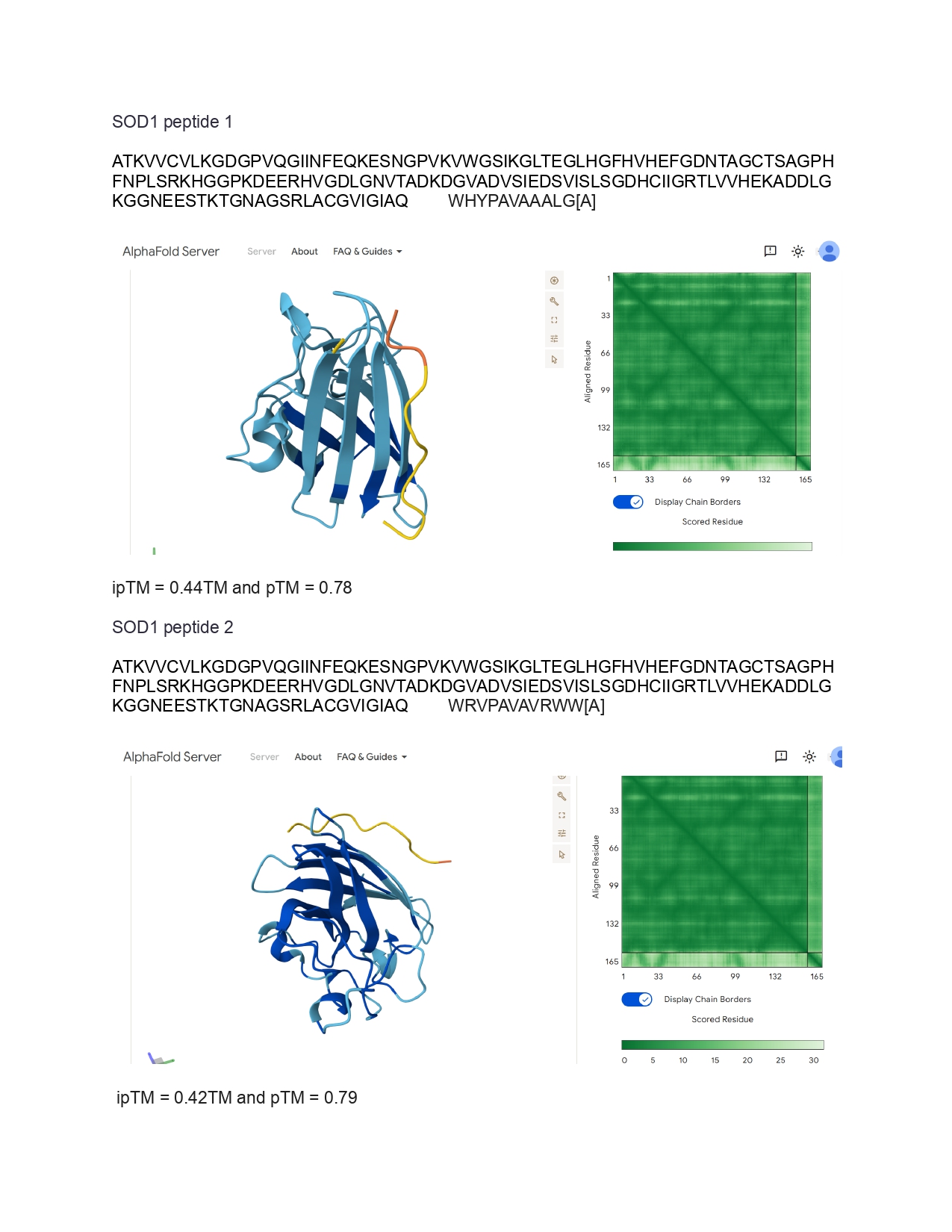
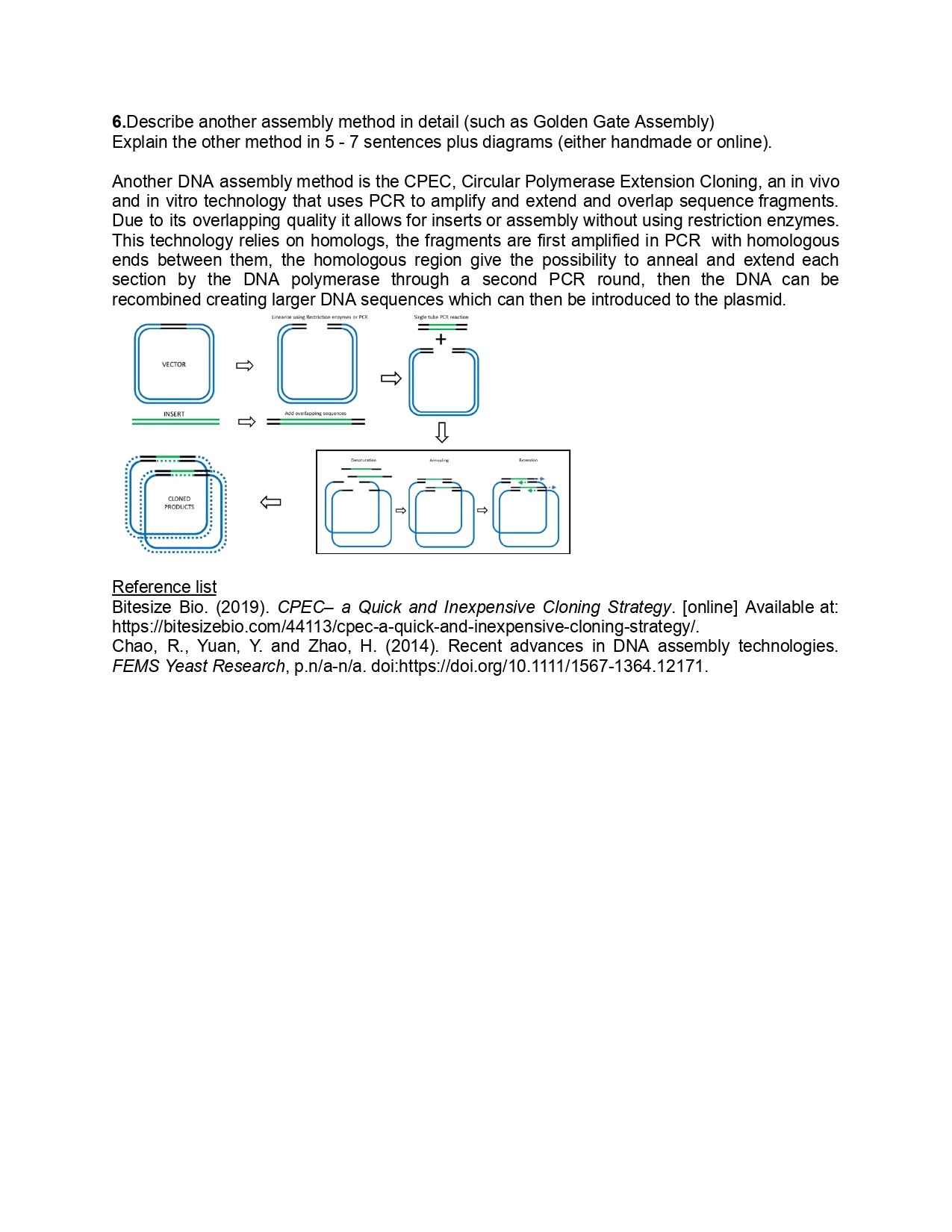
Week 9 HW: Cell Free Systems
















Homework question from Peter Nguyen
1.Write a one-sentence summary pitch sentence describing your concept.
I could use cell free systems adapted to producing inaK in order to directly inoculate glaciers with the aim to preserve and boost ice formation, it would help glaciers rebuild and be more resistant to increasing temperatures caused by climate change. However, I would be interested in pushing the idea of geotextiles already helping preserve glaciers and design a living material, with inaK with a boosted ice nucleation function to create proactive glacier covering actively working to rebuild and preserve glacier ice.
2.How will the idea work, in more detail? Write 3-4 sentences or more.
The inaK would be synthesized through a cell free model, using an alternative to E. coli which could resist and be active in sub-zero conditions (to stay active in glaciers), such as Oleispira antarctica a psychrophiles bacteria which has evolved to have specialized ribosomes and enzymes able to remain flexible and functional in a frozen environment. Oleispira antarctica contains unique chaperone (Cpn60 and Cpn10) preventing protein misfolds in frozen temperatures. This cell design would include pores in the membrane so it can stay alive in the textile by having an ATP source of input.
These then freeze dried cells would be put into a textile (inoculated during the making of the textile), the textile can be brought to location and installed on the glacier and then be rehydrated to allow ice nucleation of the glacier to begin. This would permit me to create a live material that would be dormant in production and transportation and control its freezing function (preventing the textile from accidentally freezing its surrounding).
Note that because I am working with ice nucleation there might be challenges in freeze drying these cells.
Reference List
Ferrer, M. et al. (2003) ‘Low temperature-induced systems failure in Escherichia coli: Insights from rescue by cold-adapted chaperones’, Journal of Biological Chemistry.
Cui, Y. et al. (2022) ‘Cell-free PURE system: evolution and achievements’, Biodesign Research.
D’Amico, S. et al. (2006) ‘Psychrophilic microorganisms: challenges for life’, EMBO reports.
