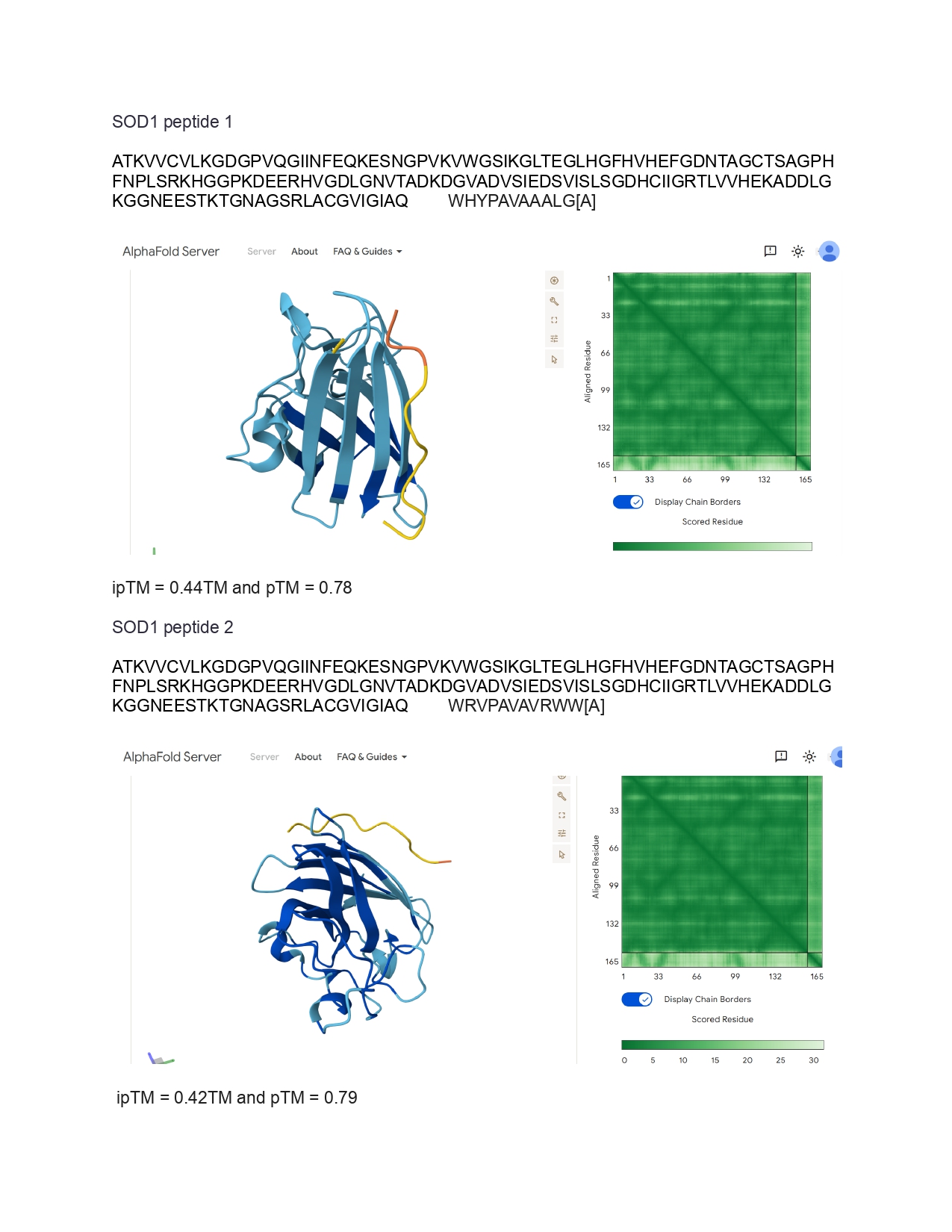
Individual Final Project
iKe
Abstract
Mountain glaciers are melting progressively due to climate change and human activity. I am inspired by the glaciers of the Italian Alps where my family is from and throughout generations have seen firsthand the glaciers progressively disappear. Glaciers are vital ecosystems which contribute to protecting nature and human existence.
I aim to use my knowledge in textiles, biology and biodesign to help preserve and rebuild glaciers using ice nucleating proteins (INPs).
I theorise that inoculating glaciers with modified INPs using cell free synthesis would help improve ice catalyzation and make glacier ice more resilient to face increasing temperatures. This method will work as a defensive tool helping restore natural and healthy glacier cycles benefiting a wider ecosystem and battling climate change.
Project overview
Growing up I spent a lot of time in the Italian Alps where part of my family originates from, within only 23 years of life I have seen first hand the glaciers of mountains surrounding me disappear progressively, my mother sees an even bigger decrease and my grandfather a shocking decrease. With my background in textiles, my current studies in Biodesign, my curiosity for biology and now partaking in HTGAAA I will conceptualise a project combining textiles and biology as a means to create a tool which could help prevent the fast disappearance of glaciers.
Glaciers are vital elements to regulating the earth’s temperature, they are ecosystems of their own across the world. As pointed out by Glacier Preservation, “glaciers are essential for sustaining millions of people by providing fresh water, supporting hydropower generation, and playing a key role in environmental stability. However, as climate change accelerates, glaciers are retreating at an unprecedented rate, threatening water security, energy infrastructure, and increasing natural hazards like flooding and avalanches.” (Un-glaciers.org. (2026). Glacier Preservation is the Key to Ensuring the Security of Water, Energy, and Environmental Resources. [online] Available at: https://www.un-glaciers.org/en/articles/glacier-preservation-key-ensuring-security-water-energy-and-environmental-resources.).
My project aims to ethically preserve glaciers and help them naturally rebuild while respecting their natural cycles and having minimal interference.
I want to work with ice nucleating proteins, specifically the inaK strands, they are commonly found in nature and have the function to catalyze ice. These INPs should be used to improve ice formation in glaciers and sustain ice levels while they face a rise in temperatures.
I aim to work using a cell free synthesis method, potentially improve the inaK function through gene mutation and through different levels of test innoculate glaciers with inaK.
Aim 1 synthesize inaK ice nucleating protein (INP)
- Using UniProt source records (O30611) the ice nucleating protein DNA sequence of inaK (found in Pseudomonas Syringae - pathogenic)
- Refine DNA sequence through codon optimization using Twist
- Using Benchling and Twist create a plasmid for the inaK INP sequence
- Attempt cell free protein synthesis for inaK INP. This method provides more stability and less toxicity. Using one of the following methods
- Detergent process
- Liposome process
- Nanodisc process
Aim 2 Further development of INPs in glaciers
- Consider genetically modifying inaK in order to increase ice catalyzation capacities.
- Use peptide motifs to improve the ice catalyzation function, enhancing or stabilizing the ice binding surface through crystal lattice. INPs like inaK a large and repetitive proteins, using peptide motifs allows to focus the INP.
- Tandem N-terminal repeats: improve sequence stability enabling to boost ice nucleation
- Aline rich helical peptides: freezing at higher temperatures due to aggregation of high concentration of aline rich peptides
- TxT motif : uses hydroxyl to anchor water
- SLT motif : stabilizes the organisation of water interface
- Y motif: helps align water molecules
Could this method boost ice formation further without interrupting the natural cycle of glacier ice melting and rebuilding?
Aim 3 Future INP output reinforcing glacier ice formation
- Introduce successfully synthesized protein into a living material inspired by existing geotextile
- Observe if protein is able to stay alive. Observe how it may catalyse ice independantly
- Place living material over a sample of ice and observe the effect of the INP on ice. Is there an increase in ice formation?
- Hopefully obtain sample of a glacier and test the success rate of glacier inoculation of inaK.
- Introduce technology to glaciers
- If this technology allows glaciers to rebuild stronger and be less prone to melting this would have a significant positive impact on reversing climate change
Note that if CFPS fails, then attempt in vivo synthesis and introduce inaK to new host, Pseudomonas Fluorescens, which does not contain any ice-nucleotides, already exist within glaciers and presents no risk to nature. Transcribing new DNA data from one Pseudomonas to another will be either as they are part of the same species. Additionally, because Pseudomonas Fluorescens already exist within glaciers it facilitates horizontal gene transfer but also avoids disrupting the natural balance and ecosystem of the glaciers.
Within this project I will also focus a section of my research in ethics, analyzing the impact my work could have on glaciers and their ecosystems. I aim to be educated about environmental laws and bioethics.
I recognise that within the context of this project I would need the expert knowledge of a glaciologue able to provide me with specific and reliable data to use within my research which would allow me to work as ethically, sustainably and durably as possible.
Literature review
- Existing methods for glacier preservation
Start by reviewing current glacier-preservation approaches and their limitations. This gives the project a real-world anchor and shows why controlling ice nucleation could matter.
There are a few ongoing projects which aim to protect glaciers. Mainly they appear in the form of geotextile covering.
GlacierProtect by Naue is a glacier protection project through geotextiles, it uses sustainable raw materials to reflect up to 75% of sunlight. Glaciers already have the capacity to reflect sunlight but this new textile supports this natural ability which preserves the glaciers and by preserving glaciers will enable them better form and themselves better reflect the sunlight. This project inspires me as a way of demonstrating these innovative projects are scalable but also that true regenerative design can be achieved with a mindset of supporting glaciers to promote their rebuild and growth where in turn they could thrive and function as they were meant to.
Ponte di Legno Tonale glacier project with a similar technology to Naue, the Presena glacier has been protected since 2008 after observing significant damage to the glacier in 2000 by “covering Presena glacier with geotextile fabric covers, which are able to reduce by 50% the melting of snow and ice during summer months” (Ponte di Legno Tonale, n.d.). This project demonstrating chronological progress shows the real potential and benefit of protecting glaciers and assisting them to rebuild naturally.
Furthermore, many companies are innovating and making use of ice nucleating proteins in snow canons to increase production of snow. Snowmax International has made great progress in this direction by using INP to make water free at higher temperatures because of a higher concentration of nuclides, increase ice catalyzation speed as it attaches to nuclides, and be more resistant in time as it makes use of less water therefore reducing evaporation capacity and due to its smaller crystal structures from stronger iconic bonds. According to GoldBio, “P. syringae’s ice-nucleating activities have long been used to make artificial snow. Products including Snomax® use the proteins derived from the outside of bacteria to enhance the snow generated by snow blowers. One study showed that Snomax® increases the amount of snow made by a snow blower by as much as 90% (Snomax® International, 2013)” (Christner et al., 2008) leading to my next point of research.
While this is mostly destined to ski track upkeep and not oriented to ice production it demonstrates that INPs are being used on a larger scale and prove to be effective. These technologies are also proving that they have no negative repercussions on the environment as they make use of all natural compounds.
Pseudomonas syringae is a bacteria which contains ice nucleating protein strands which have the ability to catalyze the formation of ice in sub zero temperatures. The main issue with pseudomonas syringae is that it is a pathogen in agriculture as it infects crops. However, would it be possible to extract the ice nucleating protein strands from the bacteria, use pseudomonas syringae as a model to synthesize ice nucleating protein or genetically modify the bacteria to neutralize its harmful effect to plants in order to use this genetically modified or synthesised bacteria as an ice forming agent to help preserve glaciers.
Instant ice packs are innovations of modern medicine where a sealed pouch of water is activated through an endothermic chemical reaction transforming surrounding heat into ice. Would it be possible to upscale this chemical reaction or let it inspire some biomimicry which through textiles and biology could use the increasingly hot temperature of the earth to transform it into ice on the surface of glaciers to help preserve them.
Snow Seeds by Tanay Wadokar is a project presented by a 2025 graduate of the MA Materials Future at Central Saint Martins (UAL, London). Tanay Wadokar created snow board stickers with cloud seeding technology. This would allow the deposit of ice nuclei while snowboarding which would enable locals and tourists to enjoy the inter sport while preserving mountains and glaciers by regenerating snowfalls. It is not about limiting the life we know today but rather by shaping it in a sustainable proactive way.
Reference list
Arnold, D.L. and Preston, G.M. (2019). Pseudomonas syringae: enterprising epiphyte and stealthy parasite. Microbiology, 165(3), pp.251–253. doi:https://doi.org/10.1099/mic.0.000715.
Boztas, S. (2024). Pumped up: will a Dutch startup’s plan to restore Arctic sea-ice work? The Guardian. [online] 27 Feb. Available at: https://www.theguardian.com/environment/2024/feb/27/climate-crisis-arctic-ecosystems-environment-startup-plan-pump-restore-melting-sea-ice-caps.
Christner, B.C., Morris, C.E., Foreman, C.M., Cai, R. and Sands, D.C. (2008). Ubiquity of Biological Ice Nucleators in Snowfall. Science, 319(5867), pp.1214–1214. doi:https://doi.org/10.1126/science.1149757.
