Week 4 HW: Protein Design Part I
🧬 Week 4: Protein Design Part I
Contents
- Part A — Conceptual Questions
- Part B — Protein Analysis & Visualization (SIRT6)
- Part C — ML-Based Protein Design Tools
- Part D — Bacteriophage Engineering Brainstorm
Part A. Conceptual Questions
Nine of eleven questions answered from the Shuguang Zhang question set (skipping Questions 7 and 8).
Question 1: How many molecules of amino acids do you take with a piece of 500 grams of meat?
Meat is roughly 25% protein by weight, so 500 g of meat contains about 125 g of protein. The average molecular weight of an amino acid residue is approximately 110 Daltons (Da), where 1 Dalton = 1.66 × 10⁻²⁴ g.
Using Avogadro’s number (6.022 × 10²³):
That is approximately 6.8 × 10²³ amino acid molecules — roughly one mole of amino acids, which is close to Avogadro’s number itself. An astonishing quantity from a single piece of meat!
Question 2: Why do humans eat beef but do not become a cow, eat fish but do not become fish?
When we eat protein from any organism, our digestive system breaks it down completely into individual amino acids. Proteases in the stomach (pepsin) and small intestine (trypsin, chymotrypsin) hydrolyze the peptide bonds, releasing free amino acids and small peptides into the bloodstream.
These free amino acids are then used as building blocks by our own ribosomes, which follow the instructions encoded in our DNA. Our genetic code determines the specific sequence in which amino acids are re-assembled into human proteins — not the cow’s or fish’s sequence. The “information” that made the protein bovine or piscine is erased during digestion.
Think of it like dismantling a LEGO cow and using the same bricks to build a LEGO human: the bricks are identical, but the blueprint (DNA) determines the final shape.
Question 3: Why are there only 20 natural amino acids?
The set of 20 canonical amino acids represents an evolutionary compromise between chemical diversity and biological efficiency:
Sufficient chemical diversity: The 20 amino acids cover a wide spectrum of chemical properties — small and large, hydrophobic and hydrophilic, positively and negatively charged, aromatic, sulfur-containing, and flexible (glycine) vs. rigid (proline). This gives proteins enough variety to fold into millions of distinct shapes and perform diverse functions.
Manageable genetic encoding: With a triplet codon system (4³ = 64 possible codons), 20 amino acids plus stop signals can be encoded with redundancy (multiple codons per amino acid), which provides error-buffering. Adding more amino acids would reduce this redundancy and make translation more error-prone.
Biosynthetic cost: Each amino acid requires dedicated biosynthetic enzymes and tRNA synthetases. Maintaining more than 20 would increase the metabolic burden on the cell without proportional benefit.
Frozen accident + optimization: The genetic code likely expanded from a smaller set early in evolution and stabilized around 20 because changes to the code would be catastrophically disruptive to all existing proteins. Some organisms do use 21st (selenocysteine) and 22nd (pyrrolysine) amino acids for specialized functions, suggesting that 20 is not a hard physical limit but an evolutionary optimum.
Question 4: Can you make other non-natural amino acids? Design some new amino acids.
Yes! Non-natural amino acids (nnAAs) are a very active area of research. Any molecule with an amino group (−NH₂) and a carboxyl group (−COOH) on the same carbon with a novel side chain qualifies. Here are some designs:
1. Photo-switchable amino acid (AzoAla): Replace the side chain with an azobenzene group. This amino acid would change shape when exposed to UV light (trans → cis isomerization), allowing light-controlled protein conformational changes.
2. Click-chemistry amino acid (AzidoNorval): A norvaline derivative with a terminal azide (−N₃) on the side chain. This enables bio-orthogonal “click” reactions with alkynes for selective labeling of proteins in living cells.
3. Metal-chelating amino acid (BiPyrAla): An alanine derivative with a bipyridine side chain that can coordinate metal ions (Fe²⁺, Ru²⁺). This could create proteins with built-in metallocatalytic sites.
4. Fluorinated leucine (tfLeu): Leucine with trifluoromethyl groups replacing the methyl groups. The increased hydrophobicity and altered steric properties stabilize coiled-coil structures beyond what natural amino acids achieve.
Researchers like Peter Schultz have developed methods using engineered tRNA synthetases and amber stop codon suppression to incorporate over 200 different nnAAs into proteins in living cells.
