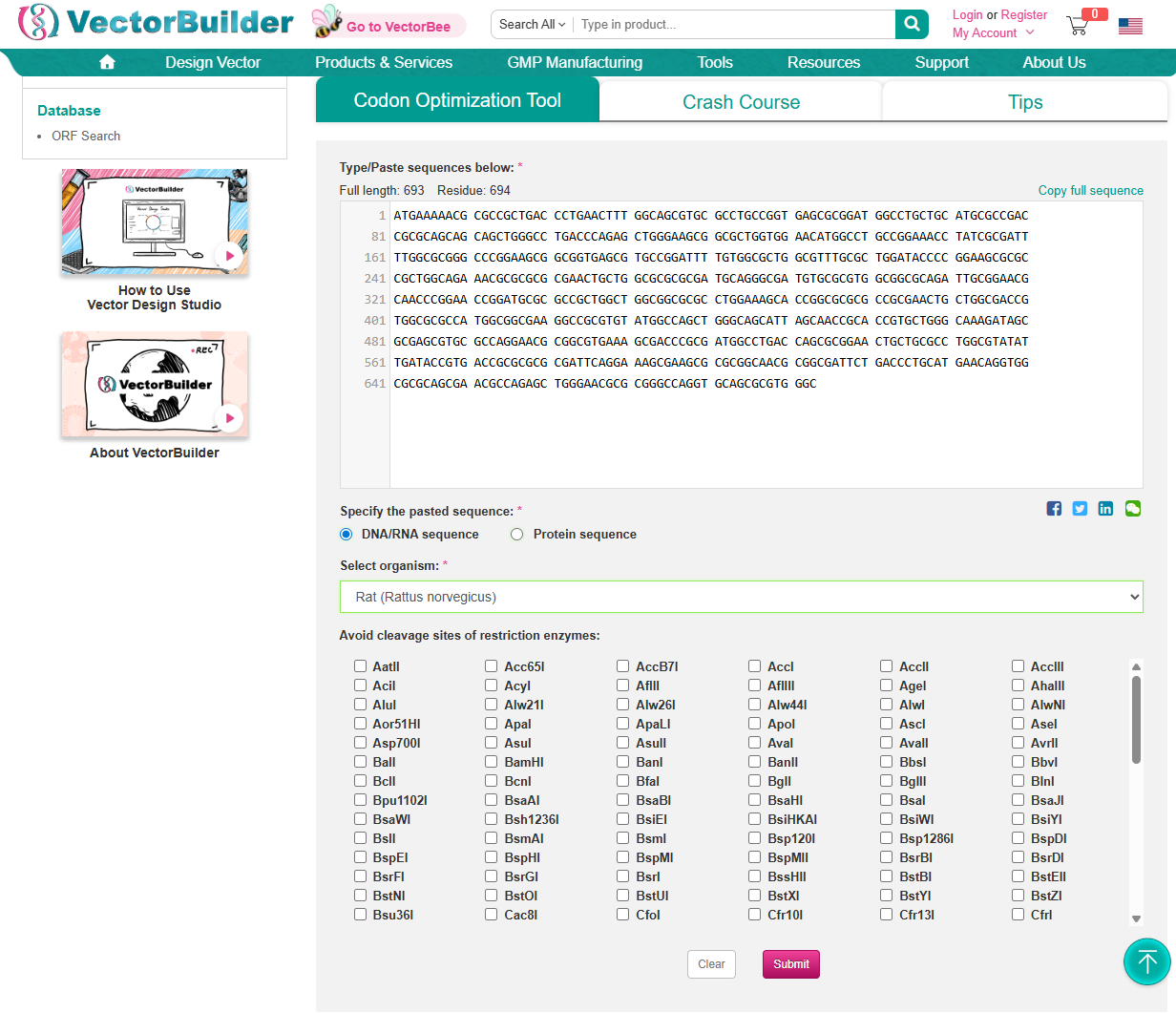

Using https://en.vectorbuilder.com/tool/codon-optimization.html to Condon Optimise DNA/RNA sequence Selected Organism: Rat (Rattus Norvegicus)
In your own words, describe why you need to optimize codon usage. Which organism have you chosen to optimize the codon sequence for and why?
Codon Optimization is required to improve the efficiency of translation of mRNA to Protein. It involves modifying the nucleotide sequence of a gene to replace rare or less-favoured codons with more frequently used codons in the host organsim. The organism I have chosen is a Rat and this is because the codon usuage differs from the orginal source of the DdrC gene. The codon optimisation will replace codons that are rare in rats with preferred rat codons allowing fro faster translation, higher protein expression and more efficient product of the DdrC protein.
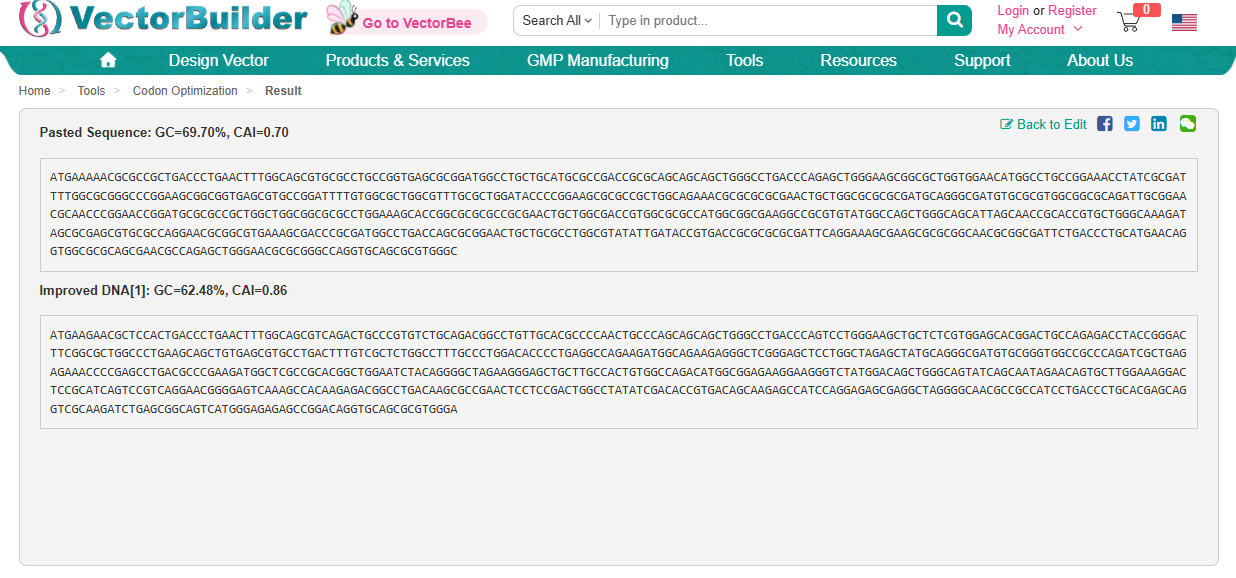

ATGAAGAACGCTCCACTGACCCTGAACTTTGGCAGCGTCAGACTGCCCGTGTCTGCAGACGGCCTGTTGCACGCCCCAACTGCCCAGCAGCAGCTGGGCCTGACCCAGTCCTGGGAAGCTGCTCTCGTGGAGCACGGACTGCCAGAGACCTACCGGGACTTCGGCGCTGGCCCTGAAGCAGCTGTGAGCGTGCCTGACTTTGTCGCTCTGGCCTTTGCCCTGGACACCCCTGAGGCCAGAAGATGGCAGAAGAGGGCTCGGGAGCTCCTGGCTAGAGCTATGCAGGGCGATGTGCGGGTGGCCGCCCAGATCGCTGAGAGAAACCCCGAGCCTGACGCCCGAAGATGGCTCGCCGCACGGCTGGAATCTACAGGGGCTAGAAGGGAGCTGCTTGCCACTGTGGCCAGACATGGCGGAGAAGGAAGGGTCTATGGACAGCTGGGCAGTATCAGCAATAGAACAGTGCTTGGAAAGGACTCCGCATCAGTCCGTCAGGAACGGGGAGTCAAAGCCACAAGAGACGGCCTGACAAGCGCCGAACTCCTCCGACTGGCCTATATCGACACCGTGACAGCAAGAGCCATCCAGGAGAGCGAGGCTAGGGGCAACGCCGCCATCCTGACCCTGCACGAGCAGGTCGCAAGATCTGAGCGGCAGTCATGGGAGAGAGCCGGACAGGTGCAGCGCGTGGGA
3.4. You have a sequence! Now what?
What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both.
Cell-free protein synthesis - Using extracted cellular machinery
DNA is inserted into a reaction mixture and the sequence starts with ATG
Transcription machinery with RNA polymerase which binds ot the DNA and transcribes it into mRNA
mRNA is produced, containing codons
Translation of mRNA with reaction mixture containing: Ribosomes, tRNA molecules, 20 amino acids and ATP
Translation stars with Ribosomes binding to the mRNA, the translation starts at the codon AUG, and each codon is read in order. tRNA brings the correct amino acids and the amino acids are joined to form a polypetide chain.
Translation ends and the protein is released.
The protin then folds into the correct shape and becomes functional.
3.5. (Optional) How does it work in nature/biological systems?
- Describe how a single gene codes for multiple proteins at the transcriptional level.
A single gene can code for multiple proteins at the transcription level due to alternative splicing, it is a process where different sections of the a gene’s DNA can be selected and combined in different ways. This then results in the product of different proten variants with unqiue functions (Science of Bio Genetics, 2023).
- Try aligning the DNA sequence, the transcribed RNA, and also the resulting translated Protein!!! See example below.
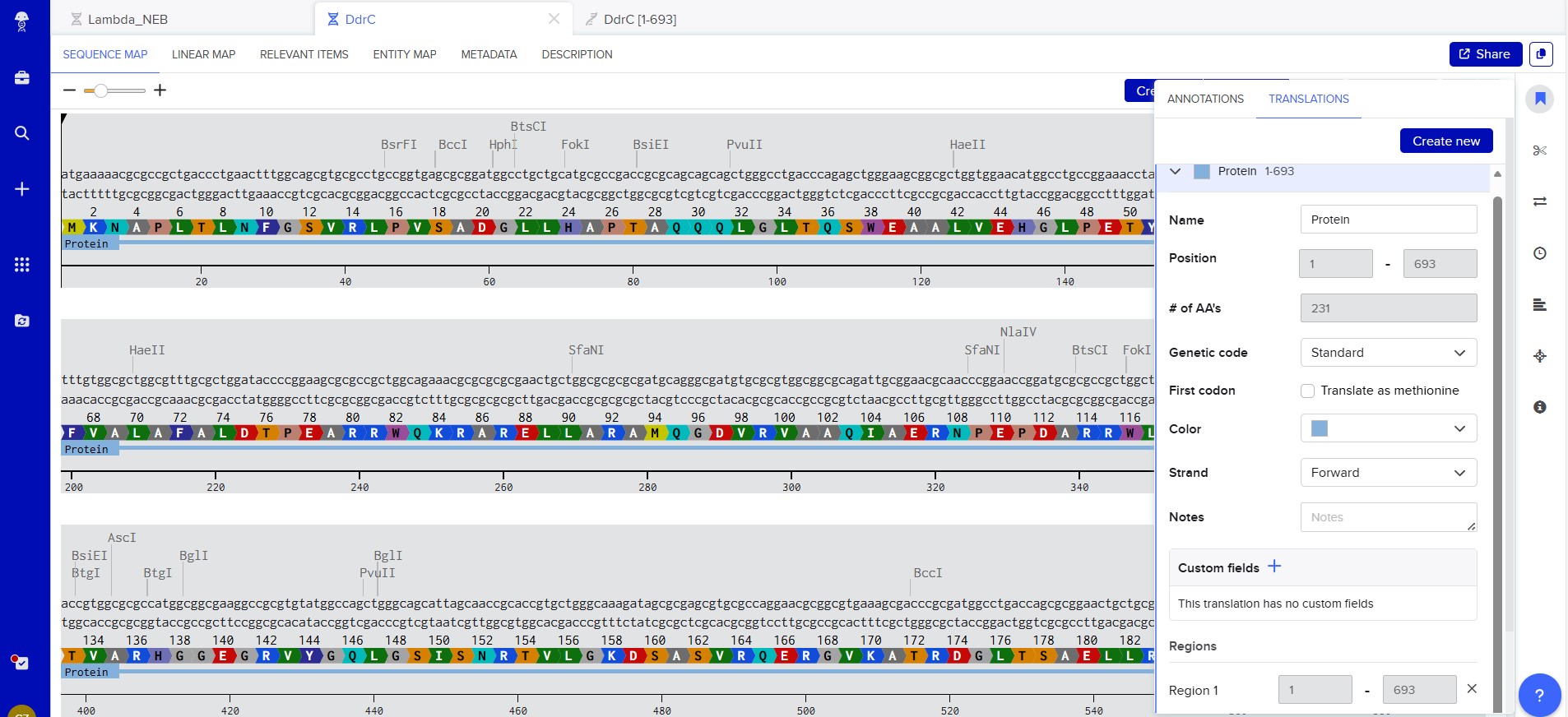