Dr. Tobias Weidner, Dr. Janine Fröhlich-Nowoisky (2016). The effect of bacterial ice nuclei. [online] Www.mpg.de. Available at: https://www.mpg.de/10470442/ice-formation-bacteria-syringae.
James Dalton, Global Head, Water and Wetlands Team, IUCN (2025). Protecting glaciers – our most effective natural water manager. [online] IUCN. Available at: https://iucn.org/blog/202503/protecting-glaciers-our-most-effective-natural-water-manager.
McDonough, F. (n.d.). What is Cloud Seeding? [online] Desert Research Institute. Available at: https://www.dri.edu/cloud-seeding-program/what-is-cloud-seeding/.
Ponte di Legno Tonale. (n.d.). The protection of Presena Glacier. [online] Available at: https://www.pontedilegnotonale.com/en/pontedilegno-tonale-what-to-see/the-protection-of-presena-glacier/.
Roeters, S.J., Golbek, T.W., Bregnhøj, M., Drace, T., Alamdari, S., Roseboom, W., Kramer, G., Šantl-Temkiv, T., Finster, K., Pfaendtner, J., Woutersen, S., Boesen, T. and Weidner, T. (2021). Ice-nucleating proteins are activated by low temperatures to control the structure of interfacial water. Nature Communications, [online] 12(1), p.1183. doi:https://doi.org/10.1038/s41467-021-21349-3.
Snomax.com. (2015). FAQ - Snomax. [online] Available at: https://www.snomax.com/faq.html.
Steroplast Healthcare (2022). How Do Instant Ice Packs Work? [online] www.steroplast.co.uk. Available at: https://www.steroplast.co.uk/knowledge-base/how-do-ice-packs-work.html.
Un-glaciers.org. (2026). Glacier Preservation is the Key to Ensuring the Security of Water, Energy, and Environmental Resources. [online] Available at: https://www.un-glaciers.org/en/articles/glacier-preservation-key-ensuring-security-water-energy-and-environmental-resources.
Wadokar, T. (2025). Snow Seeds - Tanay Wadodkar - UAL Showcase. [online] Arts.ac.uk. Available at: https://ualshowcase.arts.ac.uk/project/635735/cover [Accessed 9 Feb. 2026].
Wellpott, V. and Wellpott, V. (2025). Glacier protection with geotextiles – A sustainable solution for the future. [online] Naue - Geosynthetics | Digtal Engineering Software | Installation services. Available at: https://www.naue.com/glacier-protection-with-geotextiles-a-sustainable-solution-for-the-future/.
- Fundamentals of ice nucleation
Explain the basic mechanism of ice nucleation, including:
- homogeneous vs heterogeneous nucleation
- the temperatures typically required
- why nucleation at higher subzero temperatures is valuable
Ice nucleation is a mechanism where water molecules transform from a liquid state to a solid crystalline lattice, an existing nucleus or template needs to surpass the energy barrier of crystallization.
In a pure water sample, containing no foreign particles, water molecules can spontaneously collide and form stable crystals, this is homogeneous nucleation. In contrast a heterogeneous nucleation is when a foreign compound, an ice nucleator, forms a physical surface or template for the water molecules to attach themselves to which significantly lowers the energy needed for freezing. In the case of Pseudomonas syringae the INPs provide a heterogeneous surface for ice to form itself on.
Ice nucleation temperature will depend on the context in which ice nucleation is occurring, typically for pure water samples can stay liquid till -40°C in a supercooled state before freezing. With common nucleators such as minerals the water sample can freeze at about -10°C to -15°C. In the case of the inaK INP which I am studying, the ice catalyzation can be triggered at -2°C to -5°C, demonstrating the potential of using inaK compared to other INP. Being able to catalyze ice at higher subzero temperatures shows potential for environmental protection, as seen in the Snowmax project, helping to produce snow at higher temperatures, essential for protecting glaciers and reducing the ice loss. Additionally, they can have an atmospheric impact, because INPs are so effective they can truly affect weather patterns, similarly to cloud seeding technology they can trigger cloud formation and influence precipitation at warmer altitudes. For companies such as Snowmax the ice nucleation technologies allow them to produce the same amount of snow using less energy than they would need at regular ice nucleation temperature of -10°C or -15°C as machines don’t need to compensate for the temperature.
Reference List
Biotreks (2021) ‘Ice nucleation proteins – a synthetic pathway to alleviate ice loss’, Biotreks, (e202111).
GoldBio (n.d.) The Extraordinary Bacterial Proteins That Make Snow. Available at: https://www.goldbio.com/blogs/articles/the-extraordinary-bacterial-proteins-that-make-snow (Accessed: 25 April 2026).
Hudait, A. et al. (2024) ‘Hierarchical assembly and environmental enhancement of bacterial ice nucleators’, Proceedings of the National Academy of Sciences (PNAS), 121(18), p. e2409283121. doi: 10.1073/pnas.2409283121.
Pandey, R. et al. (2016) ‘Ice-nucleating bacteria control the order and dynamics of interfacial water’, Science Advances, 2(4), p. e1501630. doi: 10.1126/sciadv.1501630.
Roeters, S. J. et al. (2024) ‘Polyol-Induced 100-Fold Enhancement of Bacterial Ice Nucleation Efficiency’, The Journal of Physical Chemistry C, 128(15). doi: 10.1021/acs.jpcc.4c07422.
Schmid, D. (2026) Glacier Blankets Could Help Prevent Melting. [Online Video]. 25 April. Available at: https://www.youtube.com/watch?v=hKT_SGK2qtY (Accessed: 25 April 2026).
USNSJ (n.d.) ‘Wonders of the Invisible World: Pseudomonas syringae, the Ice Maker’, University of Southern North Science Journal, 2(2).
Experimental Data Figure (2026) Intact Mass Spectra: Native vs. Denatured sfGFP Analysis.
Experimental Data Figure (2026) Sequence Coverage Report: 88% Confirmation of sfGFP.
Experimental Data Figure (2026) CDMS Spectrum of KLH Oligomeric States.
- Types of ice-nucleating particles
Summarize the major categories of ice-nucleating particles:
- mineral
- biological
- organic/macromolecular
- engineered materials
Then position biological ice nucleators as the most relevant class for your work.
Different types of ice nucleation particles exist, they can be organized in categories.
The mineral particles which are mostly inorganic atmospheric aerosols like minerals and soot. They are the most common INPs in the atmosphere but are not considered very efficient as they require the very low temperatures of -10°C to -15°C to initiate heterogeneous nucleation.
The biological ice nucleators which consist of bacteria such as the Pseudomonas syringae containing inaK. They are the most effective ice nucleators as they can trigger ice at higher temperatures, between -2°C and -5°C. Like the inaK protein they are usually very efficient due to their repetitive genetic code improving crystal lattice capability.
Organic and macromolecular compounds which refer to non-living organic substances. Polyols are an example of macromolecular compounds which are not always ice nucleators but can improve the efficiency of existing biological nucleators by up to 100 fold.
Engineered materials, however, are synthetic substances designed to imitate natural ice nucleation. Silver iodide for instance is a synthetic protein used in cloud seeding.
Biological nucleators such as inaK are the most relevant for my work as, on one hand, they prove to be more efficient for ice nucleation at higher sub zero temperatures which will be essential when working on glaciers confronted with rising temperatures. On the other hand, they have a better chance of non disruptive inoculation into glaciers as they are more likely to assimilate naturally to the existing biological nucleators already present in glaciers.
Reference list
Biotreks (2021) ‘Ice nucleation proteins – a synthetic pathway to alleviate ice loss’, Biotreks, (e202111).
GoldBio (n.d.) The Extraordinary Bacterial Proteins That Make Snow. Available at: https://www.goldbio.com/blogs/articles/the-extraordinary-bacterial-proteins-that-make-snow (Accessed: 25 April 2026).
Hudait, A. et al. (2024) ‘Hierarchical assembly and environmental enhancement of bacterial ice nucleators’, Proceedings of the National Academy of Sciences (PNAS), 121(18), p. e2409283121. doi: 10.1073/pnas.2409283121.
Lukas, M. et al. (2025) ‘A New Class of Fungal Ice-Nucleating Proteins with Bacterial Ancestry’, ChemRxiv. doi: 10.26434/chemrxiv-2025-73058.
Pandey, R. et al. (2016) ‘Ice-nucleating bacteria control the order and dynamics of interfacial water’, Science Advances, 2(4), p. e1501630. doi: 10.1126/sciadv.1501630.
Roeters, S. J. et al. (2024) ‘Polyol-Induced 100-Fold Enhancement of Bacterial Ice Nucleation Efficiency’, The Journal of Physical Chemistry C, 128(15). doi: 10.1021/acs.jpcc.4c07422.
Schmid, D. (2026) Glacier Blankets Could Help Prevent Melting. [Online Video]. 25 April. Available at: https://www.youtube.com/watch?v=hKT_SGK2qtY (Accessed: 25 April 2026).
USNSJ (n.d.) ‘Wonders of the Invisible World: Pseudomonas syringae, the Ice Maker’, University of Southern North Science Journal, 2(2).
- how ice-nucleating proteins classified?
- by source class (bacterial, fungal, plant, and insect)
- by activity type (Class A, B, C by nucleation temperature / aggregate size)
- by protein family/named variants (InaZ, InaK, InaV, InaQ, InaW, etc.)
Ice nucleating proteins are classified according to biological origin, physical assembly of the cell membrane and specific genetic variants.
Source class classification identifies INPs found across organisms and how they utilize their protein for different ecological advantages. The bacterial class is the most studied and their main function is to facilitate precipitation or cause frost damage on plants causing various nutrients to release, the most common bacteria is the Pseudomonas syringae which is recognized as pathogenic to nature. The fungal class is a newly recognized class usually sharing bacterial ancestry but can have different structural arrangements. The plant source is used to facilitate water uptake or environmental interaction at sub zero temperatures. The insect class refers to certain freeze tolerant insects which naturally produce INPs in order to control where and when ice forms on in their bodies to prevent lethal intracellular freezing.
INPs can also be classified according to activity type, or functional classes, categorized A, B and C according to the temperature at which ice nucleation is activated, this is directly tied to the size o the protein aggregate. Class A corresponds to the highly efficient INPs which nucleate ice between -2°C and -5°C, such as inaK, this type of nucleation requires very large and ordered protein aggregates and is usually connected to the presence of specific membrane lipids. Class B refers to moderately efficient INPs catalyzing ice between -7°C and -9°C, they appear as intermediate sized protein clusters. Class C identifies the less efficient INPs nucleating ice below -10°C, the activity is linked to smaller protein clusters or individual INP monomers.
Finally, INPs can be classified according to the protein family and various species of bacteria contain different genetic orthologs of the ice nucleation genes. There is the Pseudomonas family, including inaZ, inaK, inaV, inaQ, they often share similar genetic structure such as N-terminal anchors, repetitive fragments and C-terminals. There is the Erwinia and Pantoea families originating from Gram-negative bacteria found in plants, including inaA, inaU and inaE. Additionally, as aforementioned there is the non-bacterial families such as the fungal or the insect families.
Reference list
Biotreks (2021) ‘Ice nucleation proteins – a synthetic pathway to alleviate ice loss’, Biotreks, (e202111).
GoldBio (n.d.) The Extraordinary Bacterial Proteins That Make Snow. Available at: https://www.goldbio.com/blogs/articles/the-extraordinary-bacterial-proteins-that-make-snow (Accessed:
25 April 2026).
Hudait, A. et al. (2024) ‘Hierarchical assembly and environmental enhancement of bacterial ice nucleators’, Proceedings of the National Academy of Sciences (PNAS), 121(18), p. e2409283121. doi: 10.1073/pnas.2409283121.
Lukas, M. et al. (2025) ‘A New Class of Fungal Ice-Nucleating Proteins with Bacterial Ancestry’, ChemRxiv. doi: 10.26434/chemrxiv-2025-73058.
Pandey, R. et al. (2016) ‘Ice-nucleating bacteria control the order and dynamics of interfacial water’, Science Advances, 2(4), p. e1501630. doi: 10.1126/sciadv.1501630.
Schmid, D. (2026) Glacier Blankets Could Help Prevent Melting. [Online Video]. 25 April. Available at: https://www.youtube.com/watch?v=hKT_SGK2qtY (Accessed: 25 April 2026).
USNSJ (n.d.) ‘Wonders of the Invisible World: Pseudomonas syringae, the Ice Maker’, University of Southern North Science Journal, 2(2).
- Why one INP is used over another
Do not just list INPs. Compare them critically. Ask:
- Why is one INP preferred over another?
- Is the choice based on nucleation temperature?
- ease of expression?
- stability?
- safety?
- membrane dependence?
- prior characterization in literature?
This section should justify your own protein choice.