3.What societal challenge or market need will this address?
This addresses the environmental, social and political issue of melting glaciers caused by climate change, which only increases the power of climate change as glaciers are key factors in slowing climate change. We are actively losing biodiversities and ecosystems and doing very little about it. It is not seen as a profitable income so little motivation is inputted. However, in the longterm, this irreversible damage done to our nature will actively make climate change worse, and there will be many destructive environmental, social and economic consequences driven by this overlooked issue.
4.How do you envision addressing the limitation of cell-free reactions (e.g., activation with water, stability, one-time use)?
Working at very large scale, scale of the glacier, it can be a challenge to efficiently rehydrate the living material as it would be very energy consuming and costly to do it manually, but, if the living geotextile is strategically implemented at the right time of the year (early spring, already when they glacier coverings are usually installed) then nature itself through rain could activate the material naturally. The aim is to limit the human labor impact and simply give nature a tool to reinforce what it already knows how to do.
Considering the one time use issue, geotextile coverings which are already used to protect glaciers are removed and installed yearly according to their natural ice melting and forming cycles. The next step of my research would be to find a way to keep the textile created and reabsorb it with new inaK cell free protein systems when it is needed next. The goal is to create a regenerative textile and closed loop system to avoid waste through one time solutions.
Homework question from Ally Huang
1.Provide background information that describes the space biology question or challenge you propose to address. Explain why this topic is significant for humanity, relevant for space exploration, and scientifically interesting. (Maximum 100 words)
I am interested in exploring the purpose of ice nucleation in cell free design, freeze dried or not, taking the shape of a multipurpose textile which can be used as an alternative to current voluminous refrigeration tools or a freezing textile to activate. In space stations like the ISS a lot of research relies on lab samples being preserved in sub zero temperatures, from human research samples to organisms or protein crystals. Within research some experiments need cold induced phase changes to be activated or triggered. This technology could also be used for food or medical supplies.
2.Name the molecular or genetic target that you propose to study. Examples of molecular targets include individual genes and proteins, DNA and RNA sequences, or broader -omics approaches. (Maximum 30 words)
The genetic target of this project would be the inaK ice nucleating protein, commonly found in Pseudomonas syringae, with a wide potential of freezing functions.
3.Describe how your molecular or genetic target relates to the space biology question or challenge your proposal addresses. (Maximum 100 words)
The challenge is optimizing cold packaging and storage systems, a material which would require less space or a material able to be activated once in space again having less constraints in terms of space while travelling. The inaK offers a variety of possibilities in a cell free system whether freeze dried or not as it has a focused and controlled function to freeze. According to the development of the product it can be chosen at what temperature it freezes or activates and how resistant it can be to external temperatures. Creating a highly controlled and bespoke design for certain use in space allows for better control on the research done in space, every aspect of the research can be tailored in hopes to improve success rates of experiments. InaK is a relatively easy INP to work with.
4.Clearly state your hypothesis or research goal and explain the reasoning behind it. (Maximum 150 words)
I am interested in creating polyvalent designs with multiple usages and applications, this project aims to find an optimal alternative refrigerating system which can have bespoke qualities specific to in space research. As small of a detail it might seem every aspect and tool of experiments impacts the result of research and can lead to better efficiency, results or unexpected breakthroughs. During a space mission all equipment has to be optimized due to lack of space and need for many items and a polyvalent tool that can respond to a wide range of uses can help with the space optimization.
5.Outline your experimental plan - identify the sample(s) you will test in your experiment, including any necessary controls, the type of data or measurements that will be collected, etc. (Maximum 100 words)
I would design a cell free system for the inaK ice nucleating protein, freeze dry some and then create living textiles, some active and some dormant. The practicality of a textile is that it can be molded, cut, sewn, layered to adapt to any existing object which would then need a freezing function.
I can control the amount of inaK for the freezing rate needed, experiment with the different temperatures it can freeze at and the different temperatures it can stay frozen at, I can explore the threshold of the inaK.
I would then test the reactivation rates, how much water is needed and how long it would take.
Week 10 HW: Advanced imaging & measurement technology
HTGAA Week 10 Advanced imaging & measurement technology
Final Project
1.Please identify at least one (ideally many) aspect(s) of your project that you will measure. It could be the mass or sequence of a protein, the presence, absence, or quantity of a biomarker, etc.
As for this project I aim to use inaK for ice production I would like to measure the ice nucleation ratio and efficiency of the inaK protein. Additionally, I would like to measure the temperatures inaK can resist to, on its own and as a supplement to an ice sample.
If my initial experiments are successful I would like to measure the inaK ratio innoculated into ice to find the most optimal inaK quantity needed.
2.Please describe all of the elements you would like to measure, and furthermore describe how you will perform these measurements.