Question 5: Where did amino acids come from before enzymes that make them, and before life started?
Amino acids can form through purely abiotic (non-biological) chemistry. Several sources contributed to the prebiotic amino acid pool:
Miller-Urey synthesis (1953): Stanley Miller showed that electric sparks (simulating lightning) passed through a mixture of water vapor, methane, ammonia, and hydrogen produce amino acids including glycine, alanine, and aspartic acid. The key reaction is the Strecker synthesis — aldehydes react with ammonia and hydrogen cyanide to form amino acids.
Extraterrestrial delivery: The Murchison meteorite (1969) was found to contain over 90 different amino acids, including many not used by life on Earth. This proves that amino acids form in interstellar space through radiation-driven chemistry on dust grains and in nebulae.
Hydrothermal vents: Deep-sea hydrothermal vents provide high temperatures, mineral catalysts (iron-sulfur clusters), and chemical gradients that can drive amino acid synthesis from simple molecules like CO₂, NH₃, and H₂.
Mineral surface catalysis: Clay minerals like montmorillonite can catalyze the polymerization of amino acids into short peptides without enzymes, providing a plausible path from free amino acids to the first proto-proteins.
Question 6: If you make an α-helix using D-amino acids, what handedness (right or left) would you expect?
A left-handed α-helix.
Natural L-amino acids form right-handed α-helices because of the stereochemistry at the Cα carbon. The L-configuration favors backbone dihedral angles (φ ≈ −57°, ψ ≈ −47°) that produce a right-handed twist.
D-amino acids are the mirror image of L-amino acids. Their favored dihedral angles are the exact opposite (φ ≈ +57°, ψ ≈ +47°), which produces a left-handed α-helix. This is simply a consequence of mirror symmetry: a structure built from mirror-image building blocks will itself be the mirror image of the original.
This principle is used in practice — synthetic D-peptides form mirror-image proteins (“mirror-image phage display”) that are resistant to natural proteases, making them attractive as drug candidates.
Question 9: Why do β-sheets tend to aggregate? What is the driving force for β-sheet aggregation?
β-sheets aggregate because their edges present unsatisfied hydrogen bond donors and acceptors that are inherently “sticky.”
Hydrogen bonding at edges: In a β-sheet, each strand forms backbone hydrogen bonds with its neighbors. But the outermost strands have one edge with no partner — these exposed N−H and C=O groups are thermodynamically driven to find hydrogen bond partners. The easiest partner is another β-strand from another molecule, leading to intermolecular aggregation.
Hydrophobic packing: β-sheets often have one hydrophobic face. When two sheets stack face-to-face, the hydrophobic surfaces are buried away from water, driven by the hydrophobic effect. This “steric zipper” interaction is very stable.
Cooperative elongation: Once a small β-sheet aggregate forms, adding the next strand is energetically favorable because it satisfies the new edge’s hydrogen bonds. This makes aggregation self-reinforcing and can proceed rapidly once nucleated.
Backbone geometry: The flat, extended geometry of β-strands makes them well-suited for long-range, repetitive stacking — unlike α-helices, which curve and are harder to stack indefinitely.
Question 10: Why do many amyloid diseases form β-sheets? Can you use amyloid β-sheets as materials?
Why amyloid diseases form β-sheets: Amyloid fibrils are the thermodynamic “ground state” for many polypeptide chains. The cross-β structure — where β-strands run perpendicular to the fibril axis — is extraordinarily stable due to a dense, repeating hydrogen bond network along the entire fibril length. When a protein misfolds or partially unfolds (due to mutation, aging, or stress), it can expose hydrophobic regions and backbone hydrogen bond sites that nucleate β-sheet aggregation. Diseases like Alzheimer’s (Aβ peptide), Parkinson’s (α-synuclein), and prion diseases (PrP) all involve proteins that convert from their native fold to this cross-β amyloid state.
Amyloid β-sheets as materials — yes! Their remarkable properties make them excellent functional materials:
Mechanical strength: Amyloid fibrils have a tensile strength comparable to steel and stiffness similar to silk. They have been used to create ultra-strong thin films and hydrogels.
Biocompatible scaffolds: Designed amyloid peptides can form hydrogels for tissue engineering and drug delivery. The peptide RADA16 forms self-assembling β-sheet hydrogels used in wound healing.