Choosing an INP for a project should depend on functional performance (its nucleation temperature capacity), the ease of expression and genetic stability, the membrane and lipid dependence, the stability of the INP and its environmental robustness, and, its safety and regulatory approval.
I am choosing to work with inaK as it offers more genetic stability, efficiency, reliability and it already has records of it being used in synthetic biology. Inak is frequently used as it has a superior compatibility with surface display, it is easier to anchor secondary protein to a cell surface using inaK over other INPs, the N-terminal of the inaK is highly optimized for integration into Gram-negative bacterial membranes and will be less likely to produce misfolds when anchoring. Additionally, inaK offers better fusion stability as it preserves its own folding and ice nucleating properties even if fused with larger fluorescent reporters such as sfGFP. Furthermore, inaK is a Class A INP with the ability to freeze at higher subzero temperatures it also offers better predictability and consistency as it has been more studied. According to Jung H.C. in his study ‘Expression of Candida antarctica lipase B on the surface of Escherichia coli using InaK anchoring motif’ , Enzyme and Microbial Technology, (1998), inaK’s N-terminal domain is a superior anchor for displaying functional enzymes on the surface of bacteria. Shi H. also demonstrates in his study ‘A novel surface display system using the ice nucleation protein InaK-N as an anchor for the directed evolution of a highly active organophosphorus hydrolase’ , Applied and Environmental Microbiology, (2015), that inaK’s anchor is very stable even in harsh environmental conditions, reinforcing the idea of using them for glacier blanket technologies. Moreover, Li Q. also highlights in his research ‘Surface display of Vitreoscilla hemoglobin on Escherichia coli using InaK-N and its effects on cell growth’, Letters in Applied Microbiology, (2009), that inaK can be combined with complex proteins while maintaining its ice nucleation functionality and the physiological health of the host cell. The codon optimized inaK sequence has already been well characterized in E. coli systems making it easy to use in synthetic biology. Furthermore, the study by Roeters in 2024 in The Journal of Physical Chemistry C shows inaK has a sensitivity to enhancers when maximum efficiency is needed, inaK can have the ability to undergo a 100-fold enhancement in the presence of compounds such as polyols as they can improve stability of the hydration order in the highly repetitive segments of inaK.
Overall, inaK is the second INP to freeze at higher subzero temperatures after inaZ, both inaK and inaZ have been well characterized but inaK is more compatible with synthetic biology and it could easily be fused with an sfGFP, the inaK has extensively been studied. Inak is potentially the best choice amongst INPs as it responds very well to being modified and can be easily tailored according to what the end use is.
In terms of working with safety regulations, working with Pseudomonas syringae would involve more difficulties as it is a classified pathogenic bacteria, however, Snowmax makes use of it in its inactive form which its safety has been EPA regulated. Additionally, the DNA sequence has already been recorded on UniProt meaning I can directly synthesize it using Benchling and Twist avoiding safety issues as the inaK protein is not pathogenic.
Reference list
Biotreks (2021) ‘Ice nucleation proteins – a synthetic pathway to alleviate ice loss’, Biotreks, (e202111).
GoldBio (n.d.) The Extraordinary Bacterial Proteins That Make Snow. Available at: https://www.goldbio.com/blogs/articles/the-extraordinary-bacterial-proteins-that-make-snow (Accessed: 25 April 2026).
Jung, H.C. et al. (1998) ‘Expression of Candida antarctica lipase B on the surface of Escherichia coli using InaK anchoring motif’, Enzyme and Microbial Technology, 22(5), pp. 348–354.
Hudait, A. et al. (2024) ‘Hierarchical assembly and environmental enhancement of bacterial ice nucleators’, Proceedings of the National Academy of Sciences (PNAS), 121(18), p. e2409283121. doi: 10.1073/pnas.2409283121.
Li, Q. et al. (2009) ‘Surface display of Vitreoscilla hemoglobin on Escherichia coli using InaK-N and its effects on cell growth’, Letters in Applied Microbiology, 49(1), pp. 71–76.
Lukas, M. et al. (2025) ‘A New Class of Fungal Ice-Nucleating Proteins with Bacterial Ancestry’, ChemRxiv. doi: 10.26434/chemrxiv-2025-73058.
Roeters, S. J. et al. (2024) ‘Polyol-Induced 100-Fold Enhancement of Bacterial Ice Nucleation Efficiency’, The Journal of Physical Chemistry C, 128(15). doi: 10.1021/acs.jpcc.4c07422.
Schmid, D. (2026) Glacier Blankets Could Help Prevent Melting. [Online Video]. 25 April. Available at: https://www.youtube.com/watch?v=hKT_SGK2qtY (Accessed: 25 April 2026).
Shi, H. et al. (2015) ‘A novel surface display system using the ice nucleation protein InaK-N as an anchor for the directed evolution of a highly active organophosphorus hydrolase’, Applied and Environmental Microbiology, 81(15), pp. 5128–5135.
USNSJ (n.d.) ‘Wonders of the Invisible World: Pseudomonas syringae, the Ice Maker’, University of Southern North Science Journal, 2(2).
- Motifs necessary for ice nucleation
Describe the sequence motifs associated with INP function and explain which ones are believed to be essential for activity.
INPs share general features, here I will focus on the bacterial INPs. Bacterial INPs have a three domain structure and repetitive motifs. All bacterial INPs have a 𝞫-helix fold shaped by tandem repeats. INPs also all have exact spacing of Threonine and Serine residues matching ice lattice, this geometric spacing quality is a requirement for the protein to function as a template. The domain structure is always comprises a non repetitive N-terminal domain, (hydrophobic region which can anchor the protein to the outer membrane), a non repetitive C-terminal domain (hydrophilic tail assisting in protein stability and folding) and a central repetitive domain CRD ( the engine of the protein formed of tandem repeats which organize water molecules). The CRD is structured in hierarchy repeats composed of 16 amino acids which are then grouped in larger 48 residue periodicities. The entirety of the CRD is essential for ice nucleation at higher temperatures. Additionally, nucleation activity cannot occur without the 𝞫-helix fold meaning any mutations brought to the CRD has to be done in a way that it does not affect the 𝞫-helix fold outcome, this structurally essential. Moreover, the Threonine rich motif is essential to creating a water binding surface, if Threonine is replaced with non-polar residues then the protein might still fold correctly but risks losing nucleation capabilities, this is essential for functionality. Finally, the amount of repetition is essential as for instance a single 16 residue motif cannot nucleate ice, a certain mass of residue motif is required, explaining why inaK for instance has such a long repeat and demonstrates how it correlates to it being a very effective INP, this is essential for efficiency.
Reference list
Hudait, A. et al. (2024) ‘Hierarchical assembly and environmental enhancement of bacterial ice nucleators’, Proceedings of the National Academy of Sciences (PNAS), 121(18), p. e2409283121. doi: 10.1073/pnas.2409283121.
Pandey, R. et al. (2016) ‘Ice-nucleating bacteria control the order and dynamics of interfacial water’, Science Advances, 2(4), p. e1501630. doi: 10.1126/sciadv.1501630.
Roeters, S. J. et al. (2024) ‘Polyol-Induced 100-Fold Enhancement of Bacterial Ice Nucleation Efficiency’, The Journal of Physical Chemistry C, 128(15). doi: 10.1021/acs.jpcc.4c07422.
USNSJ (n.d.) ‘Wonders of the Invisible World: Pseudomonas syringae, the Ice Maker’, University of Southern North Science Journal, 2(2).
- Effect of motif repetition
Examine how repeat number affects nucleation activity. This is a key question. Do not assume “more repeats is always better.” Instead ask:
- Is there a minimum repeat number for activity?
- Is there an optimal number?
- Does activity plateau after a certain length?
- Does repeat number affect aggregation, folding, or membrane presentation?
This could become a central research question.
Indeed, the idea that more repeats involve better or more nucleation is not so linear, the number of repeats defines which class the INP belongs to and how much nucleation it can produce.
As aforementioned a 16 residue motif cannot produce any ice nucleation, there is a threshold of 15 to 20 repeats, which is the length of a typical CRD, to achieve the most basic level of ice nucleation at temperatures lower than -10°C. Below this number of repeats it is likely the protein could fold into a 𝞫-helix but it won’t catalyze ice as it will not have enough of a surface area to hold water molecules against temperature fluctuations. In order for the INP to function it needs an area large enough to stabilize ice catalyzation, this is referred to as the critical ice nucleus which is the smallest cluster of water molecules possible able to transform into a crystal rather than melt. The optimal number depends fully on the temperature the INP is supposed to be activated at or resist to. There is a correlation between the length of the sequence with which class the INP belongs to, the longer sequence belong to higher classes and vice versa. But, as much as longer repeats improve the capacity to nucleate at higher temperatures, too long of a sequence is likely to cause instability and become prone to genetic recombination and misfolding which can lead to failure to function. The activity does in fact reach a plateau as once the protein is long enough to stabilize a critical ice nucleus adding more repeats will not improve its function but rather might diminish its effectiveness, within each A B C class the INPs will plateau at their maximum capacity of repeats. Overcoming a plateau would depend more on aggregation capacity rather than length. The amount of repeats can impact aggregation, folding and membrane presentation. For class A the repeat number is essential to aggregation, longer repetitive sequences offer a better surface for proteins to stack on, without enough repeats proteins will not stack properly and won’t be very effective. The repeat number also impacts the folding process as the more the repeat number increases the larger the stress on the cell is for folding, the ribosome is challenged to reproduce with high fidelity a highly repetitive sequence. Finally, if the repetitive domain becomes too important it can become too heavy or too hydrophobic for the N-terminal anchor to be able to successfully connect the protein to the outer membrane, the protein might end up stuck in the cytoplasm where the INP function becomes useless.
Reference list
Biotreks (2021) ‘Ice nucleation proteins – a synthetic pathway to alleviate ice loss’, Biotreks, (e202111).
Hudait, A. et al. (2024) ‘Hierarchical assembly and environmental enhancement of bacterial ice nucleators’, Proceedings of the National Academy of Sciences (PNAS), 121(18), p. e2409283121. doi: 10.1073/pnas.2409283121.
Ling, M. L. et al. (2018) ‘The constructive role of protein repeats in ice nucleation’, Nature Communications, 9, p. 3314.
Pandey, R. et al. (2016) ‘Ice-nucleating bacteria control the order and dynamics of interfacial water’, Science Advances, 2(4), p. e1501630. doi: 10.1126/sciadv.1501630.
Roeters, S. J. et al. (2024) ‘Polyol-Induced 100-Fold Enhancement of Bacterial Ice Nucleation Efficiency’, The Journal of Physical Chemistry C, 128(15). doi: 10.1021/acs.jpcc.4c07422.
- Mechanism of ice nucleation by INPs
Go deeper into how INPs work:
- how they organize water
- how they template ice-like ordering
- how oligomerization or clustering affects activity
- why membrane localization may matter
The molecular template and hierarchical assembly of INPs, especially inaK, able to organize water molecules into a structured solid crystal using only one protein is an incredible biological engineering mechanism of nature. An INP is able to actively organize water and direct its position through its hydrophobic and hydrophilic balanced motif. The 𝞫-helix structure provides a face with periodic motifs where the hydrophobic parts like Glycine prevent water molecules from attaching too strongly where in contrast the hydrophilic parts like Threonine groups form precise hydrogen bonds with the water molecules. This dual aspect of the surface offers a stable hydration layer as water molecules are linked to a 2D sheet imitating the surface of an ice crystal. The template ice-like ordering comes from the geometric matching of INPs’ structures with lattice matching, where spacing and distance of residues is very precisely organized in the repeat sections. Having an extremely organized and repetitive template allows the protein to use less energy and increase its nucleating ability as it facilitates the organization of the water molecules. Oligomerization or clustering affects activity by making it stronger or weaker. An individual INP is too weak to actively form ice whereas the class A INPs which are constituted of clusters have demonstrated to be more powerful, there is a correlation between the size of the repetitive section and the amount of ice the protein is able to produce. Additionally, a wider surface area can better stabilize a larger critical ice nucleus, which itself will be more resistant to higher temperatures making the INP more effective and stronger, this hierarchy is shown by Hudait in ‘Hierarchical assembly and environmental enhancement of bacterial ice nucleators’, Proceedings of the National Academy of Sciences, 2024.
Membrane localization matters because for INPs to function correctly and effectively they must be situated in the cell membrane, therefore, even through a cell free design one would still have to synthesize a cell membrane. If the INP doesn’t reach the cell membrane (thanks to the N-terminal) then it is rendered useless, for ice catalyzation to be used it needs to occur on the outside of the cell.
Reference list
Biotreks (2021) ‘Ice nucleation proteins – a synthetic pathway to alleviate ice loss’, Biotreks, (e202111).