To measure the ice production ratio and efficiency of inaK I could use differential scanning calorimetry (DSC) which measures the difference in the amount of heat required to increase the temperature in the sample compared to a reference. It measures the ice nucleation ratio by calculating the enthalpy which is the area below the peak, allowing me to understand precisely how much of the water in the cell is being converted to ice. It measures the efficiency of ice nucleation by creating an exothermic peak ( release of a burst of energy) and analysing how high of a temperature the inaK can still function. This should give me information on thermodynamic efficiency.
I can also measure nucleation temperature through a droplet freezing assay for smaller samples allowing me to test a multitude of potential solution mixes. Here a high speed camera paired with a cooling plate ( a Linkam for example) can allow me to assess how fast a droplet of a solution containing inaK can freeze. Testing this on multiple samples containing different amounts of inaK will give me a spectrum of freezing capacity to find the most optimal ratio of inaK. This experiment could be coupled with an infrared thermography technology which will capture the heat spike and nucleation rate of an inaK and understand how fast the ice nucleation spreads through the cell membrane.
Reference List
Schmid, D. et al. (2016) ‘A high-throughput assay for the characterization of ice-nucleating proteins’, Biophysical Journal. This study outlines the specific use of droplet assays to quantify InaK efficiency.
3.What are the technologies you will use (e.g., gel electrophoresis, DNA sequencing, mass spectrometry, etc.)? Describe in detail.
For these experiments I will use a Differential Scanning Calorimeter for the DSC, a high precision camera, a cooling plate and IR thermography.
Waters Part 1 - Molecular Weight
For this section I used a combination of the tools provided, my knowledge and AI assistance as I have trouble understanding math related work.
- Based on the predicted amino acid sequence of eGFP and any known modifications, what is the calculated molecular weight ?
According to Expasy I found that this sequence has a theoretical pI/Mw of 5.90 / 28006.60.
To calculate the molecular weight of this sequence I referred to the standard isotopic mass of amino acids and subtracting H²O for each peptide bond. For this amino acid sequence I found that :
-eGFP of 238 AA = 26.735.6 Da
-LE Linker of 2 AA = 242.3 Da
-x-His Tag of 6 AA = 822.8 Da
Resulting in a total molecular weight of 27800.7 Da
- Calculate the molecular weight of the eGFP using the adjacent charge state approach described in the recitation. Select two charge states from the intact LC-MS data (figure 1) and:
MW : molecular weight in Daltons
m/z : value of the peak on the x-axis of the spectrum
z : integer charge state of said peak
H+ : mass of a proton
a.Determine z for each adjacent pair of peaks $(n, n+1)$ using: $$ {\large z} = {\Large \frac{\frac{m}{z_{n+1}}}{\frac{m}{z_n} - \frac{m}{z_{n+1}}}} $$
Here I am using the adjacent peaks at 875.4421 and 903.7148.
m:zn = 875.4421
m:zn+1 = 903.7148
The lower the m/z the higher the charge versus the higher the m/z the lower the charge.
Using the provided formula
z = 903.7148 : (903.7148 – 875.4421) = 903.7148 : 28.2727 = 31.96
Charge states must be integers and the charge state for the peak at 903.7148 is z= 31
So, the peak at 875.4421 has a charge state of z+1=32
b.Determine the MW of the protein using the relationship between $\frac{m}{z_n}$, $MW$, and $z$
The base equation for the peaks in this figure is
m:z = (MW + (z x H+)) : z
Here I rearrange the formula to find MW
MW = z x (m:z) - (z x H+)
If z=31, then,
MW = 31 x(903.7148) - (31x1.008)
MW = 28 015.1588 – 31.248
MW = 27 983.91 Da
c.Calculate the accuracy of the measurement using the deconvoluted MW from 2.2 and the predicted weight of the protein from 2.1 using: $$ \text{Accuracy} = \frac{|MW_{\text{experiment}} - MW_{\text{theory}}|}{MW_{\text{theory}}} $$
Using the provided formula and substituting the values accordingly,
If,
Accuracy = (MW experimental - MWtheory) : MW theory
Then,
Accuracy = (27 982.90 - 27 782.70) : 27 782.70
Accuracy = 200.20 : 27 782.70
Accuracy = 0.0072
=0.72%
3.Can you observe the charge state for the zoomed-in peak in the mass spectrum for the intact eGFP? If yes, what is it? If no, why not?
One can observe the charge state in the zoomed in peak but not using the previous calculation method using the adjacent peak method used for the entire spectrum.
The zoomed in area presents an isotopic cluster of one charge state, therefore, the observation is focused on isotopic resolution. The other peaks represent different amounts of protons.