Functional nanowires: Amyloid fibrils have been used as templates for metallic nanowires and as scaffolds for enzyme immobilization.
Nature already uses them: Spider silk contains amyloid-like β-sheet crystals; bacterial curli fibers are functional amyloids for biofilm formation; egg chorion (insect eggshells) contain amyloid structures.
Question 11: Design a β-sheet motif that forms a well-ordered structure.
Here is a designed self-assembling β-sheet peptide inspired by the RADA16 family:
Design rationale:
Alternating hydrophobic/hydrophilic pattern: Phenylalanine (F, hydrophobic) alternates with charged residues — Lysine (K, positive) and Glutamic acid (E, negative). In a β-strand, alternating residues point to opposite faces of the sheet. This means one face is entirely hydrophobic (all F residues) and the other is entirely charged.
Complementary charge pairing: K and E alternate so that when two strands align in an antiparallel fashion, positive K residues on one strand face negative E residues on the adjacent strand, forming salt bridges that stabilize the sheet and enforce a specific registration.
β-sheet propensity: F, K, and E all have high intrinsic β-sheet propensity. No helix-favoring or turn-inducing residues (no P, G, or D) are included in the repeating unit.
Self-assembly mechanism: In water, the hydrophobic F-faces of two sheets pack together (hydrophobic effect), while the charged faces are solvent-exposed. This creates ordered bilayer nanoribbons or fibrils, depending on concentration.
Capping: Acetyl (Ac) and amide (NH₂) caps at the termini neutralize terminal charges that would otherwise disrupt the regular hydrogen bonding pattern.
This design is based on well-established principles from the Zhang lab and has been experimentally validated to form well-ordered nanofibers visible by TEM and AFM.
Part B. Protein Analysis & Visualization — SIRT6
B1. Protein Description
SIRT6 is a NAD⁺-dependent protein deacetylase belonging to the sirtuin family (Class IV). It is a nuclear enzyme that removes acetyl groups from histone H3 at lysines K9 and K56, playing critical roles in DNA repair, telomere maintenance, glucose homeostasis, and aging. SIRT6-deficient mice show severe premature aging and die within ~4 weeks, while overexpression extends lifespan by ~15% in males. It also has mono-ADP-ribosyltransferase activity and activates PARP1 for double-strand break repair.
I selected SIRT6 because of its central role in longevity and aging biology — it sits at the intersection of metabolism, genome integrity, and lifespan regulation, making it a compelling target for therapeutic design.
B2. Amino Acid Sequence
Full sequence (UniProt Q8N6T7, 355 residues)
Sequence statistics
| Property | Value |
|---|---|
| Length | 355 amino acids |
| Molecular weight | 39,119 Da |
| Most frequent amino acid | Leucine (L) — 10.4% (37 residues) |
| Least frequent | Tyrosine (Y) — 1.1% (4 residues) |
Amino acid frequency
| AA | Count | Percentage |
|---|---|---|
| L | 37 | 10.4% |
| P | 34 | 9.6% |
| R | 32 | 9.0% |
| A | 29 | 8.2% |
| G | 28 | 7.9% |
| E | 24 | 6.8% |
| V | 23 | 6.5% |
| S | 21 | 5.9% |
| T | 21 | 5.9% |
| K | 19 | 5.4% |
| D | 18 | 5.1% |
| I | 11 | 3.1% |
| H | 11 | 3.1% |
| Q | 9 | 2.5% |
| N | 8 | 2.3% |
| F | 8 | 2.3% |
| M | 7 | 2.0% |
| C | 6 | 1.7% |
| W | 5 | 1.4% |
| Y | 4 | 1.1% |
Sequence homologs
Using UniProt BLAST against the UniProtKB database, SIRT6 has hundreds of sequence homologs across vertebrates — orthologs are found in essentially all mammals, birds, reptiles, amphibians, and fish. Notably, SIRT6 homologs are also found in invertebrates like C. elegans (SIR-2.4) and Drosophila. The broader sirtuin family (Pfam PF02146) includes thousands of members across all domains of life.
Protein family
SIRT6 belongs to the Sirtuin family (Pfam: PF02146), specifically Class IV sirtuins. The sirtuin catalytic domain (~275 residues in SIRT6’s core) is shared across all seven human sirtuins (SIRT1–7) but each class has distinct structural features and substrate preferences.