Hudait, A. et al. (2024) ‘Hierarchical assembly and environmental enhancement of bacterial ice nucleators’, Proceedings of the National Academy of Sciences (PNAS), 121(18), p. e2409283121. doi: 10.1073/pnas.2409283121.
Pandey, R. et al. (2016) ‘Ice-nucleating bacteria control the order and dynamics of interfacial water’, Science Advances, 2(4), p. e1501630. doi: 10.1126/sciadv.1501630.
Roeters, S. J. et al. (2024) ‘Polyol-Induced 100-Fold Enhancement of Bacterial Ice Nucleation Efficiency’, The Journal of Physical Chemistry C, 128(15), pp. 5845–5854. doi: 10.1021/acs.jpcc.4c07422.
USNSJ (n.d.) ‘Wonders of the Invisible World: Pseudomonas syringae, the Ice Maker’, University of Southern North Science Journal, 2(2).
- Existing methods for INP production
Review how INPs are currently produced:
- native microbial production
- recombinant in vivo bacterial expression
- cell-free synthesis
- membrane-based reconstitution or display systems
INP production has progressed from harvesting wild type bacteria to synthetically engineered ones.
The native microbial production involved cultivating naturally ice nucleating bacterias such as the Pseudomonas syringae. The bacterias would be cultivated in large scale fermenters, once the desired density was reached the cell often deactivated through UV or chemical treatment to prevent environment damage from their pathogenic nature. Companies like Snomax use pelletized inactive Pseudomonas syringae.
Recombinant in vivo bacterial expression is safer, non pathogenic, process of inserting the INP gene into a lab strain such as E.coli or B.subtilis, the protein will be expressed in the cytoplasm or led to the outer membrane.
Cell free protein synthesis is used to produce INPs from DNA templates and added to a cell free solution, this allows to bypass the need for living cells and only requires the mechanical components such as the ribosomes, enzymes and amino acids extracted from a cell. This method is faster and offers better control for longer, repetitive proteins.
Lastly, membrane based reconstitution or display systems are methods which combine recombinant expression with artificial membranes. For example, proteoliposomes is when INPs are retracted and reconstituted in a synthetic lipid vesicle, a liposome. This is the current method used to synthetically produce snow and allows researchers such as Hudait to study the different lipid types and understand the clustering required for high temperature activity.
Reference list
Biotreks (2021) ‘Ice nucleation proteins – a synthetic pathway to alleviate ice loss’, Biotreks, (e202111).
GoldBio (n.d.) The Extraordinary Bacterial Proteins That Make Snow. [Online]. Available at: https://www.goldbio.com/blogs/articles/the-extraordinary-bacterial-proteins-that-make-snow (Accessed: 26 April 2026).
Hudait, A. et al. (2024) ‘Hierarchical assembly and environmental enhancement of bacterial ice nucleators’, Proceedings of the National Academy of Sciences (PNAS), 121(18), p. e2409283121.
Jung, H.C. et al. (1998) ‘Expression of Candida antarctica lipase B on the surface of Escherichia coli using InaK anchoring motif’, Enzyme and Microbial Technology, 22(5), pp. 348–354.
Li, Q. et al. (2009) ‘Surface display of Vitreoscilla hemoglobin on Escherichia coli using InaK-N and its effects on cell growth’, Letters in Applied Microbiology, 49(1), pp. 71–76.
Roeters, S. J. et al. (2024) ‘Polyol-Induced 100-Fold Enhancement of Bacterial Ice Nucleation Efficiency’, The Journal of Physical Chemistry C, 128(15).
- Existing products that use INPs
Survey current commercial or applied uses of INPs, such as:
- artificial snow production
- freeze structuring
- possibly food or biotech applications
This helps define the translation potential of your work.
INPs are now used in a wide range of applications from environmental and ecological focused uses and research to commercial sectors. These proteins are utilized in many industries as they can accurately control the ice nucleation phase.
As aforementioned, they are often used in artificial snow production, as demonstrated by Snomax International using native bacterial proteins for snow making at temperatures where traditional machines would not be effective. The use of INPs in snow production significantly reduces the energy and water cost for ski resorts and therefore reduces the commercial impact on the environment. This application is close to my area of research as it is used within a similar context of mountains and rebuilding and reinforcing skiing tracks against rising temperatures. However, the mindset and end use varies widely, the aim to protect and preserve glaciers does not have a commercial use as it focuses on an ecological solution to climate change rather than compensating for climate change consequences human activity does not want to face. Additionally, the technology would still vary as the aim for me is not to produce snow but ice which will have different temperature requirements and a different upkeep.
My research of INPs would come closer to the geo-engineering experimental approach of considering the use of INPs for cloud seeding, where silver iodide is commonly used and can have some toxic secondary effects but INPs are biodegradable and highly efficient making them a possible sustainable alternative. These approaches are still conceptual and theoretical projects as seen in the following projects; for atmospherical cloud seeding in the Walser 2024 project ‘Fungal ice nucleation proteins open new pathways for weather modification and biopreservation’, Science Advances, or, in idea of developing glacier blankets in the case study of Biotreks 2021, ‘Ice nucleation proteins – a synthetic pathway to alleviate ice loss’ .
INPs are also commonly used in the food industry as freezing structuring and preservation technologies allowing to control the size and distribution of ice crystals. Large ice crystals can damage the texture of frozen food whereas precise ice nucleation can preserve it, the INPs are used as freeze structuring agents to form many small ice crystals simultaneously. Similarly to the artificial snow making technologies using INPs to freeze food also allows to reduce energy waste as the process is controlled and optimized.
Furthermore, INPs are used in the biotechnology or medical field for biopreservation. INPs have the ability to prevent supercooling and are used to better preserve sensitive biological samples. For example it can be used for cryopreservation where samples are frozen in a homogeneous and controlled way preventing cells from being damaged or dying. There are case studies where this technology is being experimented with in organ preservation, with controlled nucleation in liver preservation as discussed in ‘Controlled ice nucleation by ice-nucleating proteins for the cryopreservation of complex biological systems’, Biomaterials (38, pp. 11–21. doi: 10.1016/j. ), 2015, by Lee, C.Y., et al..
Reference list
Biotreks (2021) ‘Ice nucleation proteins – a synthetic pathway to alleviate ice loss’, Biotreks, (e202111).
GoldBio (n.d.) The Extraordinary Bacterial Proteins That Make Snow. [Online]. Available at: https://www.goldbio.com/blogs/articles/the-extraordinary-bacterial-proteins-that-make-snow (Accessed: 27 April 2026).
Hudait, A. et al. (2024) ‘Hierarchical assembly and environmental enhancement of bacterial ice nucleators’, Proceedings of the National Academy of Sciences (PNAS), 121(18), p. e2409283121.
Lee, C.Y., et al. (2015) ‘Controlled ice nucleation by ice-nucleating proteins for the cryopreservation of complex biological systems’, Biomaterials, 38, pp. 11–21. doi: 10.1016/j.biomaterials.2014.10.050.
Roeters, S. J. et al. (2024) ‘Polyol-Induced 100-Fold Enhancement of Bacterial Ice Nucleation Efficiency’, The Journal of Physical Chemistry C, 128(15).
Schmid, D. (2026) Glacier Blankets Could Help Prevent Melting. [Online Video]. 25 April. Available at: https://www.youtube.com/watch?v=hKT_SGK2qtY (Accessed: 27 April 2026).
Snomax International (2026) The Science of Snomax: Maximizing Snow Production Efficiency. [Online Technical Bulletin].
USNSJ (n.d.) ‘Wonders of the Invisible World: Pseudomonas syringae, the Ice Maker’, University of Southern North Science Journal, 2(2).
Walser, A. et al. (2024) ‘Fungal ice nucleation proteins open new pathways for weather modification and biopreservation’, Science Advances, 10(12). doi: 10.1126/sciadv.adl1234.
- Compare INP production platforms
Include a dedicated comparison of:
Cell-free synthesis (CFS/CFPS) vs in vivo bacterial expression
Compare them on:
- production cost
- yield
- scalability
- speed
- ease of optimization
- ability to handle toxic or membrane-associated proteins
- purity
- regulatory or biosafety concerns
- safety of the final product
- need for post-expression assembly or membrane mimicry
This comparison should not be generic. It should be tied specifically to INPs.
For this section I made use of Gemini assistant by inputting my information collected with the comparative elements needed so I could create a clear comparative table.
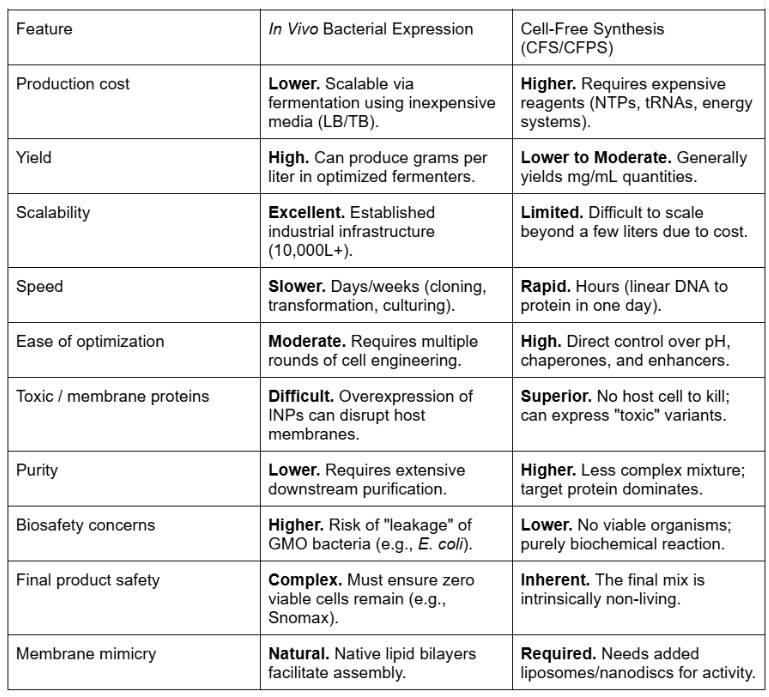

There are clearly very strong advantages to using CFPS compared to an in vivo method.
My main challenge when working with inaK, a class A INP, will be the membrane dependence issue, as class A INPs require a membrane in order to form larger ice clusters using the lipid bilayer. Class C INPs would be able to function without a membrane but would be significantly less useful for the needed output working on glacier preservation faced with the rise of temperatures as they will produce much less ice and much lower sub zero temperature. An in vivo method using bacteria naturally provides a base for the protein to attach to, however, large amounts of inaK can become toxic to the host and might just become ineffective. Moreover, CFPS offers better optimization and enhancement enabling a superior platform of the 100 fold provided by polyols. In a cell free design the polyol concentration can be precisely adjusted to avoid killing the host which can enable better maximization to template inaK. The other main issue I would be faced with using CFPS is the limited scalability as it can be more costly, using an in vivo method the INP like inaK could be inoculated into a glacier in a new host and naturally form horizontal gene transfer, which would not be possible with CFPS. Finally, CFPS has much less regulatory and biosafety issues which will be extremely relevant for my project as my technology would be directly inoculated into nature and the natural consequence this could have must be very carefully considered.
Reference list
Biotreks (2021) ‘Ice nucleation proteins – a synthetic pathway to alleviate ice loss’, Biotreks, (e202111).
Hudait, A. et al. (2024) ‘Hierarchical assembly and environmental enhancement of bacterial ice nucleators’, Proceedings of the National Academy of Sciences (PNAS), 121(18), p. e2409283121.
Li, Q. et al. (2009) ‘Surface display of Vitreoscilla hemoglobin on Escherichia coli using InaK-N and its effects on cell growth’, Letters in Applied Microbiology, 49(1), pp. 71–76.
Roeters, S. J. et al. (2024) ‘Polyol-Induced 100-Fold Enhancement of Bacterial Ice Nucleation Efficiency’, The Journal of Physical Chemistry C, 128(15).
Silverman, A.D. et al. (2020) ‘Cell-free gene expression: an expanded repertoire of applications’, Nature Reviews Genetics, 21(3), pp. 151-170. (General CFPS vs In Vivo comparison).
- Do we need the whole protein?
This is an excellent question and should be treated as a serious design hypothesis:
- Why express the full-length protein?
- Can the nucleation motifs alone work?
- Can a truncated construct reduce complexity and cost?
- What is lost if the full scaffold is removed?
- Are motifs alone sufficient for activity, or do they need correct spacing, repetition, and supramolecular assembly?
This is one of the strongest parts of your concept.
Interfering with the natural structure of an INP, here inaK, can completely shift its ability to correctly nucleate ice.