Created new DNA sequence entry and translated to Protein

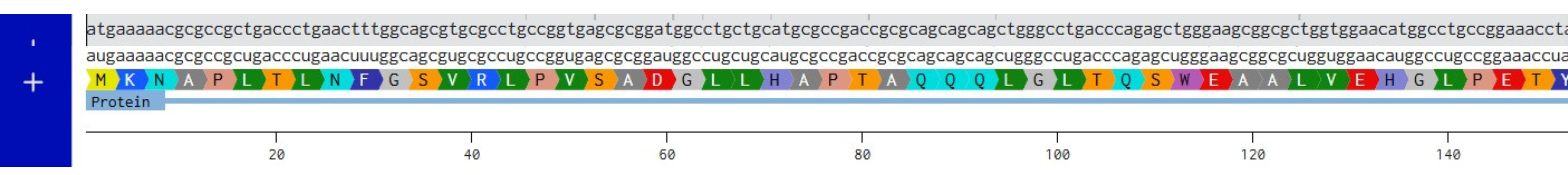
Created mRNA from DNA

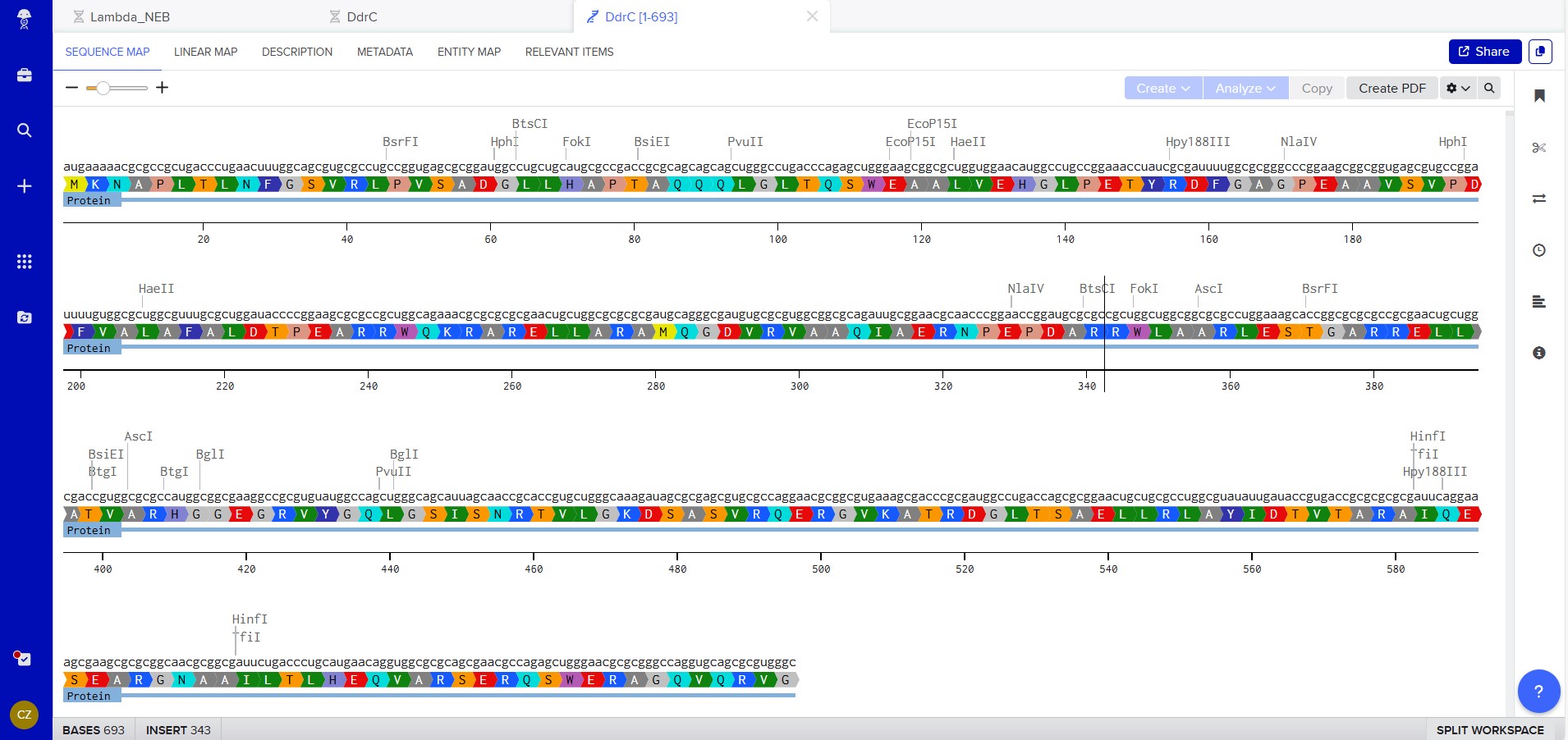
Combined DNA, mRNA and Protein
Part 4: Prepare a Twist DNA Synthesis Order
4.1. Create a Twist account and a Benchling account