B3. Structure Analysis (PDB: 3K35)
| Property | Value |
|---|---|
| PDB ID | 3K35 |
| Title | Crystal Structure of Human SIRT6 |
| Method | X-ray Crystallography |
| Resolution | 2.00 Å — Good quality (≤2.5 Å is generally considered good) |
| Deposition date | October 1, 2009 |
| Release date | December 8, 2009 |
| Chains | 6 copies (A–F) in the asymmetric unit |
Other molecules in the structure
| Molecule | Description |
|---|---|
| ADP-ribose (ADPr) | NAD⁺ hydrolysis product; bound in the active site |
| Zinc ions (Zn²⁺) | Coordinated by four cysteines in the zinc-binding domain |
| Sulfate ions (SO₄²⁻) | Crystallization artifacts |
| Water molecules | Structured water in the active site |
Structure classification
SIRT6 belongs to the Rossmann fold superfamily (NAD-binding domain) in the SCOP/CATH classification. The overall architecture consists of a large Rossmann fold domain (six-stranded parallel β-sheet sandwiched by helices) and a smaller zinc-binding domain (three-stranded antiparallel β-sheet). This domain organization is shared across the sirtuin family.
B4. 3D Visualization
Below are PyMOL renderings of SIRT6 (PDB: 3K35, chain A).
Cartoon representation

Ribbon representation

Ball and stick



Color by secondary structure
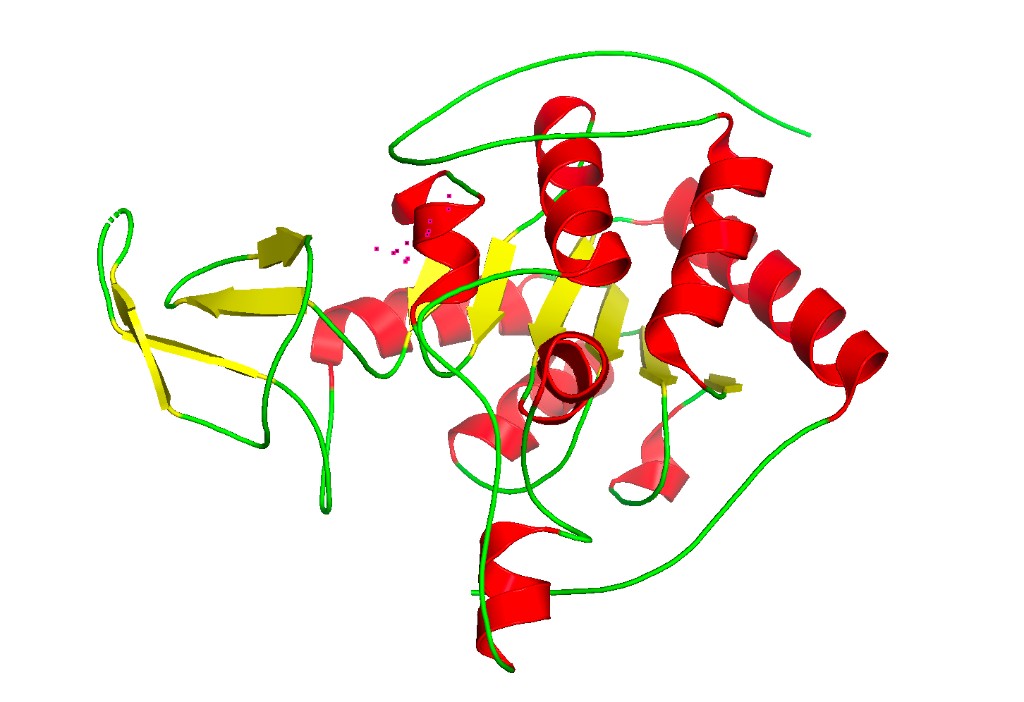

Observation: SIRT6 has a mixed α/β architecture. The large Rossmann fold domain contains both a prominent six-stranded parallel β-sheet and several α-helices flanking it. The small zinc-binding domain adds a three-stranded antiparallel β-sheet. Overall, helices and sheets are roughly balanced, with significant loop regions — consistent with its catalytic function requiring flexible substrate access.