Expressing the full protein allows for the hierarchical architecture which focuses on geometry. The N-terminal and C-terminal cannot be taken out of the structure and the first plays the essential role of an anchor and the second is essential in the folding process, with these the central repetitive domain would likely fail to nucleate ice, its primary function. Additionally, the large size of the inaK gives it stability and ensures the protein can withstand the mechanical stress it undergoes in a synthetic cell surface or within a Glacier Blanket concept for instance.
As aforementioned a single nucleation motif cannot function alone, a single motif would be too small to overcome the thermal energy of liquid water, the long continuous and repetitive surface of the protein is essential to stabilize the critical ice nucleus. Additionally the 𝜷-helix is a critical part of the structure as this is what fold the individual strings of amino acid into the 𝜷-helix shape again essential to stability and the nucleation of ice.
A truncate construct would on one hand reduce the genetic burden during PRC or translation as the sequence would be shorter and could offer higher reliability in data recovery. However, on the other hand, as aforementioned this method would simply reduce the ice nucleation capability and efficiency and essentially would downgrade the protein, likely reducing an inaK protein from a class A to a class C.
Again, removing the scaffold would render the INP useless as without the N-terminal anchor it will not be able to orient itself to the membrane and might be able to template a few water molecules but won’t be able to form crystal lattice as it will not have a flat 2D surface and the motif will randomly float within the cell. Additionally, the N-terminal anchor is also what enables clustering, without it the protein will not efficiently nucleate ice.
Thus, the motifs provide the chemical code of the hydrogen bonding patterns, the spacing and repetition provides the physical geometrical template allowing the ice to form, and, the supramolecular assembly provides the scale neede for a class A INP like inaK to nucleate ice at a higher temperature.
Reference list
Biotreks (2021) ‘Ice nucleation proteins – a synthetic pathway to alleviate ice loss’, Biotreks, (e202111).
Hudait, A. et al. (2024) ‘Hierarchical assembly and environmental enhancement of bacterial ice nucleators’, Proceedings of the National Academy of Sciences (PNAS), 121(18), p. e2409283121.
Ling, M. L. et al. (2018) ‘The constructive role of protein repeats in ice nucleation’, Nature Communications, 9, p. 3314.
Pandey, R. et al. (2016) ‘Ice-nucleating bacteria control the order and dynamics of interfacial water’, Science Advances, 2(4), p. e1501630.
Roeters, S. J. et al. (2024) ‘Polyol-Induced 100-Fold Enhancement of Bacterial Ice Nucleation Efficiency’, The Journal of Physical Chemistry C, 128(15).
- Why is a membrane or membrane-like material needed?
Do not treat membrane dependence as a side detail. It may be central. Ask:
- Is the membrane required for folding?
- for clustering?
- for orientation?
- for multivalent display?
- for stabilization of the active nucleation surface?
The membrane is not simply a container for the protein, in this case the membrane becomes a functional factor or tool. Ice nucleation produced by an INP like inaK forms on the surface of the cell at the membrane for better more effective ice catalyzation. Without a membrane imitating scaffold like a liposome or nanodisc the INP, inaK, would fail to reach its class A efficiency. The membrane is essential for clustering, a lipid bilayer would act as a fluid 2D scaffold and the inaK would be restricted to a 2D plane which increases concentration. Orientation is not possible without a membrane, the protein would simply tumble in the space and not align to each other preventing ice lattice, the N-terminal ensures proper orientation with the membrane. On a mechanical aspect the membrane offers a resilient yet flexible scaffold preventing the template from collapsing. A membrane facilitates multivalent display as it becomes a hub for ice binding sites to form simultaneously. The membrane also plays a critical role in the protein folding as the lipid tales help the N-terminal domain to work properly in a hydrophobic environment. In CFPS system the lipid membrane is vital to avoid dead aggregates, inclusion bodies, and give the protein a direction upon translation.
Overall, a membrane of membrane like material is essential for ice nucleation and makes it more stable and efficient. For my project I will need to create a synthetic cell membrane.
Reference list
Hudait, A. et al. (2024) ‘Hierarchical assembly and environmental enhancement of bacterial ice nucleators’, Proceedings of the National Academy of Sciences (PNAS), 121(18), p. e2409283121.
Pandey, R. et al. (2016) ‘Ice-nucleating bacteria control the order and dynamics of interfacial water’, Science Advances, 2(4), p. e1501630.
Roeters, S. J. et al. (2024) ‘Polyol-Induced 100-Fold Enhancement of Bacterial Ice Nucleation Efficiency’, The Journal of Physical Chemistry C, 128(15).
Schmid, D. (2026) Glacier Blankets Could Help Prevent Melting. [Online Video]. 25 April. Available at: https://www.youtube.com/watch?v=hKT_SGK2qtY (Accessed: 27 April 2026).
- Identify the literature gap
Only after the full review should you define the gap. Possible gaps may include:
- poor understanding of minimal active INP units
- unclear relationship between repeat number and activity
- overreliance on full-length membrane-associated proteins
- low-yield or expensive production systems
- lack of rationally engineered INPs with tunable freezing temperatures
- insufficient comparison between CFS and in vivo systems for INP manufacture
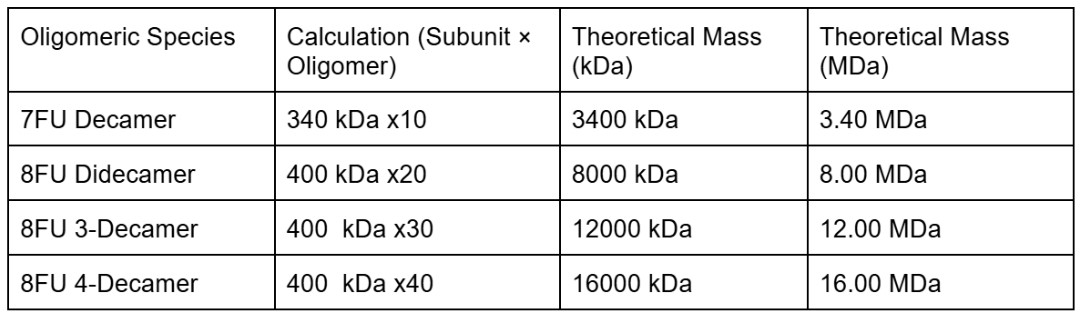
While we know how to identify INPs and how their general mechanism works there are still many existing gaps within literature. For instance, while there is an understanding of the different classes and that 8.33 MDa didecamers correlates with high temperature nucleation there is still uncertainty around the tipping point of minimal clusters, of how many proteins are actually needed. Additionally, there is a lack of high resolution structural data mapping the transition from a single 𝞫-helix monomer to a supramolecular assembly and without understanding what is the minimal viable cluster it makes it difficult to optimize synthetic cells the best efficiency with the least protein expression. On top of it, the relationship between repetitive sequence length and the freezing temperature is still to some extent speculative as INPs are usually tested on within similar context using the same amount of repeats and working within a membrane like environment, as explorations have been limited it is likely a lot of potential has not yet been explored. Moreover, the main evident gap in literature is the lack of knowledge on how to achieve class A function without relying on a membrane or membrane like environment. There is also limited information on the comparative data between in vivo and CFPS systems, currently it appears to be more of an overall idea. Finally, the potential of INPs has not been explored much further from what we know already, the idea of tailored design through the exploration of specific mutations has not been explored. In conclusion, in the context of my glacier preservation project using inaK I am faced with a few literature gaps regarding clustering requirements, precise manufacturing comparison and varied environmental application.
Reference list
Hudait, A. et al. (2024) ‘Hierarchical assembly and environmental enhancement of bacterial ice nucleators’, Proceedings of the National Academy of Sciences (PNAS), 121(18), p. e2409283121.
Ling, M. L. et al. (2018) ‘The constructive role of protein repeats in ice nucleation’, Nature Communications, 9, p. 3314.
Roeters, S. J. et al. (2024) ‘Polyol-Induced 100-Fold Enhancement of Bacterial Ice Nucleation Efficiency’, The Journal of Physical Chemistry C, 128(15).
Silverman, A.D. et al. (2020) ‘Cell-free gene expression: an expanded repertoire of applications’, Nature Reviews Genetics, 21(3), pp. 151-170.
- Define your innovation space
Since your focus is at the protein level, your innovation should be framed around (engineering the INP itself), not just producing it.
Your opportunity could be at several levels:
-Protein engineering
engineer the sequence to improve activity, stability, or manufacturability
-Chimeric design
combine active nucleation regions with scaffolds, membrane anchors, oligomerization domains, or display modules
-Minimal functional constructs
test whether shorter repeat-based designs can retain activity while reducing complexity
-Production optimization
design constructs that give higher yield and lower cost
-Tunable nucleation behavior
engineer proteins that nucleate ice at different temperatures depending on the application
Because your interest is at the protein level, I would advise you to focus the project around this core question:
What is the minimal structural and organizational requirement for efficient ice nucleation by INPs, and can we engineer a simpler, cheaper, or warmer-acting version than the natural protein?.
That question is much stronger than simply “produce an INP.”
I believe from this research that the best current option to explore relies around protein engineering and chimeric design. Structural modifications appear less likely to be successful from research.
My personal interrogation to redefine and push my aims:
Can I increase the efficiency of inaK by combining it with another compound?
sfGFP tag : increased stability and UV protection with both aspects increase ability to produce ice nucleation and be more resilient to higher temperatures (whether increasing the nucleation point or afterwards resisting warmer external temperatures)
Polyol : (Sorbitol, Glycerol, Xylitol) a hydroxyl rich molecule which can behave as a partner template, it cannot nucleate ice itself but can help organize the water molecules around the inaK repeats reducing potential entropy
Ions : adding specific ions to a CFPS design can stabilize the 𝞫-helix and inaK repeats improving nucleation
Ice Binding Domains (IBDs): added to the C-terminal help the INP adhere to the surface of another layer of ice (such as the glacier) preventing INPs to be naturally washed away by meltwater
Hydrophins : small surface active protein enable INP to spread evenly across a surface ( such as the ice layer of a glacier)
Melanin / Scytonemin : pigments which could serve as a sunscreen to the INP, protecting it from UVs would make it more resistant to melting
Lipid nanodiscs/liposomes: in CFPS combining inaK with a synthetic scaffold will increase the productivity of the N-terminal anchoring the INP with a stronger bond to the membrane
Protein cage: is a method where the INP is fused to self assembling protein cage (such as Ferritin or Encapsulin) combining inaK proteins into a larger single molecule creating a super-cluster by design rather than relying on the membrane activity
Reference list
Davies, P.L. (2014) ‘Ice-binding proteins: a remarkable capacity to adapt for life at cold temperatures’, Biochemical Journal, 458(1), pp. 9–20. doi: 10.1042/BJ20131291.
Garnham, C.P. et al. (2011) ‘A conserved water-organizing motif in ice-nucleating proteins’, Molecular Microbiology, 79(6), pp. 1419–1427. doi: 10.1111/j.1365-2958.2011.07546.x.
Govindarajan, A.G. and Lindow, S.E. (1988) ‘Size of bacterial ice-nucleation sites measured in situ by radiation inactivation’, Proceedings of the National Academy of Sciences (PNAS), 85(5), pp. 1334–1338.
Hudait, A. et al. (2024) ‘Hierarchical assembly and environmental enhancement of bacterial ice nucleators’, Proceedings of the National Academy of Sciences (PNAS), 121(18), p. E2409283121.
Lindow, S.E. et al. (1989) ‘Relationship between Ice Nucleation Frequency and inaZ Protein Content in Escherichia coli’, Molecular Plant-Microbe Interactions, 2(5), pp. 262–272.
O’Sullivan, D. et al. (2016) ‘The influence of pH, ionic strength and soluble organics on the ice nucleating ability of Pseudomonas syringae’, Atmospheric Chemistry and Physics, 16(11), pp. 7443–7454
Pandey, R. et al. (2016) ‘Ice-nucleating bacteria control the order and dynamics of interfacial water’, Science Advances, 2(4), p. e1501630. doi: 10.1126/sciadv.1501630.
Roeters, S. J. et al. (2024) ‘Polyol-Induced 100-Fold Enhancement of Bacterial Ice Nucleation Efficiency’, The Journal of Physical Chemistry C, 128(15).
Schmid, D. (2026) Glacier Blankets Could Help Prevent Melting. [Online Video]. 25 April. Available at: https://www.youtube.com/watch?v=hKT_SGK2qtY (Accessed: 27 April 2026).
Schoborg, J.A. et al. (2014) ‘Aqueous two-phase system (ATPS) for direct fractionation of proteins from cell-free protein synthesis’, Biotechnology and Bioengineering, 111(12), pp. 2405–2415.
Silverman, A.D. et al. (2020) ‘Cell-free gene expression: an expanded repertoire of applications’, Nature Reviews Genetics, 21(3), pp. 151–170. doi: 10.1038/s41576-019-0186-3.