4.2. Build Your DNA Insert Sequence
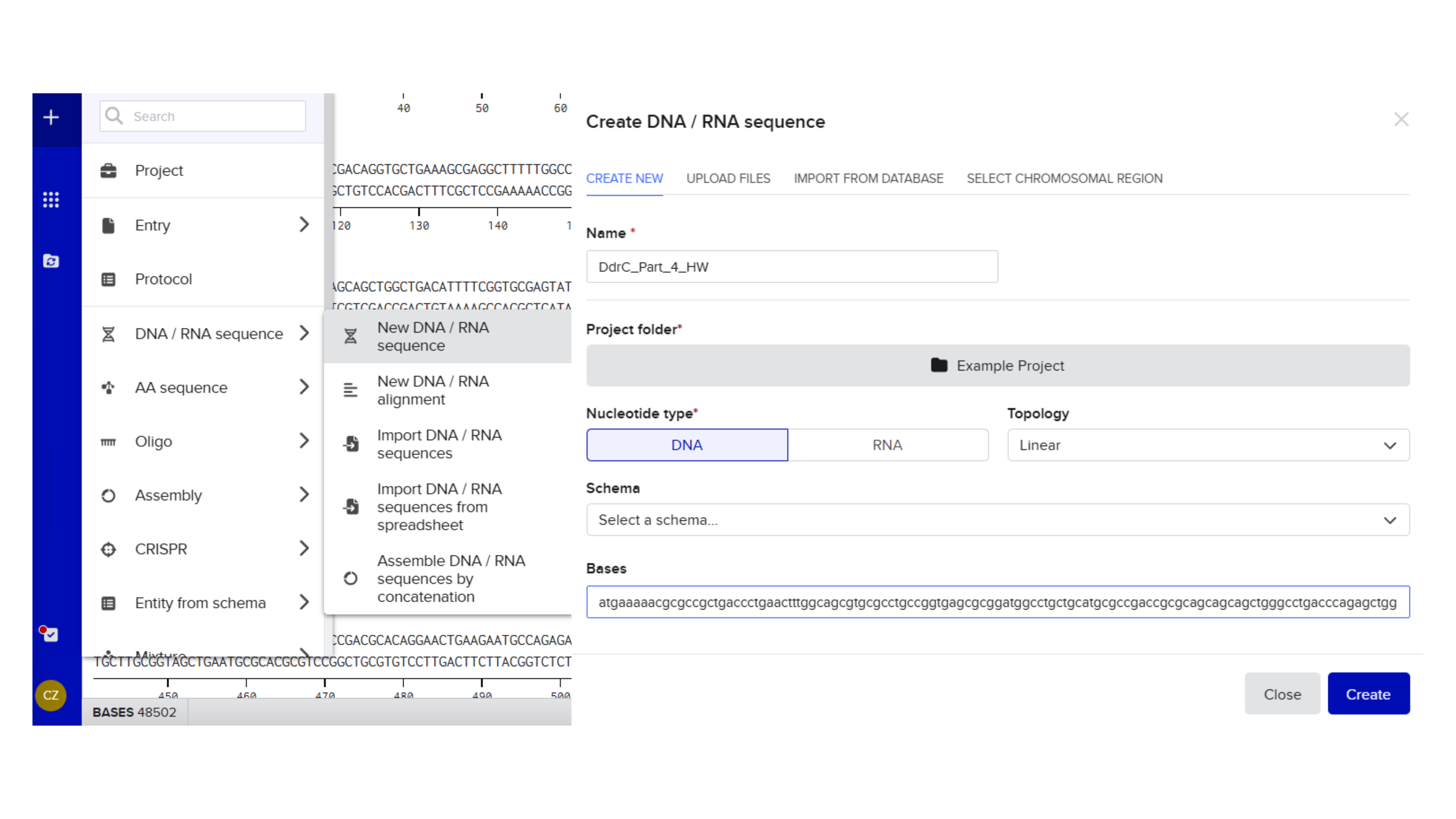

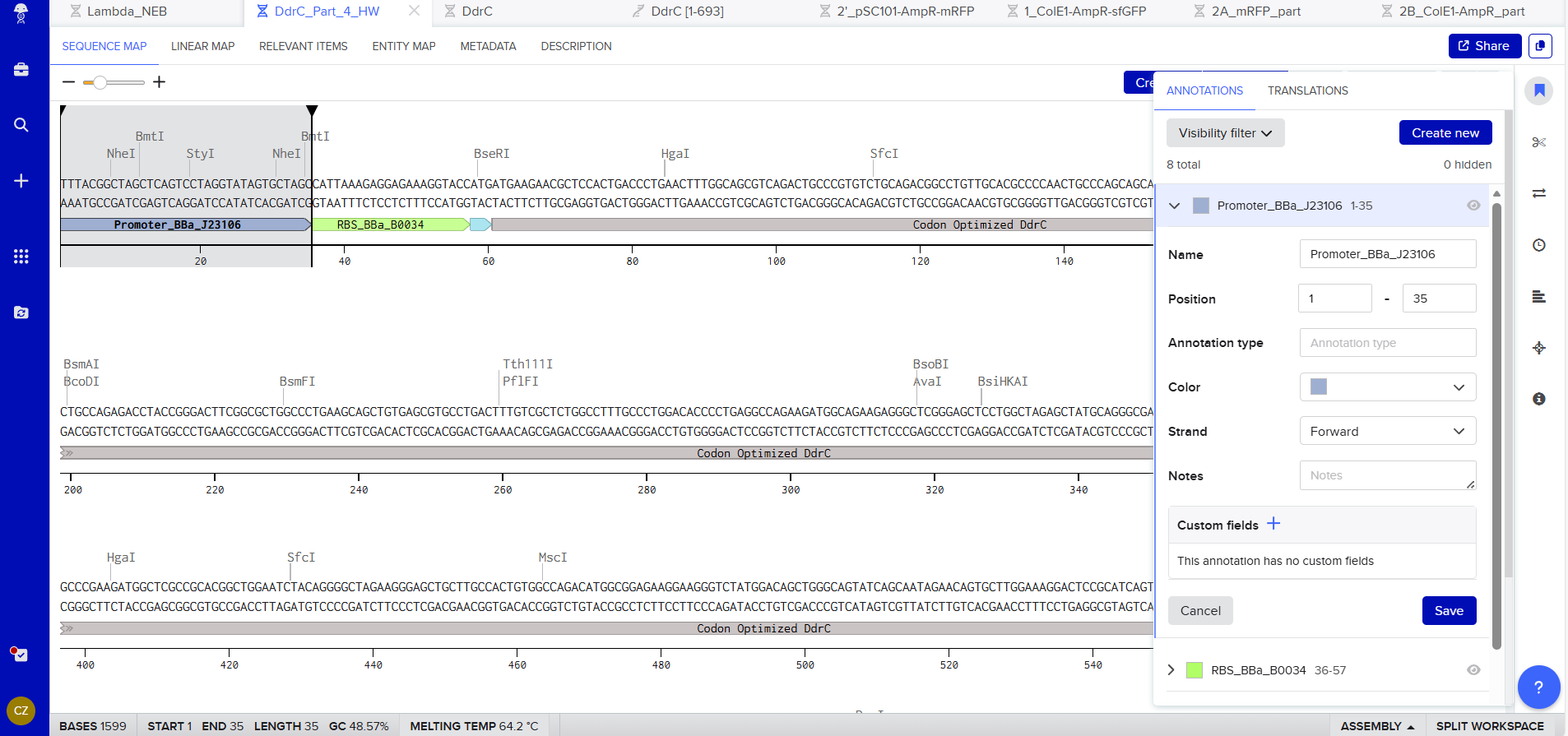

Promoter (e.g. BBa_J23106): TTTACGGCTAGCTCAGTCCTAGGTATAGTGCTAGC
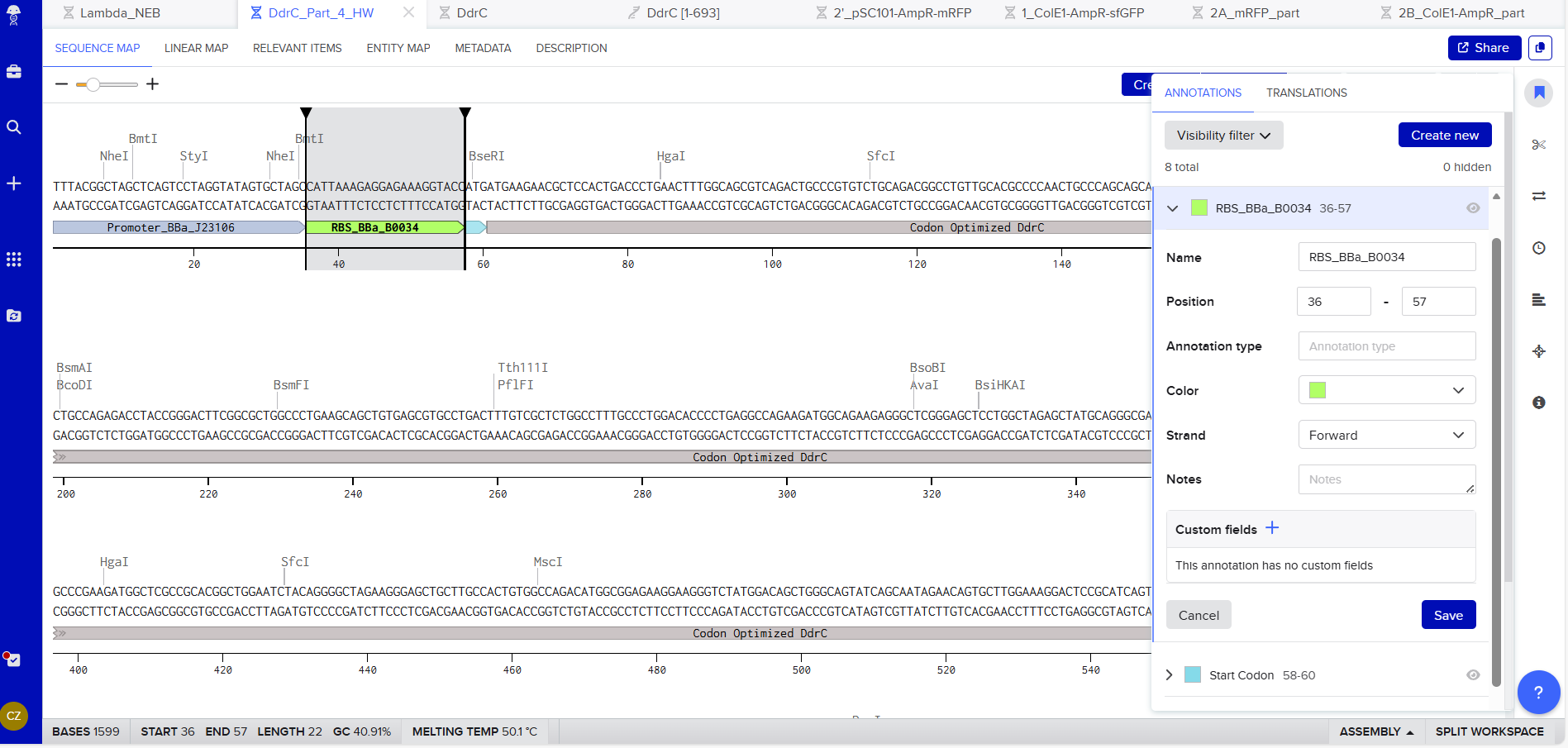

RBS (e.g. BBa_B0034 with spacers for optimal expression): CATTAAAGAGGAGAAAGGTACC
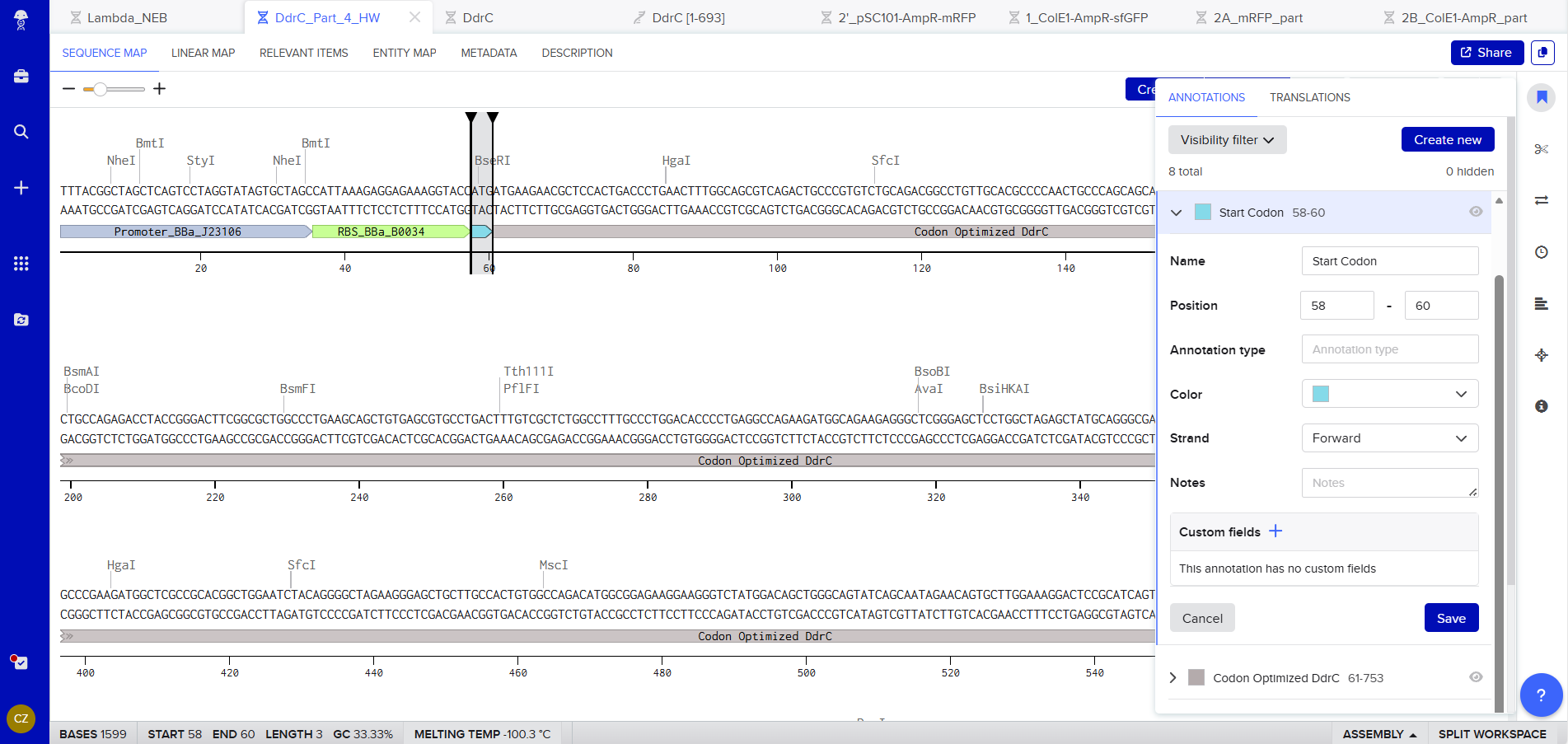