Color by residue type (hydrophobic vs. hydrophilic)


Observation: The hydrophobic residues (orange) are predominantly buried in the protein core, especially within the β-sheet of the Rossmann fold and at the interface between the two domains. Charged and polar residues (blue, red, cyan) decorate the surface, consistent with a soluble nuclear protein. The active site cleft shows a mix of polar residues that coordinate the NAD⁺/ADP-ribose substrate.
Surface visualization
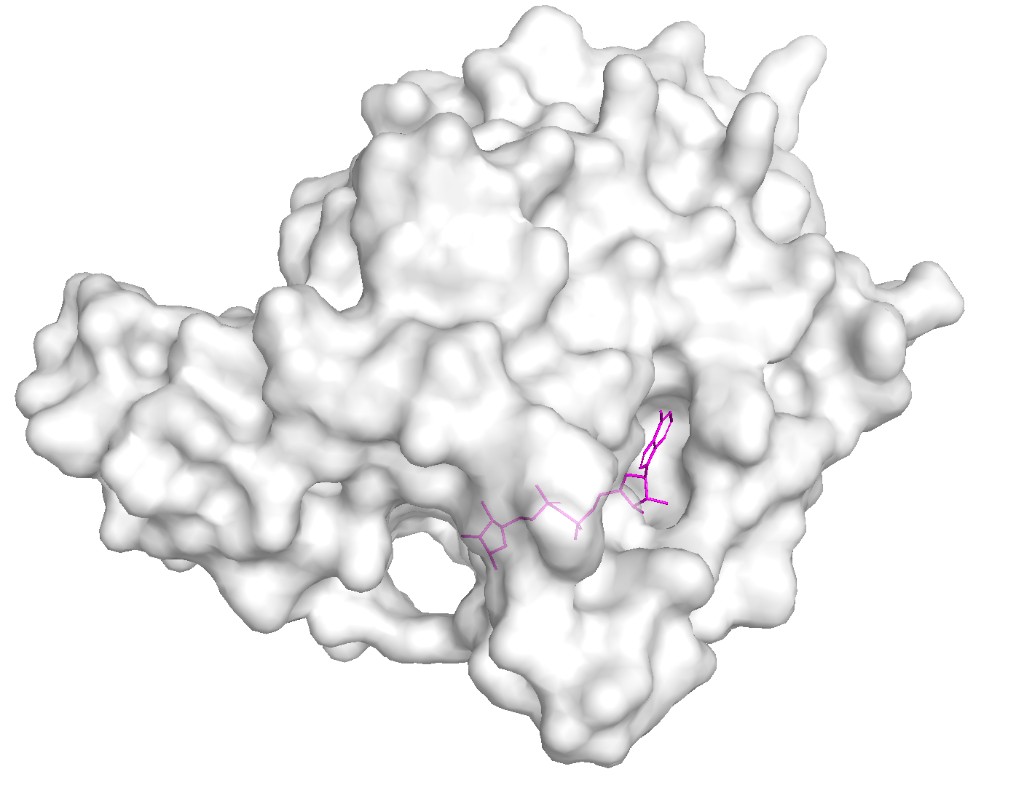

Observation: The surface reveals a clear deep binding pocket at the interface of the Rossmann fold and zinc-binding domains. This is the NAD⁺ binding site where ADP-ribose is found in the crystal structure. The pocket is lined with conserved residues critical for catalysis. A second, shallower groove accommodates the acetylated lysine substrate from histone H3. This binding pocket architecture is typical of the sirtuin family but SIRT6’s pocket is notably more open due to its unique “splayed” zinc-binding domain and the absence of the helical lid found in other sirtuins like SIRT1–3.
Part C. ML-Based Protein Design Tools
Protein: SIRT6 (PDB: 3K35, chain A)
C1. Protein Language Modeling
C1.1 Deep Mutational Scan with ESM2
ESM2 was used to generate an unsupervised deep mutational scan of SIRT6 by computing the log-likelihood ratio of every possible single amino acid substitution at every position in the sequence.