Walser, A. et al. (2024) ‘Fungal ice nucleation proteins open new pathways for weather modification and biopreservation’, Science Advances, 10(12), p. eadl1234. doi: 10.1126/sciadv.adl1234.
![cover image]()
![cover image]()

Initial project proposal






Rafined project proposal


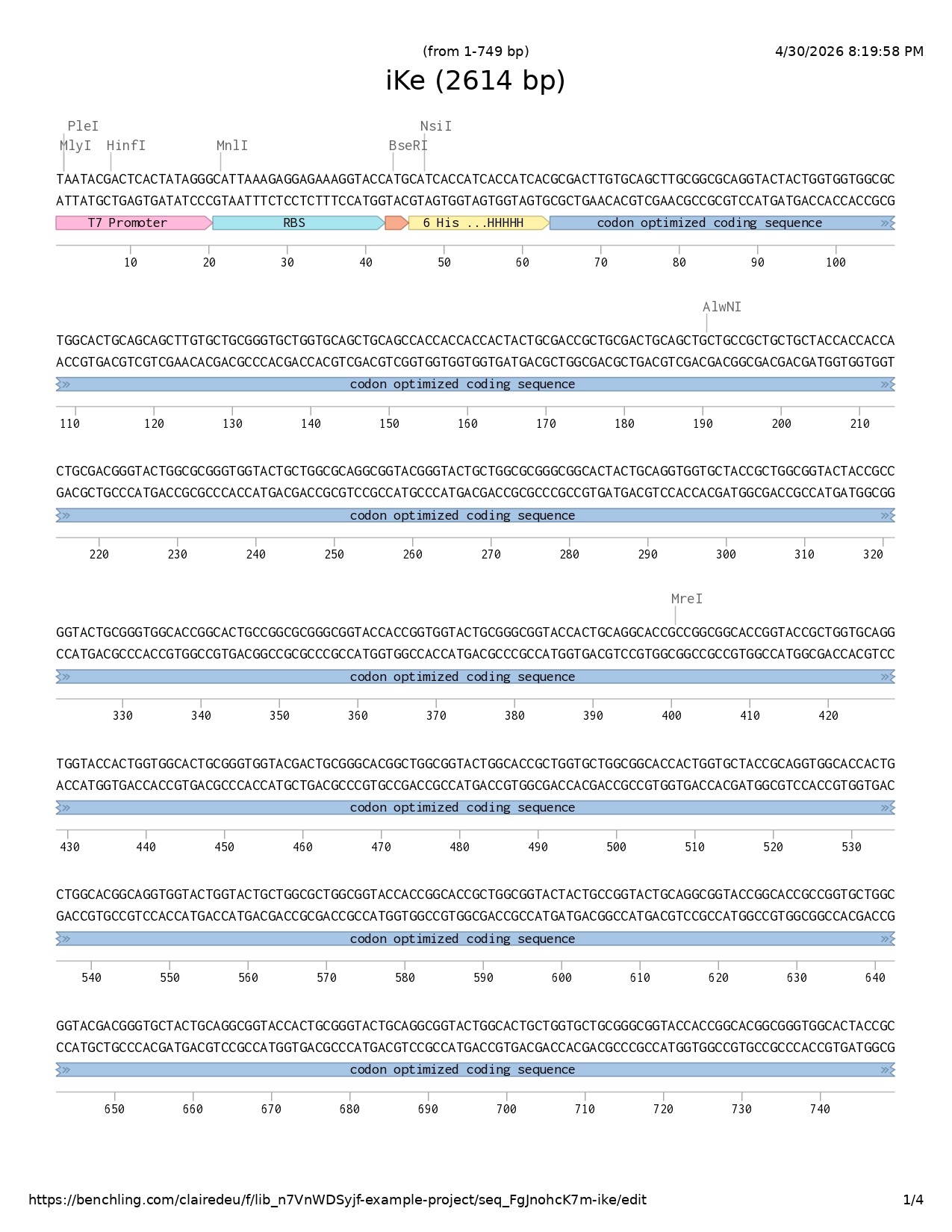
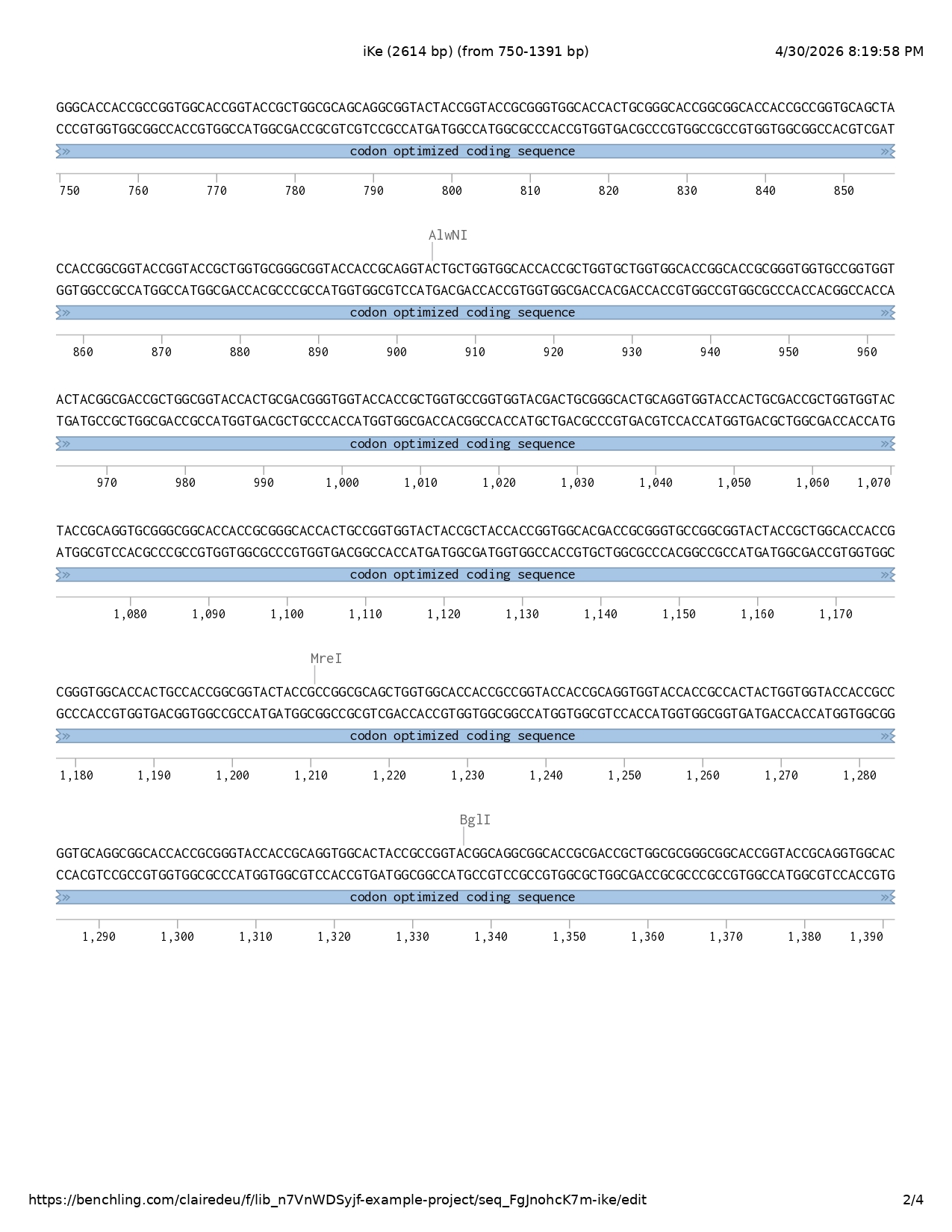
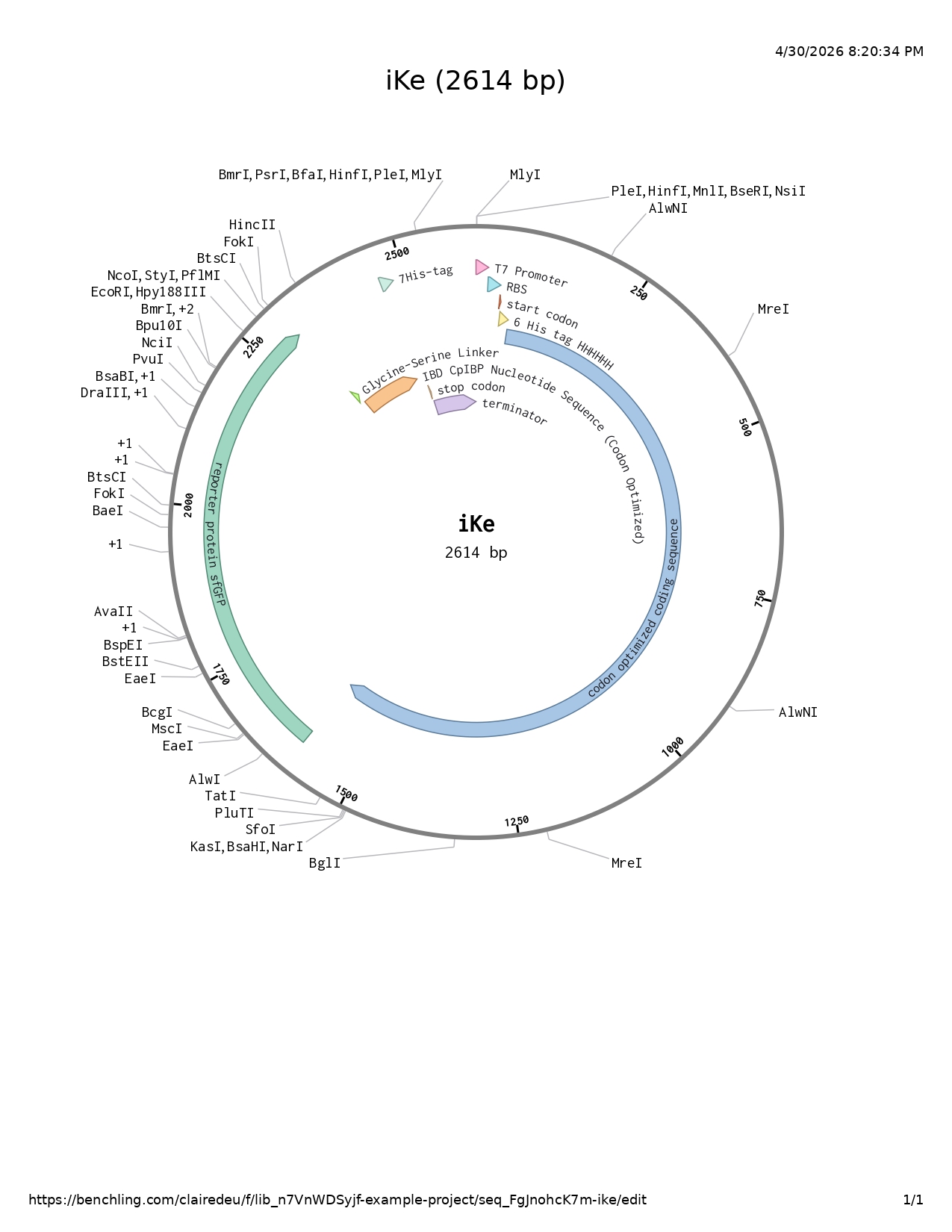
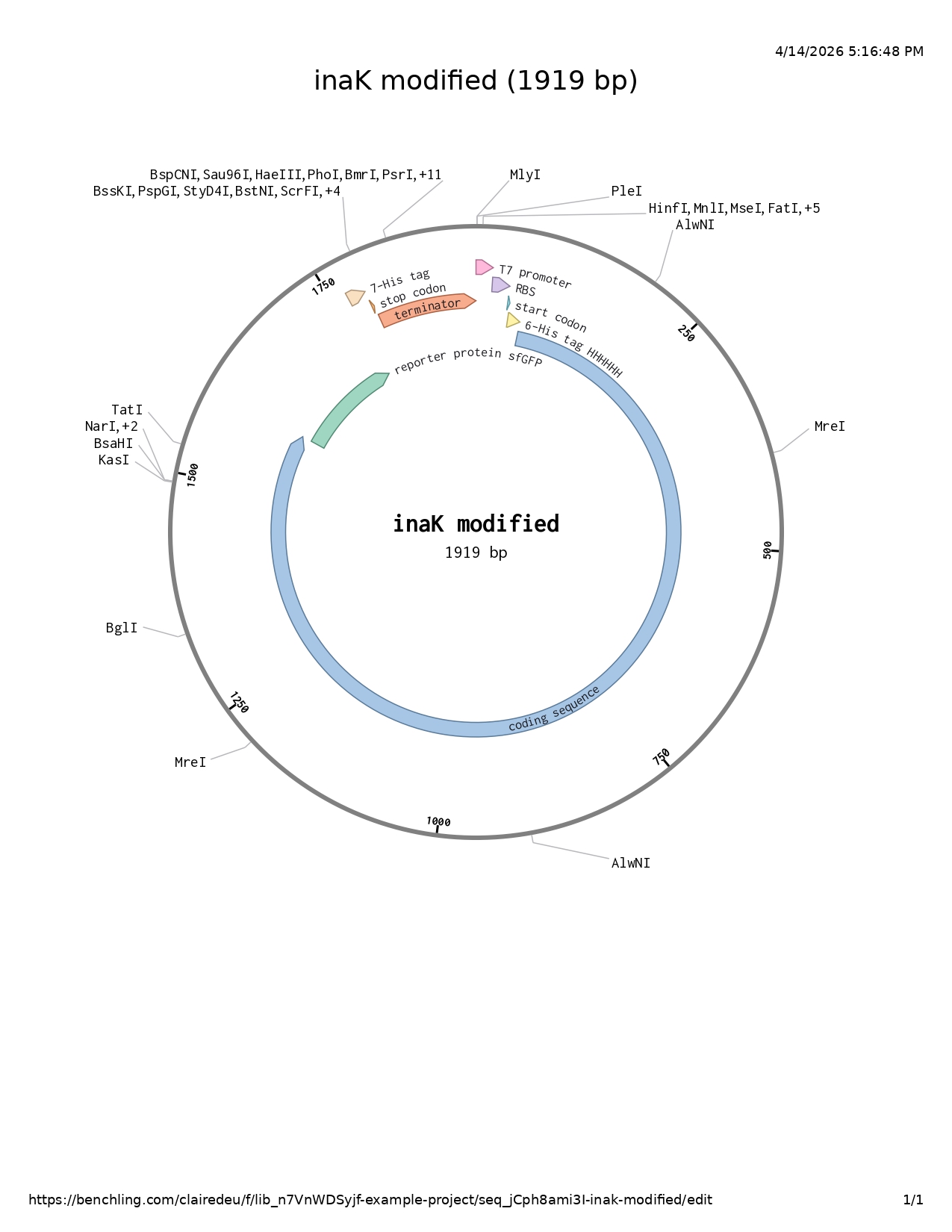
DNA sequencing
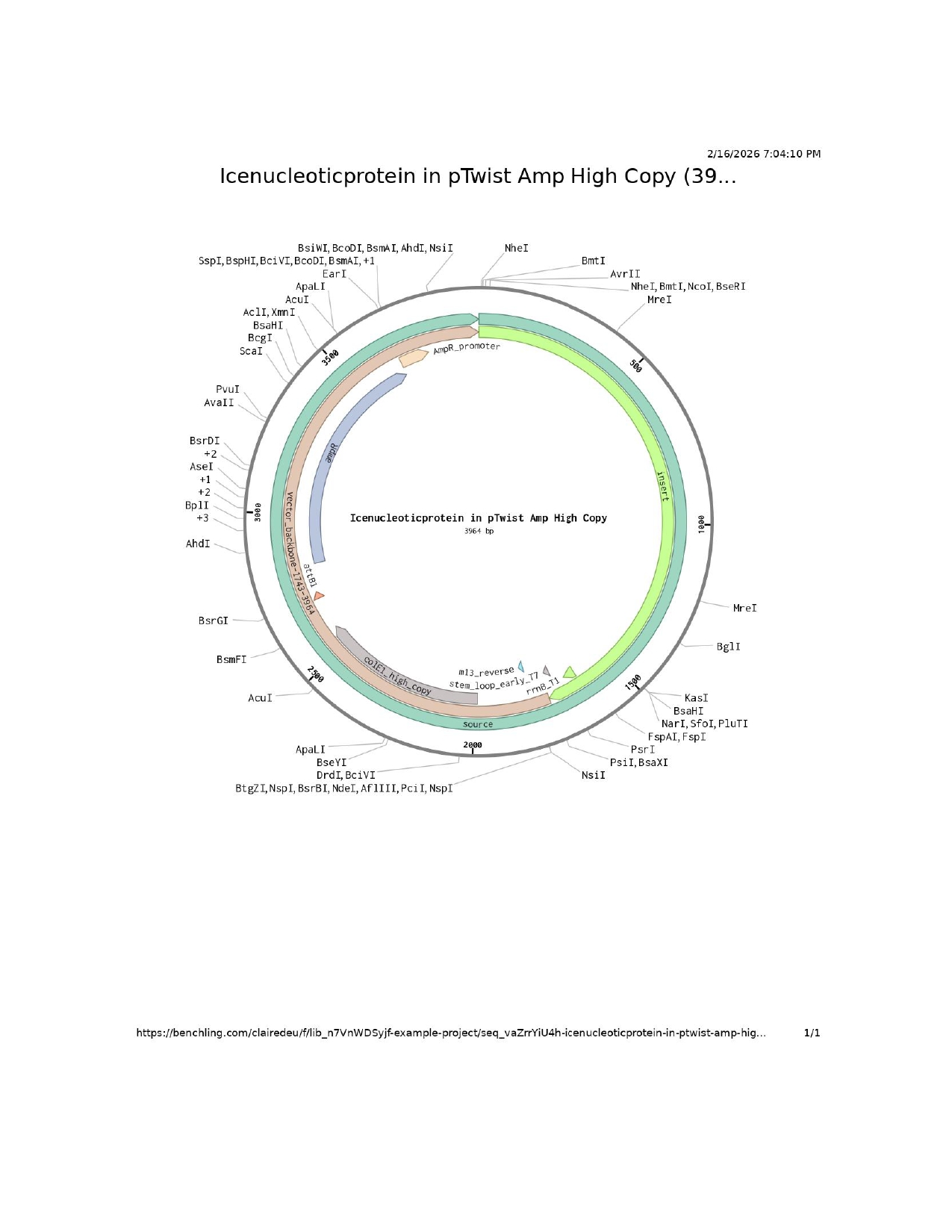
I followed the base DNA sequencing structure shown to us in week two and adapted it accordingly to my project.
- T7 promoter
- RBS
- Start codon
- 6His-tag
- Codon optimized coding sequence
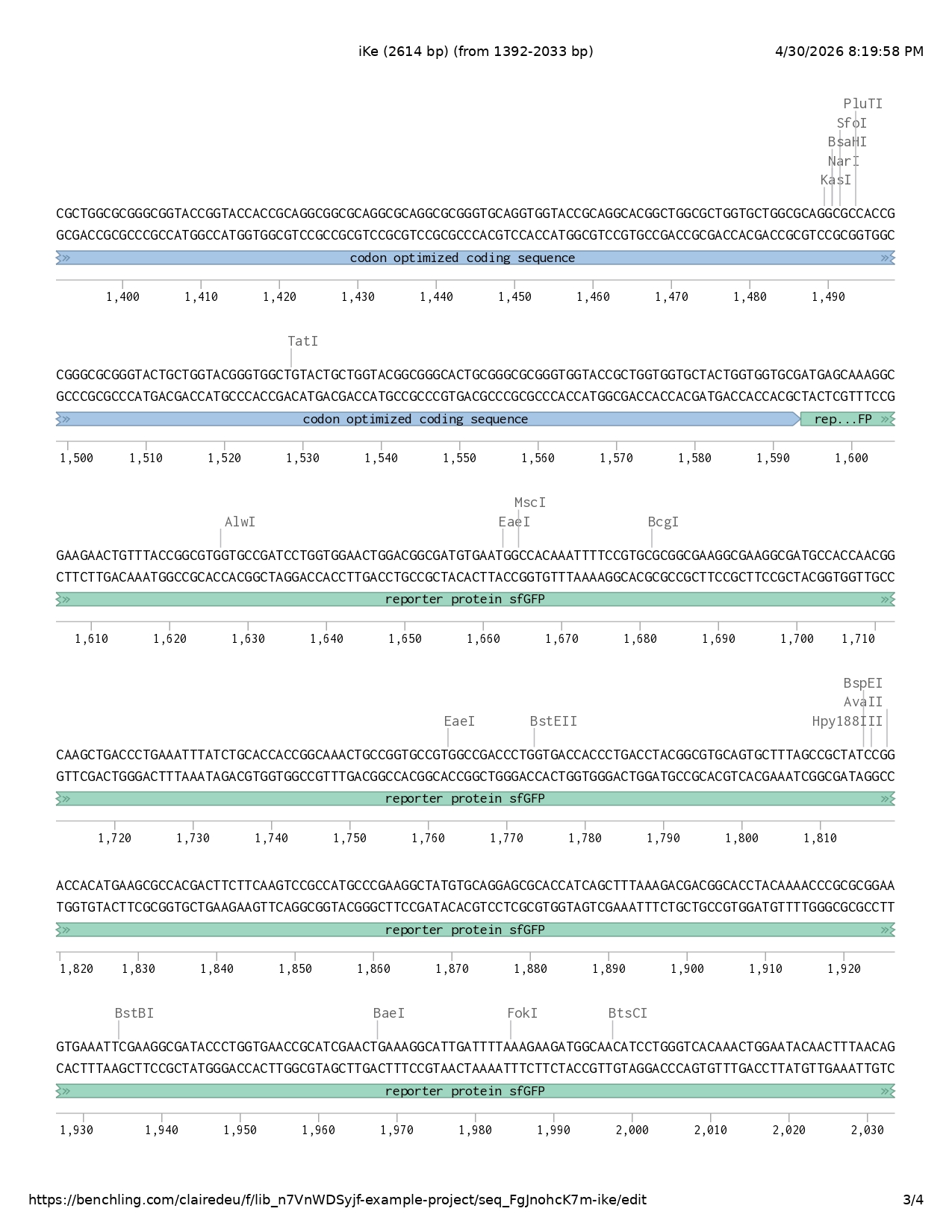
- Reporter protein sfGFP
- Glycine serine linker
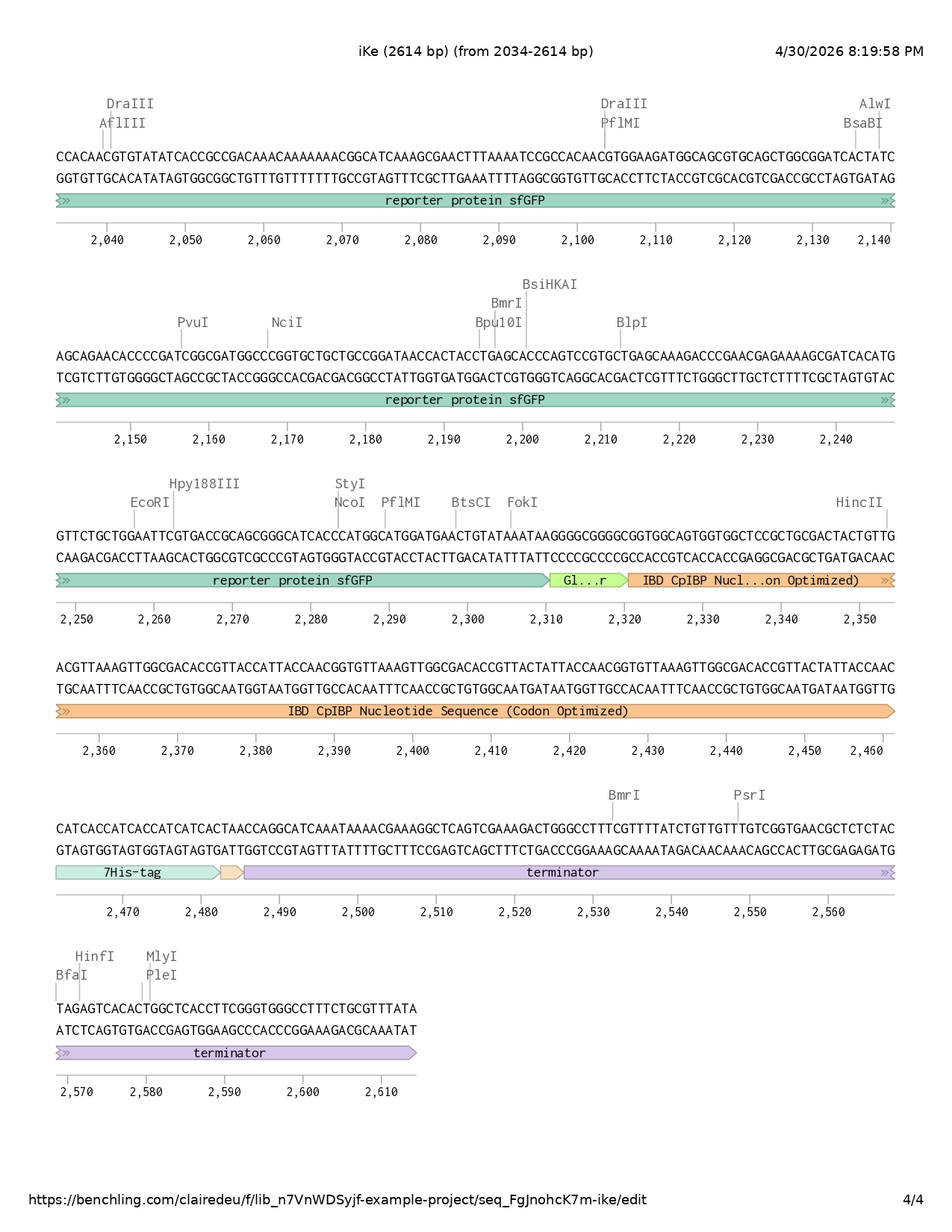
- IBD codon optimized
- 7His-tag