Start Codon: ATG
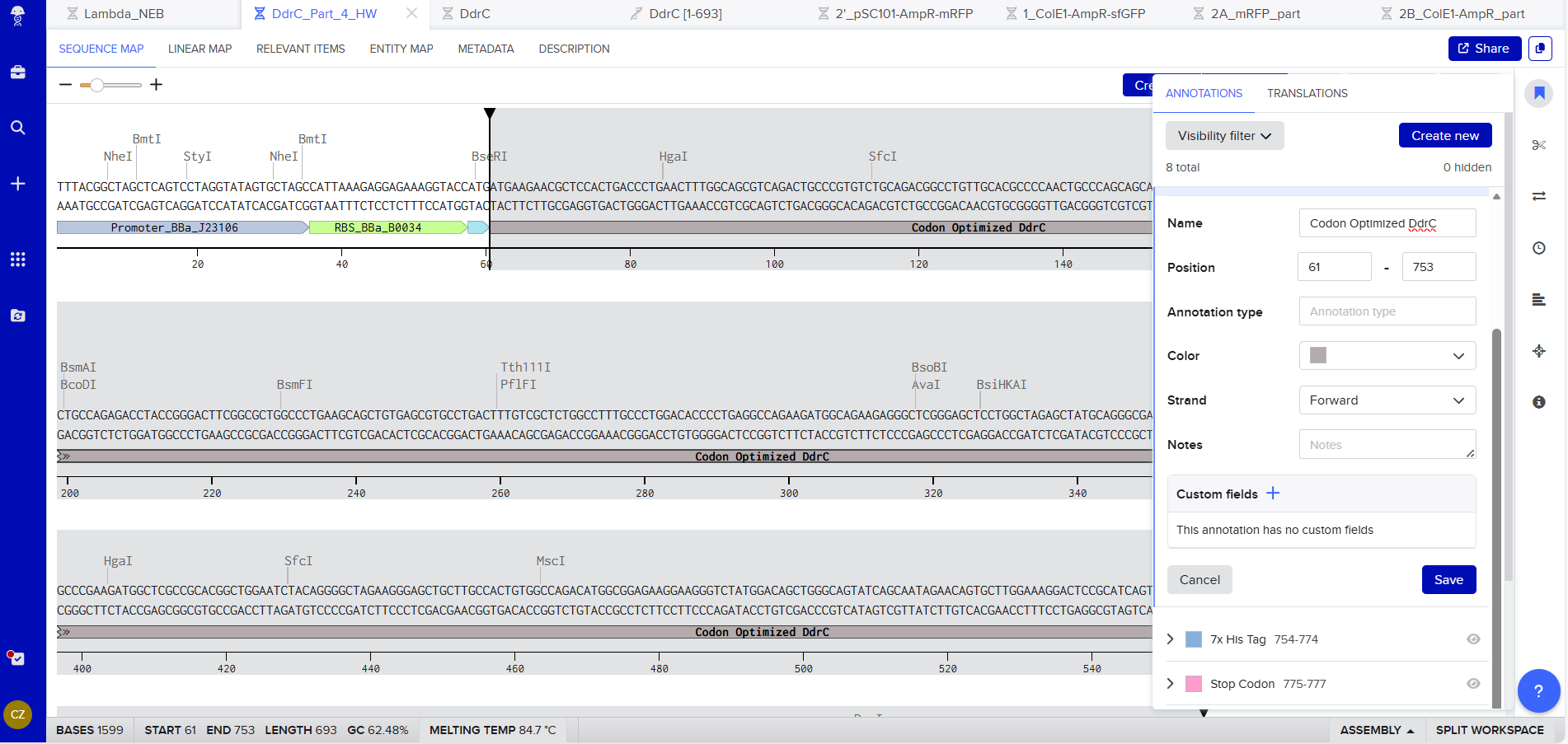

Coding Sequence codon optimized DNA
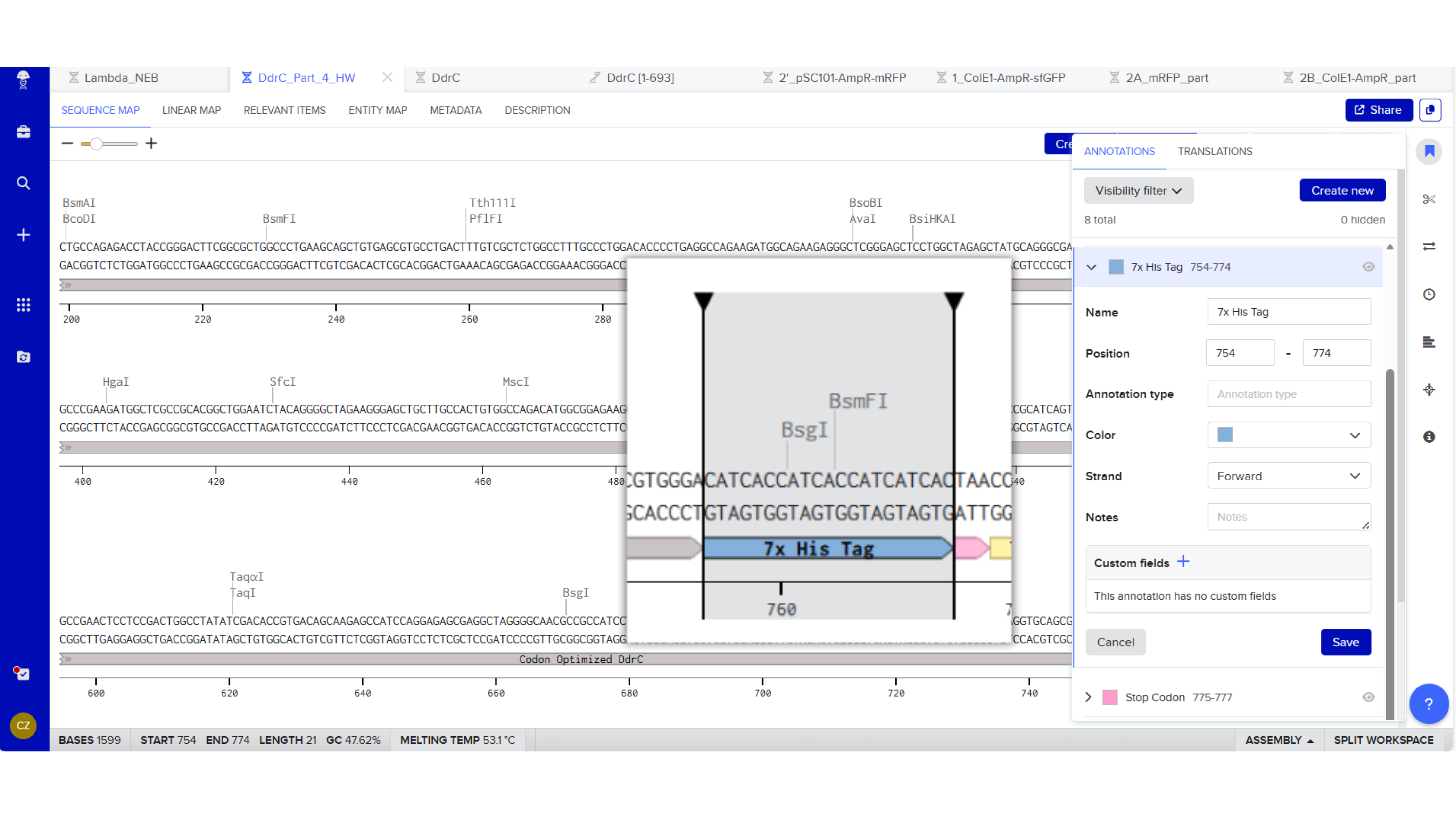

7x His Tag : CATCACCATCACCATCATCAC
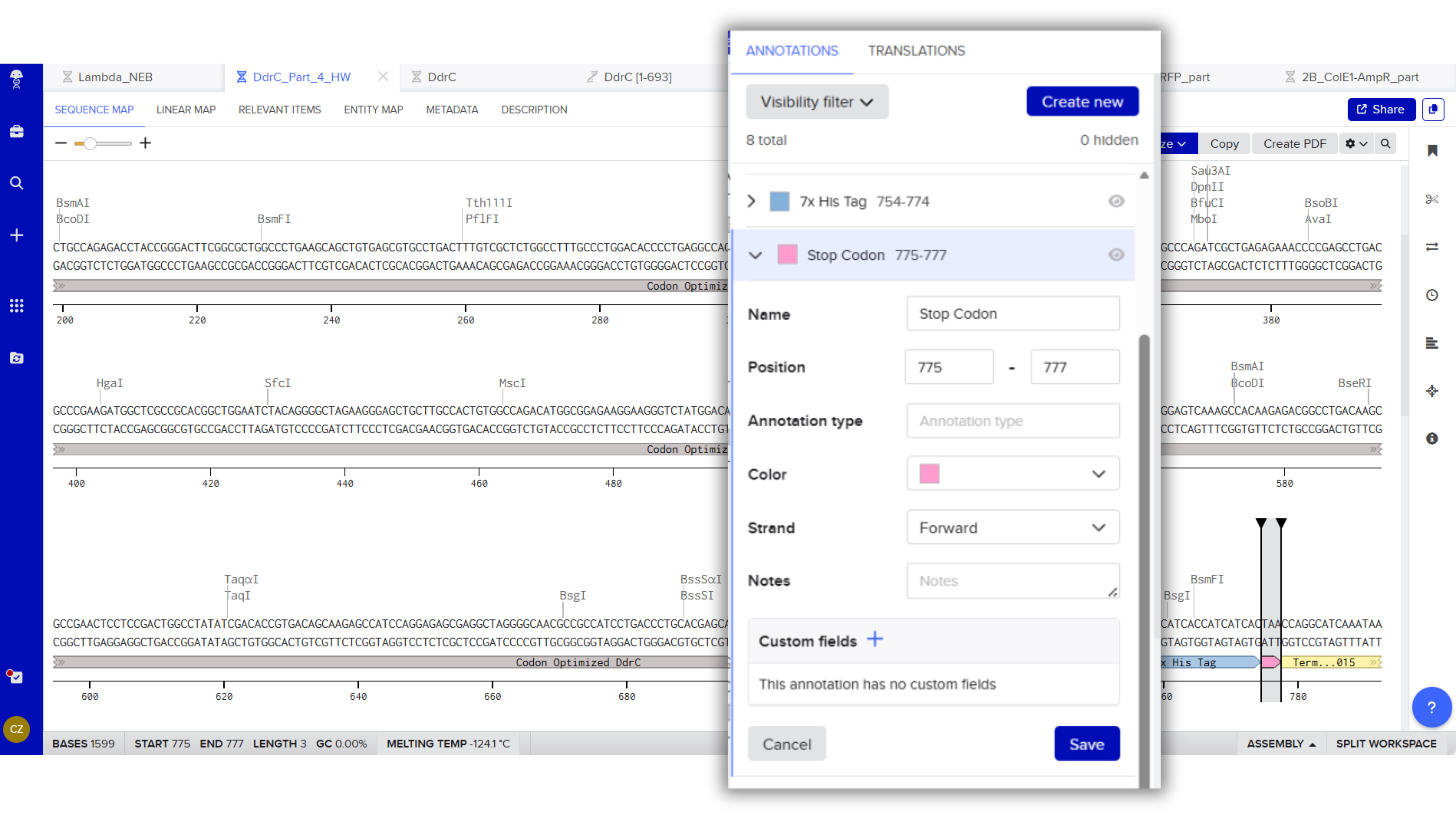

Stop Codon: TAA
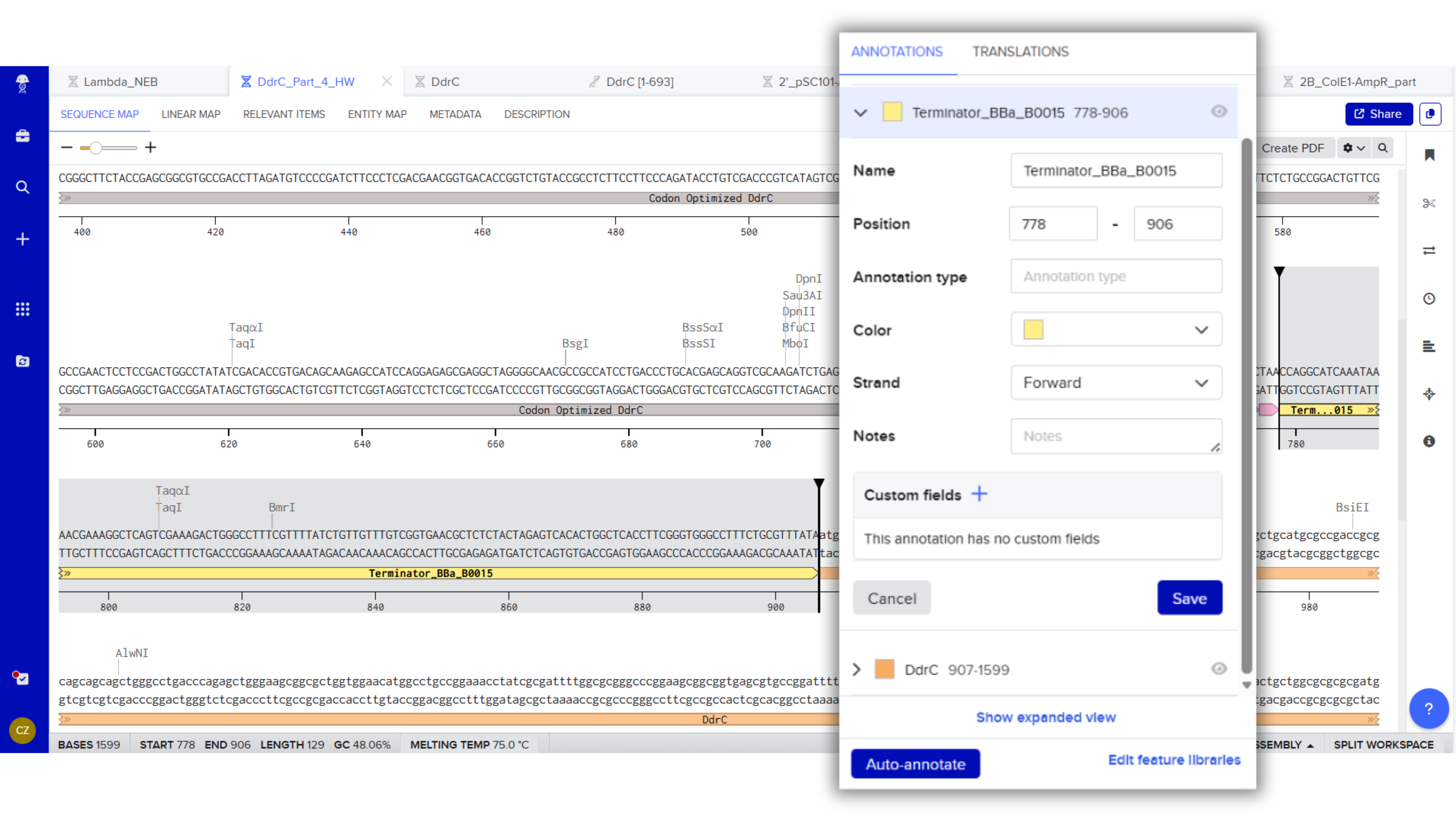