So, here, yes I can see the charge state using isotopic resolution because the peaks are distinct and separate and the instrument has high enough of a resolution to expose the isotopes separately. It is because the instrument used here is of high precision that we are able to have high resolution, if a lesser precise instrument were used then it would be very difficult to see the charge state.
The formula to calculate isotopic spacing here would be
z = 1 : Δm/z
Waters Part 2 - secondary & tertiary structure
1.Please explain the difference between native and denatured protein conformations. For example, what happens when a protein unfolds? How is that determined with a mass spectrometer? What changes do you see in the mass spectrum between the native and denatured protein analyses (figure 2)?
There is a structural difference between the native and the denatured protein conformations, the native protein conformation is a tightly folded protein, a unique three dimensional structure unique to its biological environment. The structure is compact but is held by weak non-covalent bonds. In contrast, a denatured protein conformation is the unfolded structure once the weak bonds are broken, the structure is flexible and unpredictable resulting in a random coil shape. The denatured protein no longer has a function compared to the native protein conformation.
A mass spectrometer scans the protein shape by measuring its mass and charge rather than measuring the shape directly.
The top green graph corresponds to the denatured confirmation and the bottom red chart corresponds to the native conformation.
The denatured conformation graph shows crowded peaks in the 500 to 1200 m/z range, these are low m/z values. A lower m/z equals a higher z charge and when a protein is unfolded into a random coil then the basic residues are exposed to a proton rich environment and can easily be protonated creating a protein with a very high net positive charge.
The native conformation has larger m/z values with peaks at 2545 and 2799, the higher the m/z value the lower the z charge. When the protein is still tightly folded the basic residues are buried in the hydrophobic core protecting it from its environment meaning protons cannot impact them so the protein is less protonated.
2.Zooming into the native mass spectrum of the eGFP from the Waters Xevo G3 QTof or MS (figure 3), can you discern the charge state of the peak at ~2800? What is the charge state? How can you tell?
It is possible to discern the charge state of the peak at ~2800 m/z because instrument provides a high resolution even if the image focuses on the 2525 m/z peak, we are still able to see the isotopic distribution by measuring the distance between the individual isotopes and we would be able to calculate the charge state.
Measuring the charge state of the peak at ~2800 m/z using isotopic spacing,
In a mass spectrum the individual isotopes of the same molecule differs by approximately 1 Dalton,
The distance between the isotopes here on the x-axis Δm/z can be calculated using the following formula,