- Stop codon
- Terminator
Add Sorbitol polyol to inaK master mix which will increase inaK function

Attempt at Twist order
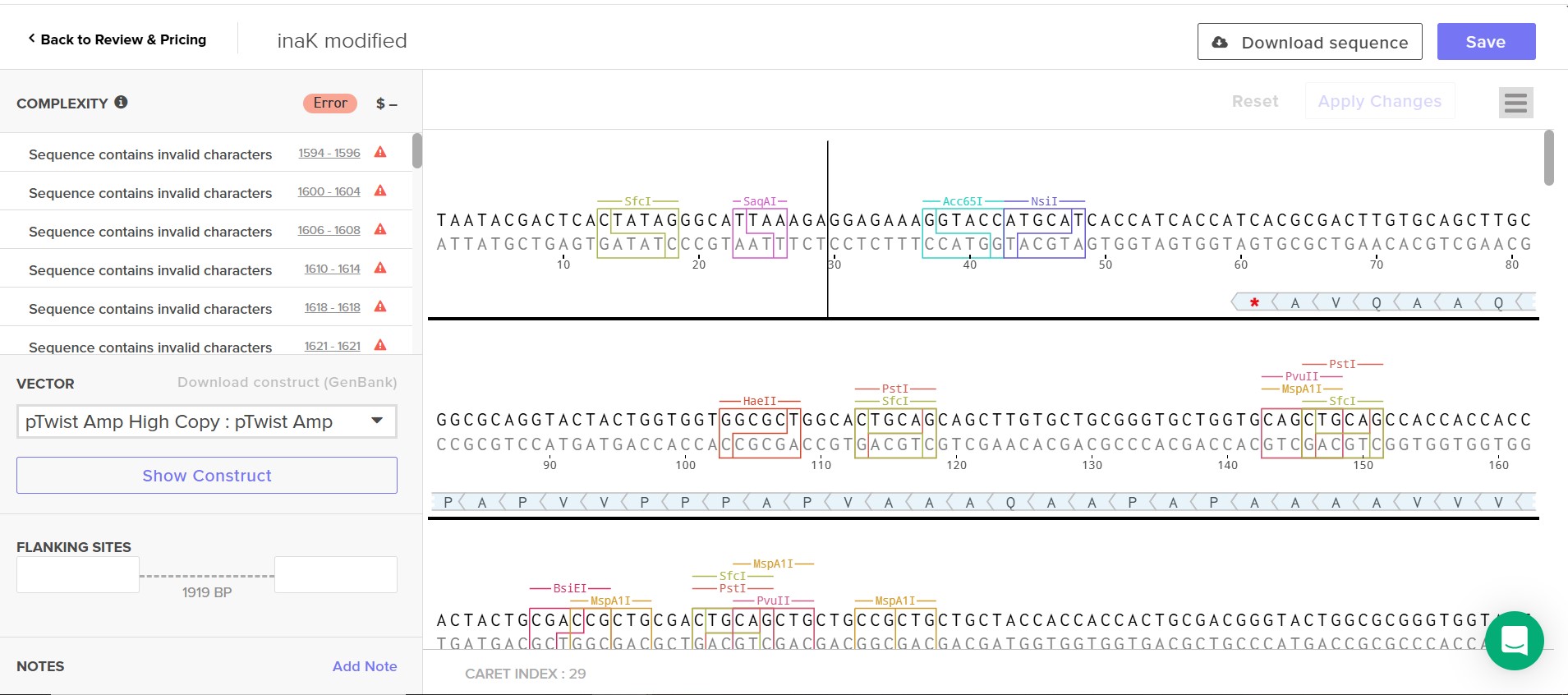

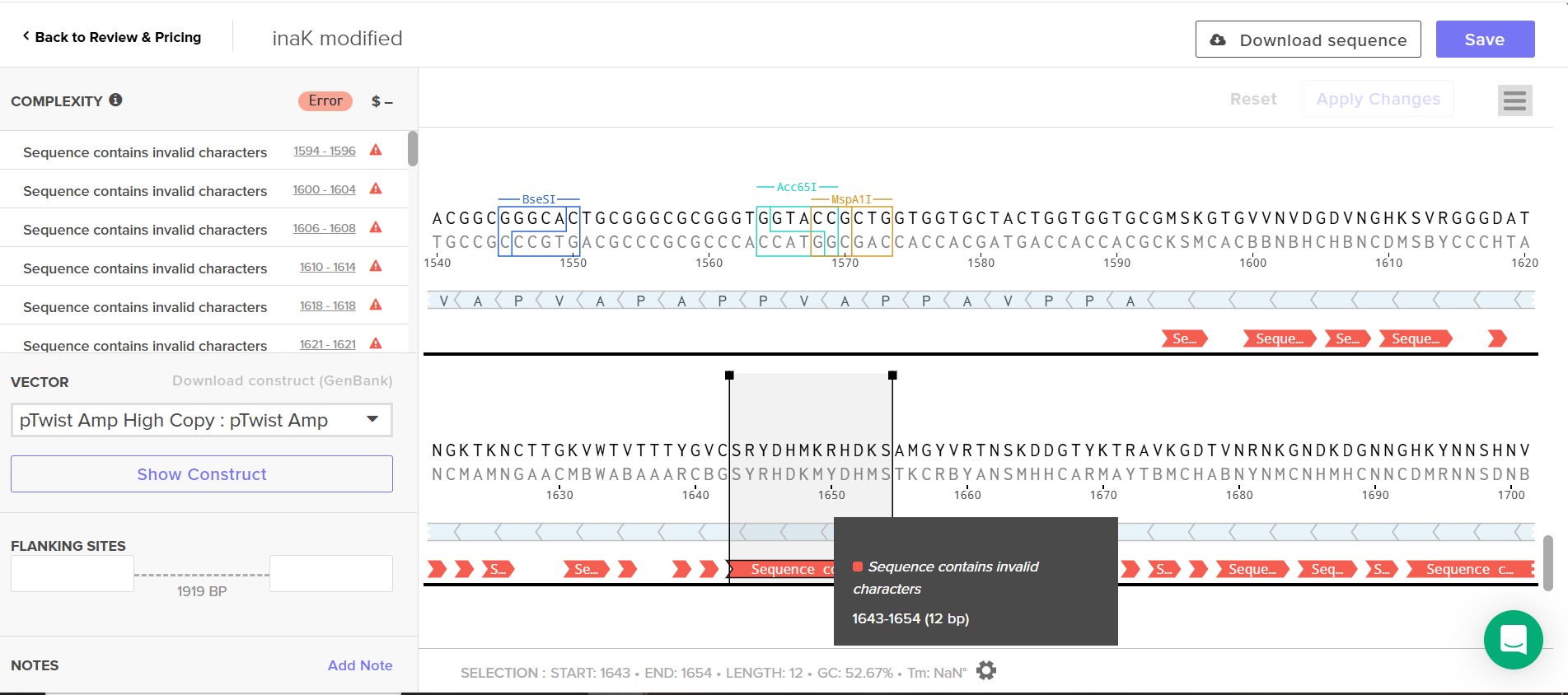
 The error found by Twist might be a reporter protein sfGFP issue as I insert the amino acid sequence rather than the nucleic acid sequence and Twist could not read it.
The error found by Twist might be a reporter protein sfGFP issue as I insert the amino acid sequence rather than the nucleic acid sequence and Twist could not read it.
Found limitation
With help of my TA Ahmad, it appears a project is already making use of the technology I was planning on innovating on, Snowmax International is already making use of ice nucleating proteins to improve ice catalyzation at higher temperatures in snow cannons to replenish skiing tracks.