Terminator (e.g. BBa_B0015): CCAGGCATCAAATAAAACGAAAGGCTCAGTCGAAAGACTGGGCCTTTCGTTTTATCTGTTGTTTGTCGGTGAACGCTCTCTACTAGAGTCACACTGGCTCACCTTCGGGTGGGCCTTTCTGCGTTTATA
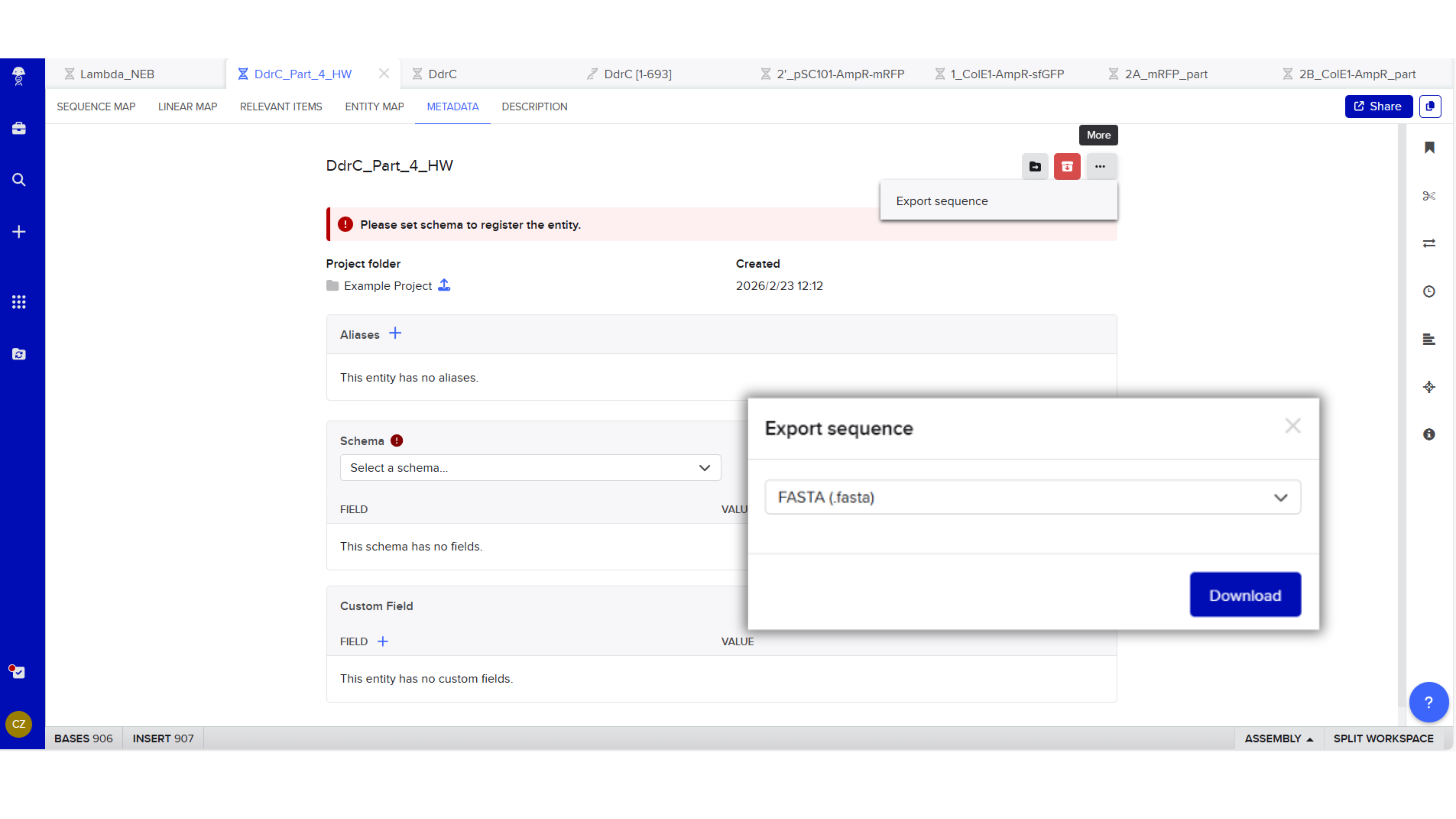

Export Sequence
4.3. On Twist, Select The “Genes” Option
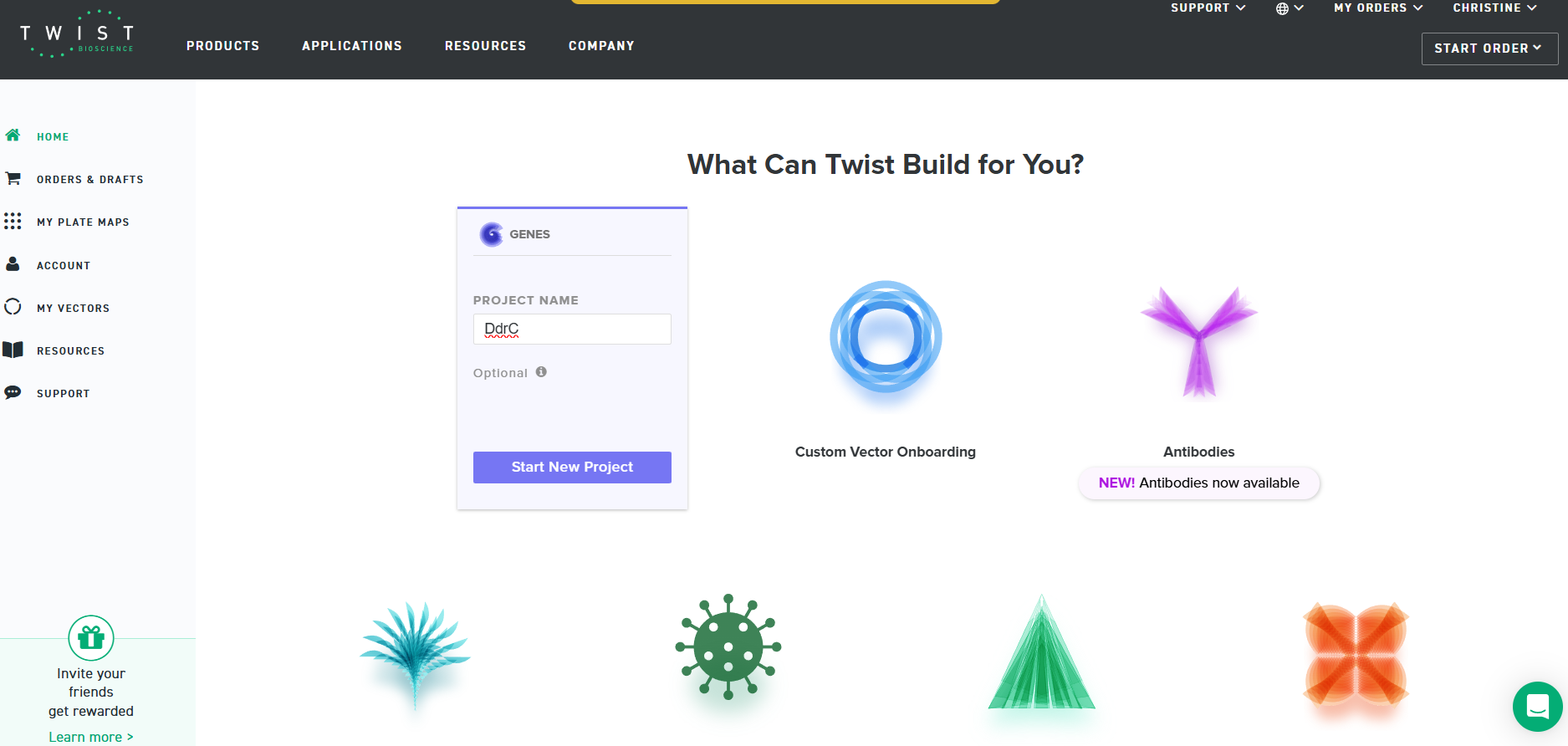