Δm/z = 1: z
Zooming in at the 2799.4199 peak same as for the 2545 peak (with a peak separation of precisely 0.1 m/z) then,
z = 1: 0.1 = 10
So the charge state of the peak at ~2800 m/z is +10
Waters Part 3 - Peptide mapping, primary structure
1.How many Lysines (K) and Arginines (R) are in eGFP? Please circle or highlight them in the eGFP sequence given in Waters Part I question 1 above. (Note: adding the sequence to Benchling as an amino acid file and clicking the biochemical properties tab will show you a count for each amino acid).


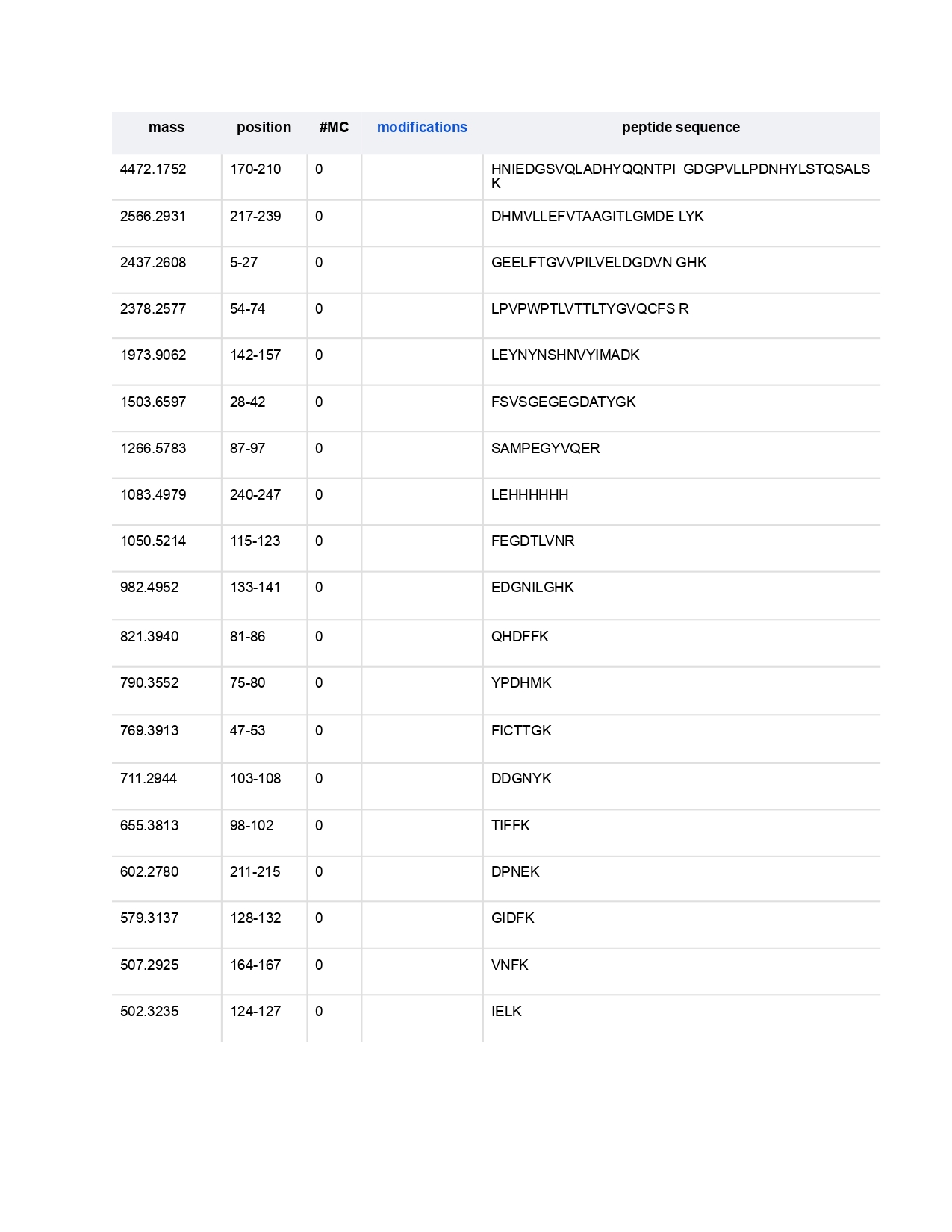
2.How many peptides will be generated from tryptic digestion of eGFP?
I found 19 peptides were generated from this sequence using the trypsin.

3.Based on the LC-MS data for the Peptide Map data generated in the lab ( please use Figure 5a as a reference) how many chromatographic peaks do you see in the eGFP peptide map between 0.5 and 6 minutes? You may count all peaks that are>10% relative abundance.
Between 0.5 and 6 minutes there are 14 distinct peaks above 10% relative abundance in this figure.
4.Assuming all the peaks are peptides, does the number of peaks match the number of peptides predicted from question 2 above? Are there more peaks in the chromatogram or fewer?
In comparison to the amount of predicted peptides, 19, there are fewer peptides in the chromatogram.
5.Identify the mass-to-charge (m:z) of the peptide shown in Figure 5b. What is the charge (z) of the most abundant charge state of the peptide (use the separation of the isotopes to determine the charge state). Calculate the mass of the singly charged form of the peptide ([M+H]+) based on its m:z and z.
The z charge of the most abundant peak in this peptide is m/z = 525.76712
To calculate the charge state,
First isotope peak 525.76712
Second isotope peak 526.25918
Calculating the spacing,
(Δm/z):526.25918 - 525.76712 = 0.49206
Using the formula,
z = 1: 0.49206≈2.03
So the charge state z of the most abundant peak of the peptide is of +2
Calculating the mass of the singly charged form of the peptide ([M+H]+),
First I need to calculate the neutral mass M using M=zx(m/z)-(zx1.00727),
1.00727 Da is the mass of a proton,
So,
M = 2 x (525.76712) - (2x1.00727)
M = 1051.53424 - 2.01454 = 1049.5197 Da
([M+H]+) can be calculated by adding one proton mass back to the neutral mass,
[M+H]+ = 1049.5197 + 1.00727 = 1050.5270 Da
6.Identify the peptide based on comparison to expected masses in the PeptideMass tool. What is mass accuracy of measurement? Please calculate the error in ppm. (Recall that Accuracy = (MW experimental - MWtheory) : MW theory)
To identify the peptide I consider that the experimental neutral molecular sight (MWexperimental) was calculated for the peak 525.76712 m/z and equaled MW: 1049.5197 Da, I will use the following peptide sequence as a theoretical tryptic digest as a comparison, LPDNHYLSTQSALSK, and considering theoretical MW (MWtheory) : 1049.5393 Da. This peptide corresponds to the residues 139–153 of the eGFP protein.