Therefore, I will keep going with my second aim of modifying certain amino acids of my DNA sequence to improve the function of inaK and find an additional way of pushing further the innovation for this project.
Redefining my aims
Aim 1 - modify my inaK sequence to be stronger, more efficient and more stable by combining to the initial design structure an sfGFP tag (contains some UV protective benefits)and an ice binding domain, additionally, add to the master mix a polyol which will help the ice binding domain stick to the surface glacier better.
Aim 2 - increase the scalability and longevity of the project by testing the modified inaK sequence by adding extra Melanin or Scytonemin to increase UV protection and hydrophins to increase better propagation on glaciers and making the ice layer more persistent in time.
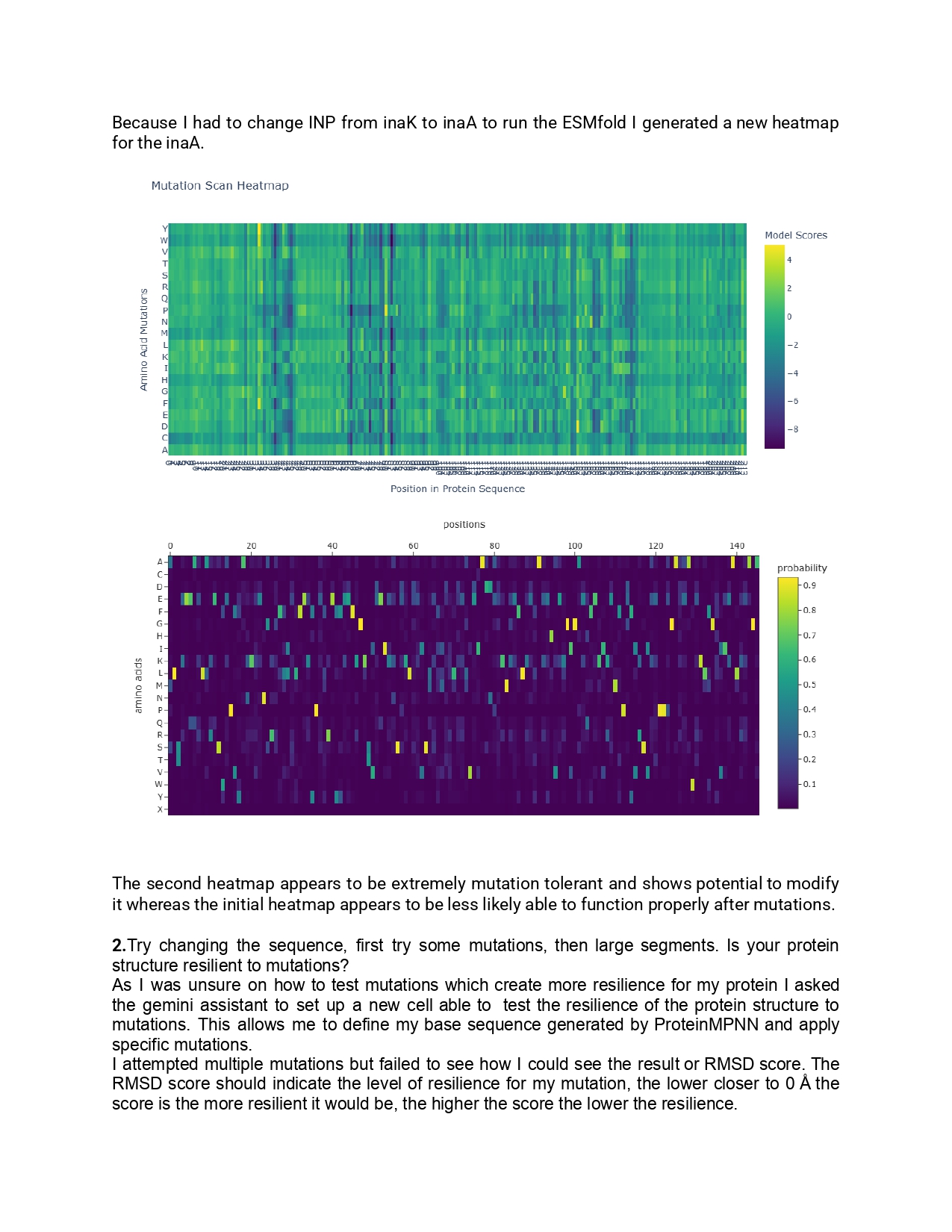
New Protein design