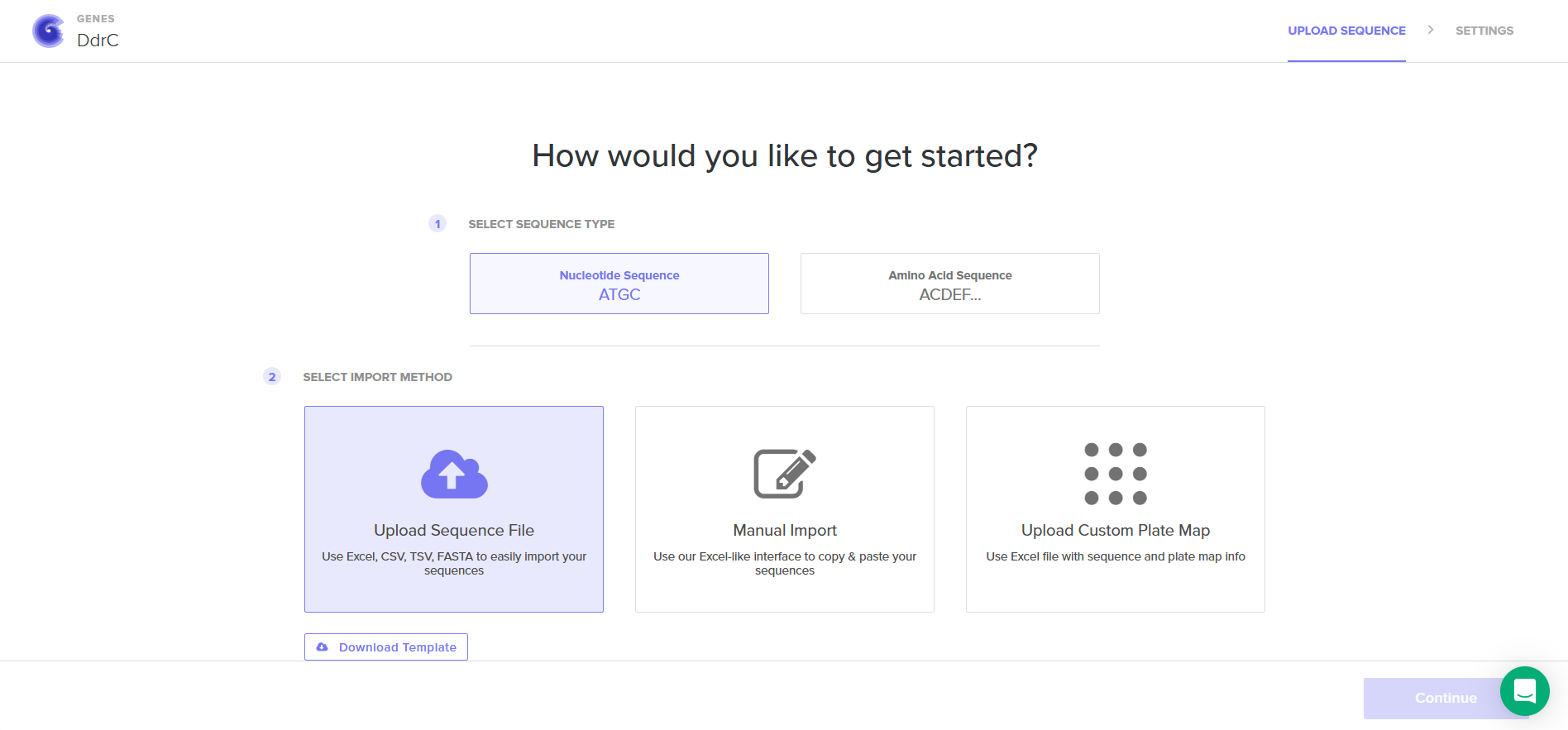

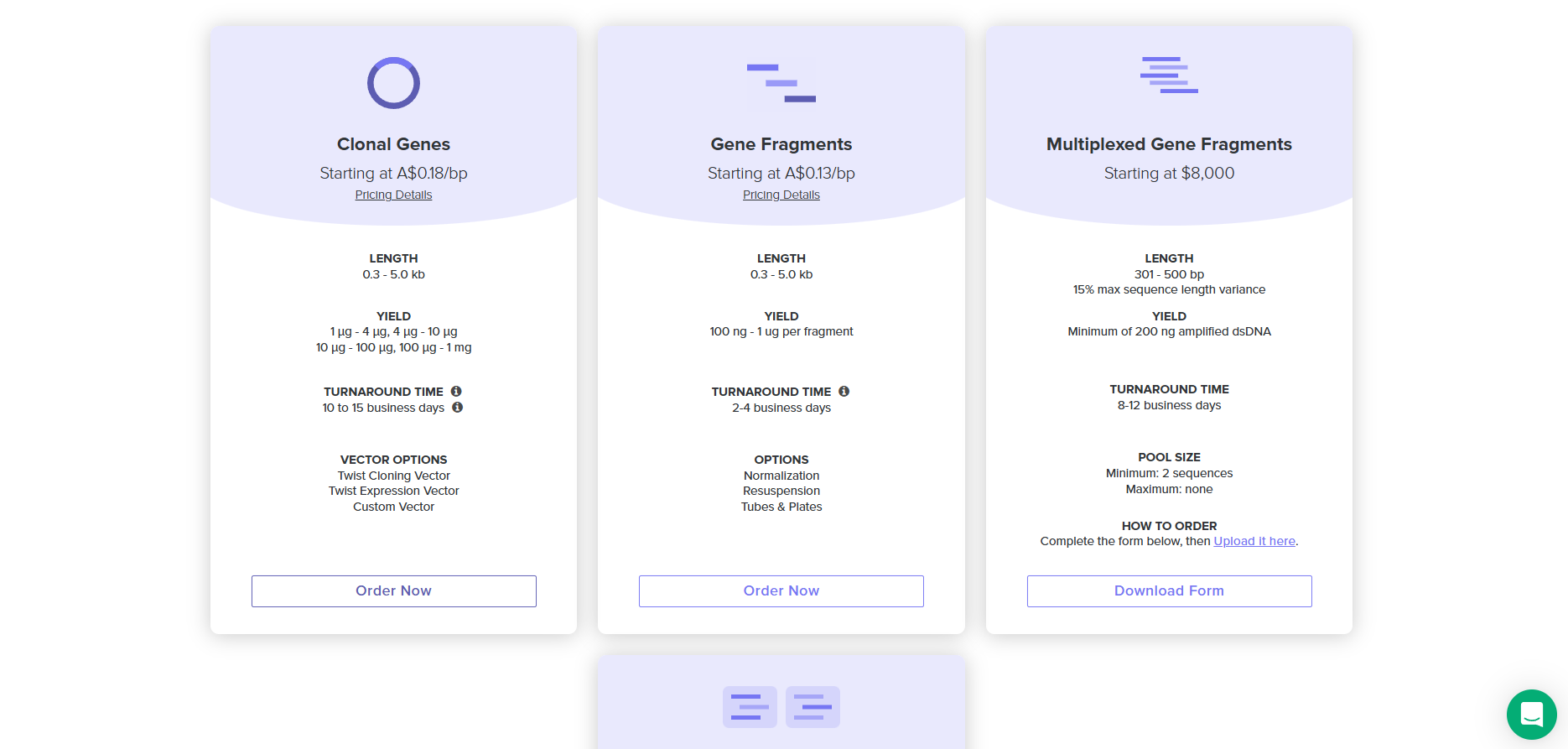
4.4. Select “Clonal Genes” option

4.5. Import your sequence
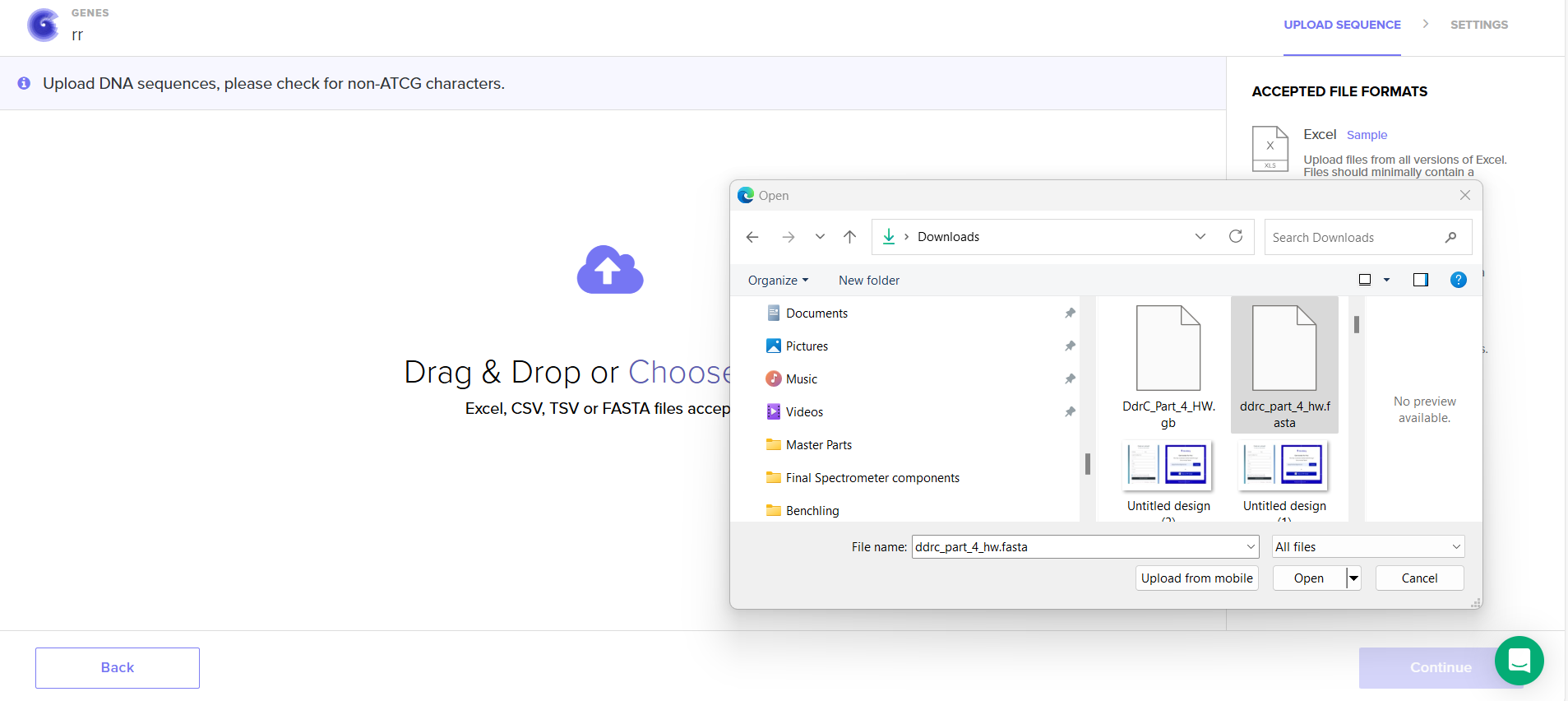

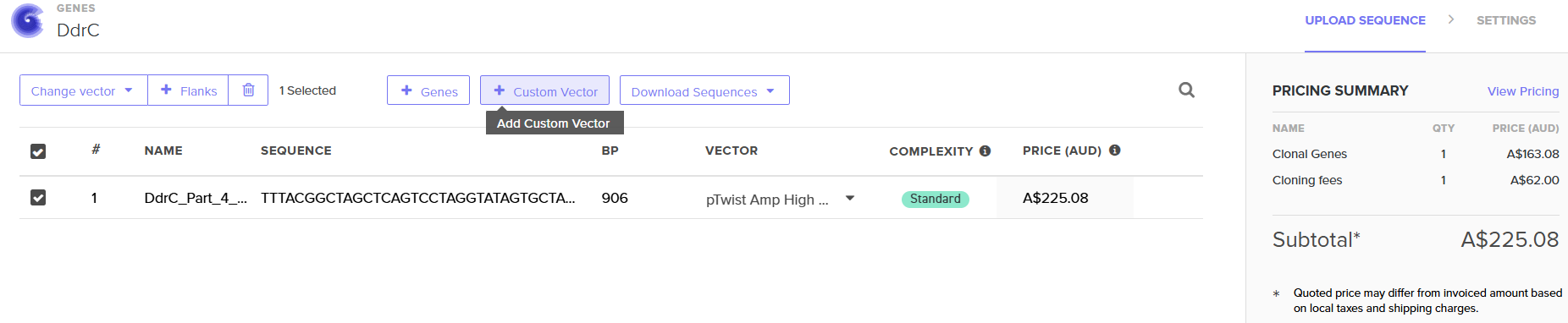

4.6. Choose Your Vector
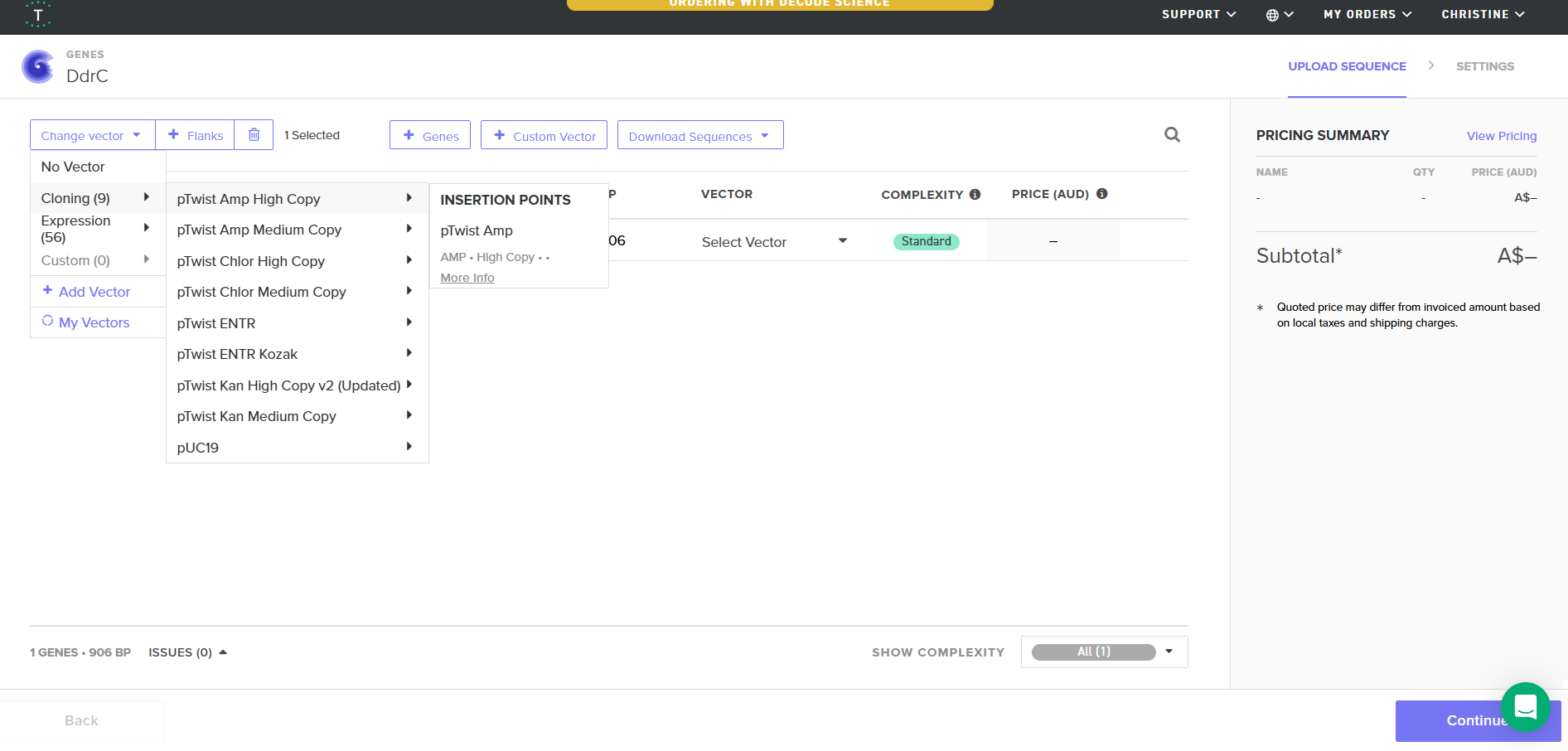

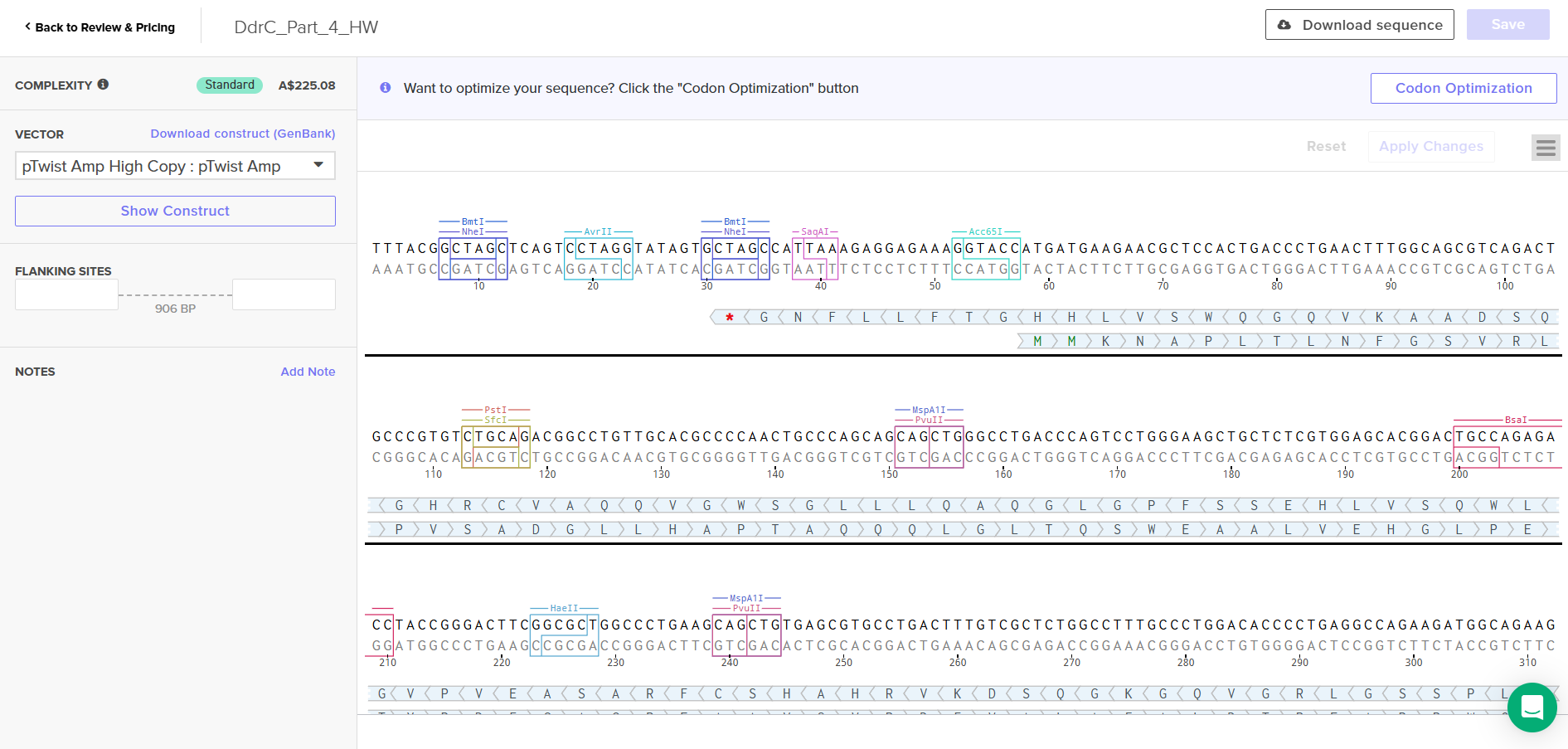

Download Construct (GenBank) and import to Benchling

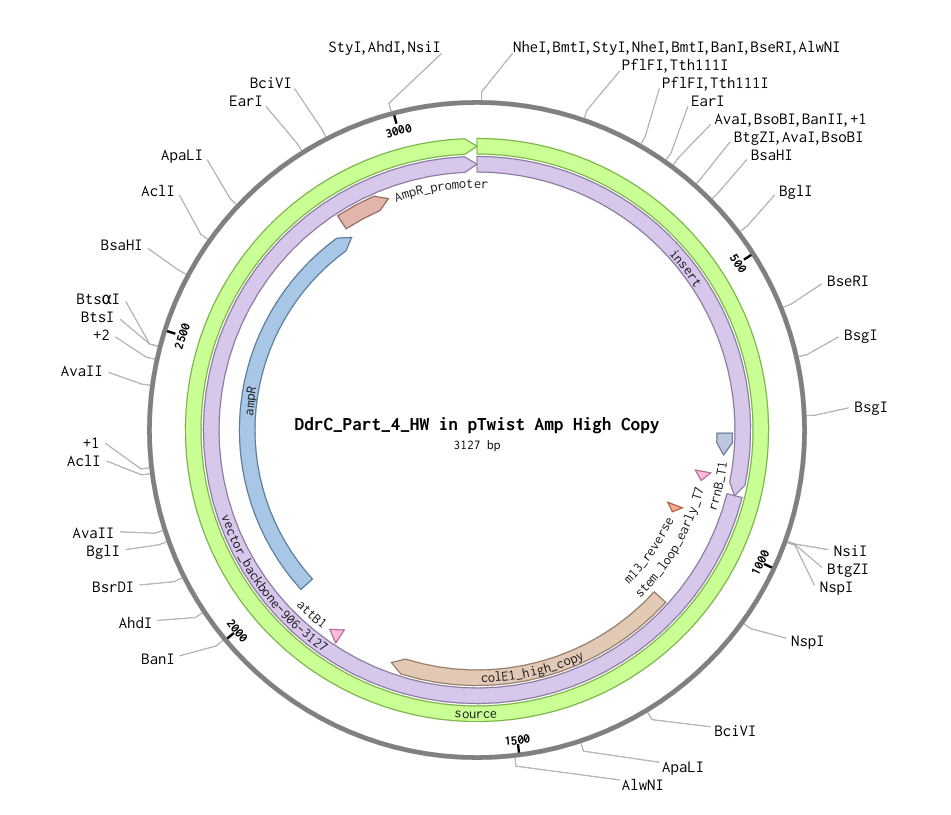

Part 5: DNA Read/Write/Edit
a) What DNA would you want to sequence (e.g., read) and why?
I want to sequence the DNA (genome) of the bacterium called Paenibacillus Vortex seen in figure 1, because of these microorganism’s social motility and ability to employ cell-to-cell signaling to prompt acitvites such as attraction and repulsion under different environmental conditons (Myers, 2012). An example of this is when grown on hard surfaces, P.vortex