To calculate the mass accuracy, error in ppm (parts per million) I will use the following formula,
Accuracy (ppm) = ((MWexperimental - MW theory) : MWtheory) x 106
Now adding the values,
MW experimental = 1049.5197
MW theory = 1049.5393
Accuracy (ppm) = ((1049.5197 - 1049.5393) : 1049.5393) x 106
Accuracy (ppm) = (-0.0196) : 1049.5393) x 106
Accuracy (ppm) = –18.67ppm
7.What is the percentage of the sequence that is confirmed by peptide mapping ? (see figure 6)
The percentage of the sequence that is confirmed by peptide mapping seems to be indicated at 88%, the blue highlighted areas are confirmed amino acids in the sequence.
Waters Part 4 - Oligomers
We will determine Keyhole Limpet Hemocyanin (KLH)’s oligomeric states using charge detection mass spectrometry (CDMS). CDMS single-particle measurements of KLH allow us to make direct mass measurements to determine what oligomeric states (that is, how many protein subunits combine) are present in solution. Using the known masses of the polypeptide subunits (Table 1) for KLH, identify where the following oligomeric species are on the spectrum shown below from the CDMS (Figure 7):
-7FU Decamer
-8FU Didecamer
-8FU 3-Decamer
-8FU 4-Decamer
I will first calculate the theoretical mass for the species using the given measures 7FU = 340 kDa and 8FU = 400 kDa. The axis of the spectrum is in MDa (megadaltons) where 1MDa = 1000 kDa.
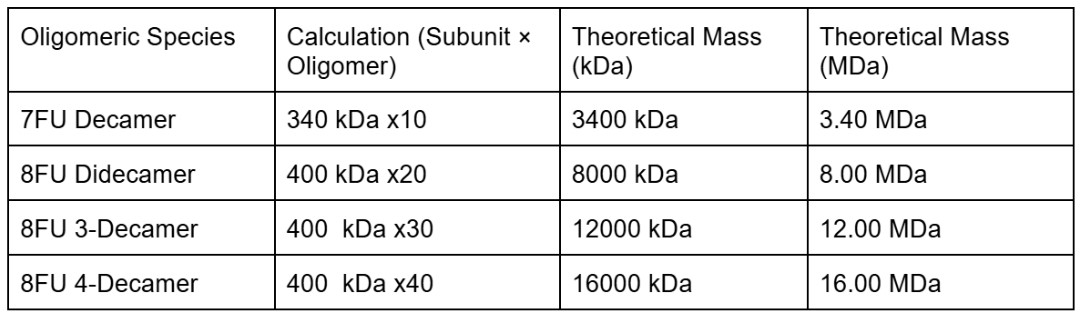

I can then identify the peaks on the spectrum
-7FU Decamer ≈ 3.40 MDa is the peak labeled 3.4 on the spectrum
-8FU Didecamer ≈ 8.00 MDa correspond to the peak labeled 8.33
-8FU 3-Decamer ≈ 12.00 MDa matches the peak labeled 12.67
-8FU 4-Decamer ≈ 16.00 MDa would appear in the cluster of small unlabeled peaks around 16 to 17 MDa on the far right side of the chart
Waters part 5 - Did I make GFP?
- Please fill out this table with the data you acquired from the lab work done at the Waters Immerse Lab in Cambridge, or else the data screenshots in this document if you were unable to have lab work done at Waters.


Week 11 HW: Building genomes
Week 11 Bioproduction & Cloud Lab
Part A - The 1.536 pixel art work canvas, collective artwork
1.Contribute at least one pixel to the global artwork
I added early on a pixel towards the top left corner. I do not have much to say about this section of the work except maybe understanding the full purpose of this exercise.
Part B - Cell Free protein synthesis, cell free reagents
- Referencing the cell-free protein synthesis reaction composition (the middle box outlined in yellow on the image above, also listed below), provide a 1-2 sentence description of what each component’s role is in the cell-free reaction.