will generate aggregates of dense bacteria that are pushed forward by repulsive chemotactic signals sensed from the cells at the back (Sirota-Madi et al., 2010). And when grown on soft surfaces these microorgranism can exhbit collective motility by forming foraging amrs that are sent out in search of food (Myers, 2012).
To achieve these cooperative ventures, P.Vortex has to be able to communicate with each other. The information exchanged are regarding its population size, myriad of indiviual environmental measurements at different locations, their internal states and their phenotypic and epigentic adjustments (Sirota-Madi et al., 2010). I want to know what sensors guides them to navigate the environment and what are the tranducers and responders that process the information into action. How do they communicate to each other?
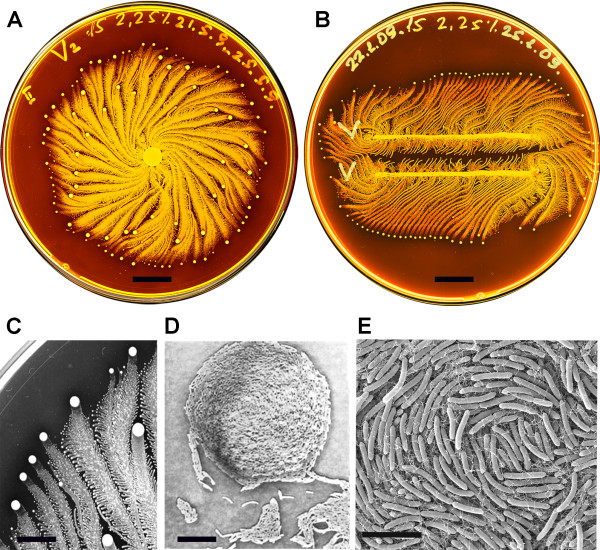

Figure 1, (Sirota-Madi et al., 2010) Colony Organisation of the P.vortex bacteria A. Whole colony view of P.vortex (grown on on 15 g/l peptone and 2.25% (w/v) agar for four days) B. Two colonies of P.vortex (involuated in two parallel lines grown on on 15 g/l peptone and 2.25% (w/v) agar) C. Colony pattern and vortex process (Magnification x20) D. Mature indivdual Vortex (Magnification x500) E. Each individual baterium has curvature (scans fro electron micropscope)
b) What technology or technologies would you use to perform this sequencing and why?
With reference to the research paper Genome sequence of the pattern forming Paenibacillus vortex bacterium reveals potential for thriving in complex environments by Sirota-Madi et al. (2010), the reserachers used a hybrid approach of 454 Life Sciences and Illumina, achieving a total of 289× coverage, with 99.8% sequence identity between the two methods (Sirota-Madi et al., 2010). This is because the reserachers claimed that p.Vortex has an exceptionally large number of signal transduction genes and a high combined regulatory score (TCS, TFs, transport, defense genes). In order to verify this, they would require a hihgly accurate genome, minimial assembly artifacts and reliable gene annotation. The hybrid approach can reduce the distortions that come from using one sequencting technology.
i) Is this your method first-, second-, or third-generation or other? How so?
Illumina and 454 Life Sciences are both second generation sequencing. 454 provides longer reads creating better contiguity whilst Illumina provides massive depth hence providing higher accuracy. The researchers chose this combination of methods because of the De novo assembly problem (no reference genome). Short reads alone (especially early Illumina reads) are difficult to assemble reliably and the hybrid allowed more reliable identification of regulatory and signal transduction genes.
ii) What is your input? How do you prepare your input? List the essential steps.
Input:
Step 1: Bacteria growth
- P. vortex grown overnight in LB medium
- 37 °C, shaking at 200 rpm
- Ensures sufficient biomass
Step 2: Cell Harvesting
- DNA extracted from 2 ml culture (~10⁹ cells/ml)
Step 3: Cell Lysis
- Cells incubated with lysozyme for 45 minutes
- Breaks down bacterial cell wall (important for Gram-positive bacteria)
Step 4: DNA Extraction
- Qiagen DNeasy Blood & Tissue Kit
- Removes proteins, lipids, and other contaminants
Step 5: DNA Elution
- Eluted in 200 µl AE buffer (10 mM Tris-HCl, 0.5 mM EDTA pH 9.0)
iii) What are the essential steps of you chosen sequencing technology, how does it decode the bases of your DNA sample?
Sequencing approach incorporated 454 pyrosequencing with Illumina Genome Analyzer.
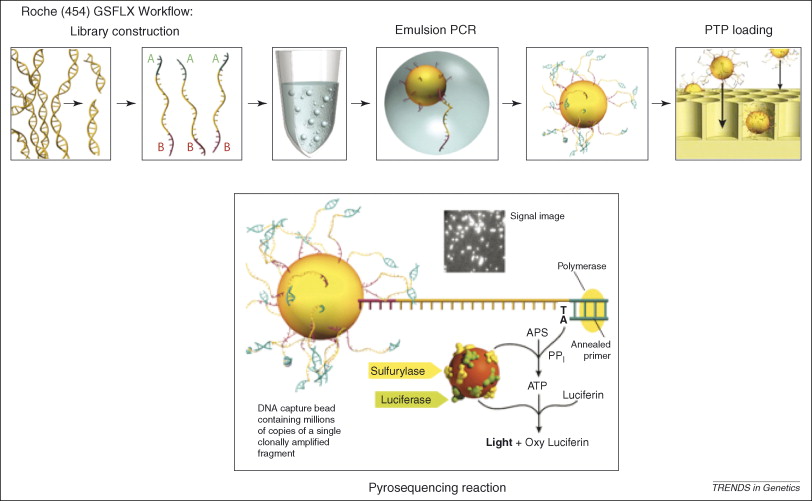
 Figure 2, (MacLean, Jones, & Studholme, 2009)
Figure 2, (MacLean, Jones, & Studholme, 2009)
Referecin MacLean, Jones, and Studholme (2009) seen in figure 2, the essential steps of 454 pyrosequencing are:
DNA fragmentation and adapter ligation
Genomic DNA is broken into small fragments and short adapter sequences are attached.
Bead attachment and emulsion PCR
Each DNA fragment is attached to a microscopic bead and amplified inside an oil droplet.
Bead loading into wells
DNA-coated beads are placed into tiny wells, one bead per well.
Sequencing by nucleotide incorporation
DNA polymerase adds nucleotides to the growing DNA strand one type at a time.
Light generation (pyrosequencing)
When a nucleotide is incorporated, pyrophosphate is released, triggering a reaction that produces light.
Signal detection and base calling
The emitted light is recorded and used to determine the DNA sequence.
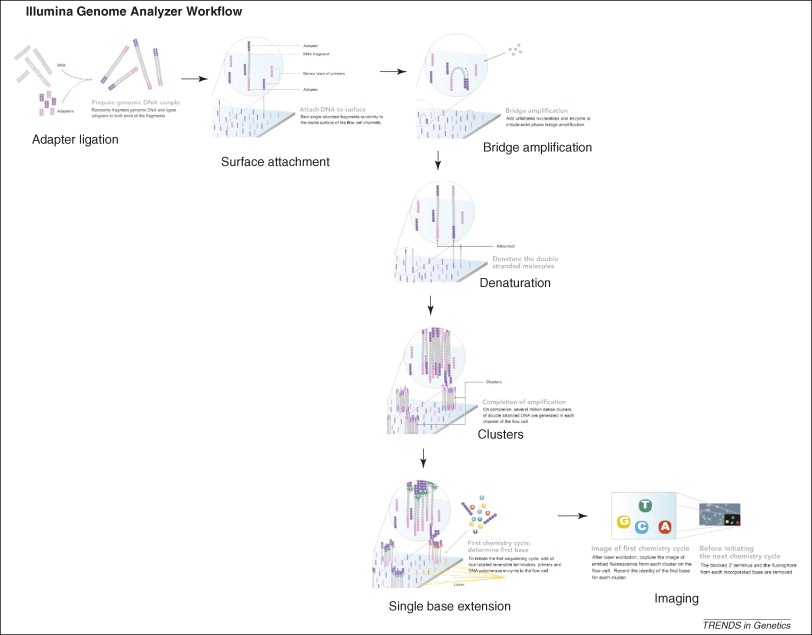
 Figure 3, (MacLean, Jones, & Studholme, 2009)
Figure 3, (MacLean, Jones, & Studholme, 2009)
Referecin MacLean, Jones, and Studholme (2009) seen in figure 3, the essential steps of Illumina Genome Analyzer sequencing are:
DNA Fragmentation
Mixture of single-stranded, adaptor oligo-ligated DNA fragments
Attachment to the flow cell
Using a microfluidic cluster station to add these fragments to the surface of a glass flow cell.
Each flow cell is divided into eight separate lanes, and the interior surfaces have covalently attached oligos complementary to the specific adapters that are ligated onto the library fragments
- Cluster amplification (bridge PCR)
- Hybridization of these DNAs to the oligos on the flow cell occurs by an active heating and cooling step
- Subsequent incubation with reactants and an isothermal polymerase amplifies the fragments in a discrete area or ‘cluster’ on the flow cell surfaces
- Flow cell is placed into a fluidics cassette within the sequencer
- Each cluster is supplied with polymerase and four differentially labeled fluorescent nucleotides that have their 3′-OH chemically inactivated to ensure that only a single base is incorporated per cycle. (MacLean, Jones, & Studholme, 2009)
- Identify the incorporated nucleotide at each cluster
- Each base incorporation cycle is followed by an imaging step, and by a chemical step removes the fluorescent group and deblocks the 3′ end for the next base incorporation cycle. (MacLean, Jones, & Studholme, 2009)
- End of the sequencing run
- The sequence of each cluster is computed and subjected to quality filtering to eliminate low-quality reads of between 32 and 40bp. (MacLean, Jones, & Studholme, 2009)
iv) What is the output of your chosen sequencing technology?
Outputs are raw sequences, base quality scores and FASTQ files
DNA WRITE
a) What DNA would you want to synthesize (e.g., write) and why?


I want to synthesise the gene encoding the luminescent protein from Aequorea vicotria jellyfish (GFP Green Fluprescent Protein) with a Plasmid to emit light natrually without electricity.
b) What technology or technologies would you use to perfom this DNA synthesis and why?
The technologies I would use to perform this DNA synethesis are CRISPR-Cas9 becasue it is extremely precise and can insert GFP exactly in the desired location. It can work in bateria, yeast and plants and allows for stable genome integration.
DNA EDIT
a) What DNA would you want to edit and why?
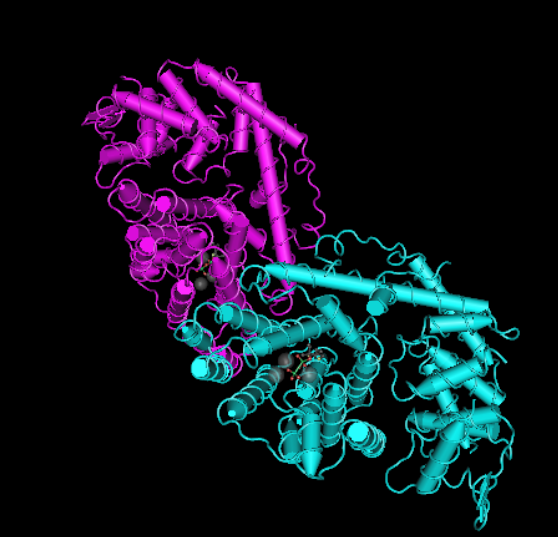

- limonene synthase (Prem Kumar, Morehouse, Yu, & Oprian, 2019)*
The protein that I can to edit is limonene synthase (citrus sinensis) because I think it is interesting to edit smells, what would happen if you edit the smell of sweet orange and grow a fruit with the edited scent? Would it result in reduced fruit quality, the fruit will have no orange smell, how does smell evolve? What are some environmental factors that can change the DNA of certain smells.
b) What technology or technologies would you use to perfom these DNA edits and why?
I could utilise base editors where I can edit the DNA without creating double strand breaks, I can introduce specific point mutations in the Limonene synthase to tweak the enzyme’s function.
Bibliorgraphy
Gueguen, E., Bruto, M., Lemaire, D., Bertrand, E., Fichant, G., & Graille, M. (2022). Structural and functional characterization of DdrC, a novel DNA damage-induced nucleoid associated protein involved in DNA compaction. Nucleic Acids Research, 50(13). https://doi.org/10.1093/nar/gkac563
MacLean, D., Jones, J. D., & Studholme, D. J. (2009). Application of “next-generation” sequencing technologies to microbial genetics. Nature Reviews Microbiology, 7(4), 287–296. https://doi.org/10.1038/nrmicro2122
Morehouse, B.R., Kumar, R.P., Matos, J.O., Yu, Q., Bannister, A., Malik, K., Temme, J.S., Krauss, I.J. & Oprian, D.D. (2019) Direct Evidence of an Enzyme‑Generated LPP Intermediate in (+)-Limonene Synthase Using a Fluorinated GPP Substrate Analog. ACS Chemical Biology, 14, 2035‑2043. https://doi.org/10.1021/acschembio.9b00514
Myers, W. (2018). Bio design: Nature, science, creativity (Paola Antonelli, Foreword). Thames & Hudson.
Sirota-Madi, A., Olender, T., Helman, Y., Ingham, C., Brainis, I., Roth, D., Hagi, E., Brodsky, L., Leshkowitz, D., Galatenko, V., Nikolaev, V., Mugasimangalam, R. C., Bransburg-Zabary, S., Gutnick, D. L., Lancet, D., & Ben-Jacob, E. (2010). Genome sequence of the pattern forming Paenibacillus vortex bacterium reveals potential for thriving in complex environments. BMC Genomics, 11, 710. https://doi.org/10.1186/1471-2164-11-710
The unique mechanism of DdrC in enhancing DNA stability. (2024, December 30). Optimise.mfm.au. Retrieved from https://optimise.mfm.au/research/the-unique-mechanism-of-ddrc-in-enhancing-dna-stability/
Unknown author. (n.d.). JellyFish [Photograph]. The Pipette Pen. http://www.thepipettepen.com/wp-content/uploads/2016/04/7338555110_bc2ba135a5_z.jpg