E. coli Lysate
- BL21 (DE3) Star Lysate (includes T7 RNA Polymerase) : offers the base molecular machinery such as ribosomes, tRNAs and enzymes for translation, the Star Lysate strain reduces mRNA degradation, and, the T7 Polymerase drives high level transcription from T7 promoters
Salts / Buffer
- Potassium Glutamate : primary salt that maintains ionic strength and provides potassium ions essential to ribosomal function and protein to nucleic acid exchange
- HEPES-KOH pH 7.5 : chemical buffer which helps maintain a stable physiological pH which affects enzymatic function of the transcription and translation machinery
- Magnesium Glutamate : magnesium ions are vital contributors to stabilizing the ribosome structure and enabling catalytic activity of the polymerases kinases
- Potassium phosphate, monobasic and dibasic : functions as a secondary pH buffer and a source of inorganic phosphate essential for the regeneration of high energy molecules such as ATP
Energy / Nucleotide system
- Ribose : serve as a carbon backbone precursor for the synthesis of nucleotides, allowing for regeneration of NTPs essential for transcription and energy transfer
- Glucose : primary metabolic energy source fueled through glycolysis allowing to regenerate the ATP and GTP essential to the good functioning of protein synthesis
- AMP / CMP / UMP : offers nucleotide building blocks for RNA synthesis and can be converted into triphosphate such as ATP, CTP, UTP needed in transcription
- GMP : from the lack of GMP might demonstrate a dependency on salvage pathways to generate GTP essential to translation
- Guanine : precursor for GMP/GTP synthesis through salvage pathways helpful to RNA synthesis and ribosomal function
Translation Mix (Amino acids)
- 17 Amino Acid Mix : provide the base building blocks to synthesize the polypeptide chain
- Tyrosine : supplied separately because of its solubility limitations, becomes an essential building block for protein synthesis once it is adapted into a usable form
- Cysteine : added separately due to its oxidation limitations, it is an essential compound in forming disulfide bonds in proteins
Additives
- Nicotinamide : serves as a precursor for NAD+ / NADH synthesis reinforcing redox balance and metabolic reactions occurring in energy regeneration
Backfill
- Nuclease Free Water : is used to adjust all the components to the desired the final reaction volume while it avoids degradation of DNA / RNA by nuclease and ensures stable transcription and translation processes
- Describe the main differences between the 1-hour optimized PEP-NTP master mix and the 20-hour NMP-Ribose-Glucose master mix shown in the slide.
The main difference between the two master mix results in found in the energy and nucleotide sourcing a the 1-hour mix makes use of the PEP and pre-synthesized NTPs for instant and high burst protein synthesis compared to the 20-hour mix uses the ribose, glucose and NMPs as precursors to regenerate energy and nucleotides throughout time. Therefore, the 1-hour mix is designed for speed and rapid prototyping in contrast to the 20-hour mix allows to better optimize the cost for effectiveness by using the Lysate’s metabolic pathway to support the reaction for an extended period of time.
Part C - Planning the global experiment, cell-free master mix design
1.Given the 6 fluorescent proteins we used for our collaborative painting, identify and explain at least one biophysical or functional property of each protein that affects expression or readout in cell-free systems. (Hint: options include maturation time, acid sensitivity, folding, oxygen dependence, etc) (1-2 sentences each)
- sfGFP : provides robust and rapid protein folding therefore the protein is less likely to aggregate enabling it to offer a strong fluorescent readout even if fused to complex proteins
- mRFP1 : is a protein with a longer maturation time signifying the fluorescence develops slower after translation and might have a delayed signaling time in shorter experiments, it has a low acidity tolerance
- mKO2 : is fast maturing and has a relative acidity tolerance, meaning the fluorescence will be less visible in a lower pH context but the fluorescence could increase in longer cell free reactions
- mTurqoise2 : is a cyan protein with for a high quantum yield and high photostability making the fluorescence outread a great signal no matter the length of the reaction, it is very sensitive to pH
- mScarlet_I : engineered for fast maturing and high brightness allowing for stronger fluorescence signals compared to older red proteins
- Electra2 : protein engineered for very fast maturation making it very useful for fast consuming energy systems in an experiment where a rapid output is needed before the mix’s energy is used up
2.Create a hypothesis for how adjusting one or more reagents in the cell-free mastermix could improve a specific biophysical or functional property you identified above, in order to maximize fluorescence over a 36-hour incubation. Clearly state the protein, the reagent(s), and the expected effect.
Could the mTurquoise2 yield be increased or accelerated through pH stabilisation.
The reagents HEPES-KOH would be increased to 100mM and Potassium Phosphate would be increased to 15mM.
This adjustment should increase the capacity of the buffer within the mater mix and should neutralize organic acid by products such as lactate and acetate generated during the 36-hour metabolism of glucose and ribosome. Because mTurquoise2 is very reactive to pH, preserving the pH environment at 7.5 would prevent the typically occurring rapid cooling of the cyan signal which usually occurs as the mix acidifies over time. Therefore, the high quantum of yield of mTurquoise2 is complete and optimized leading to a bright and stable cyan readout which won’t dim as the energy levels decrease.
3.The second phase of this lab will be to define the precise reagent concentrations for your cell-free experiment. You will be assigned artwork wells with specific fluorescent proteins and receive an email with instructions this week (by April 24). You can begin composing master mix compositions here.
- The final phase of this lab will be analyzing the fluorescence data we collect to determine whether we can draw any conclusions about favorable reagent compositions for our fluorescent proteins. This will be due a week after the data is returned (date TBD!). The reaction composition for each well will be as follows:
- 6 μL of Lysate
- 10 μL of 2X Optimized Master Mix from above
- 2 μL of assigned fluorescent protein DNA template
- 2 μL of your custom reagent supplements
Total : 20 μL reaction