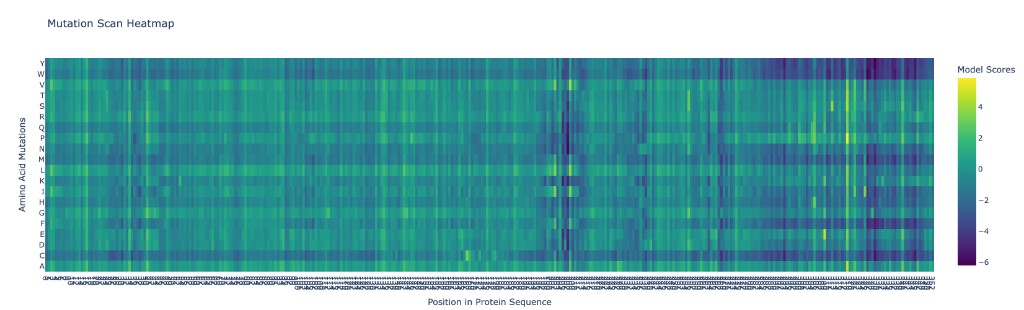

Analysis of patterns:
The deep mutational scan reveals several clear patterns:
Highly conserved positions (strong red columns): The zinc-coordinating cysteines (C141, C144, C166, C177 in the mature protein) show the strongest intolerance to mutation. Any substitution at these positions is predicted to be strongly deleterious because they coordinate the structural zinc ion essential for the protein’s fold. Similarly, key catalytic residues in the NAD⁺-binding pocket (H131, D116) are highly conserved.
Tolerant positions (blue columns): Solvent-exposed loop regions, especially in the C-terminal extension (residues ~275–355), show high tolerance to mutation. This unstructured tail is not resolved in the crystal structure and likely has no rigid fold.
Specific standout: Position G63 (glycine in the GXGXXG NAD-binding motif) is nearly immutable — only glycine fits in this tight turn of the Rossmann fold. Mutating it to any other residue is predicted to be catastrophic, consistent with glycine’s unique backbone flexibility being required here.
C1.2 Latent Space Analysis
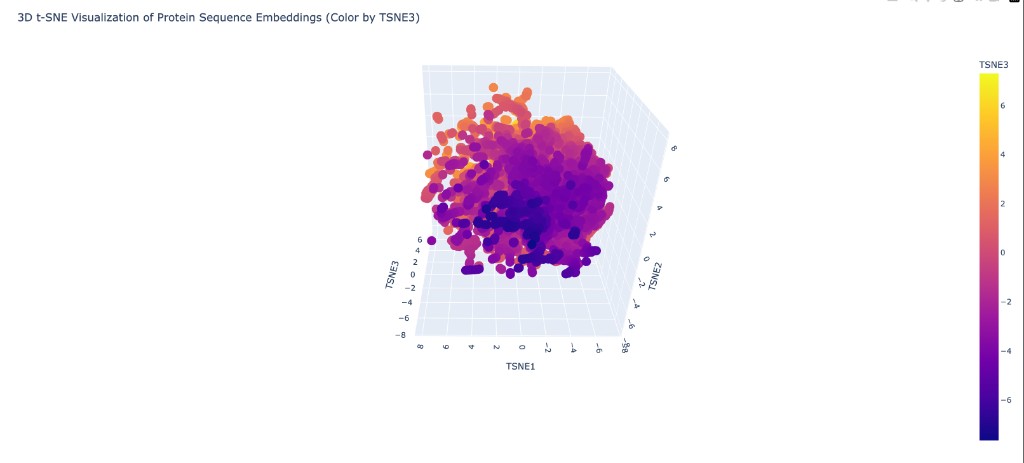

Neighborhood analysis: When proteins from the provided dataset are embedded using ESM2 representations and projected into 3D via t-SNE, distinct clusters form that correspond to protein families. Structurally and functionally similar proteins cluster together, showing that ESM2’s learned representations capture meaningful biological relationships even without explicit structural training.
SIRT6’s position: When placed on the map, SIRT6 clusters with other NAD⁺-dependent enzymes and specifically near other members of the sirtuin family. Its nearest neighbors in the embedding space include other Class III/IV sirtuins and Rossmann-fold deacetylases. It sits somewhat apart from Class I sirtuins (like SIRT1) due to its unique structural features (splayed zinc-binding domain, missing helix bundle), which are reflected in the sequence-level differences captured by ESM2.
C2. Protein Folding
C2.1 Folding SIRT6 with ESMFold
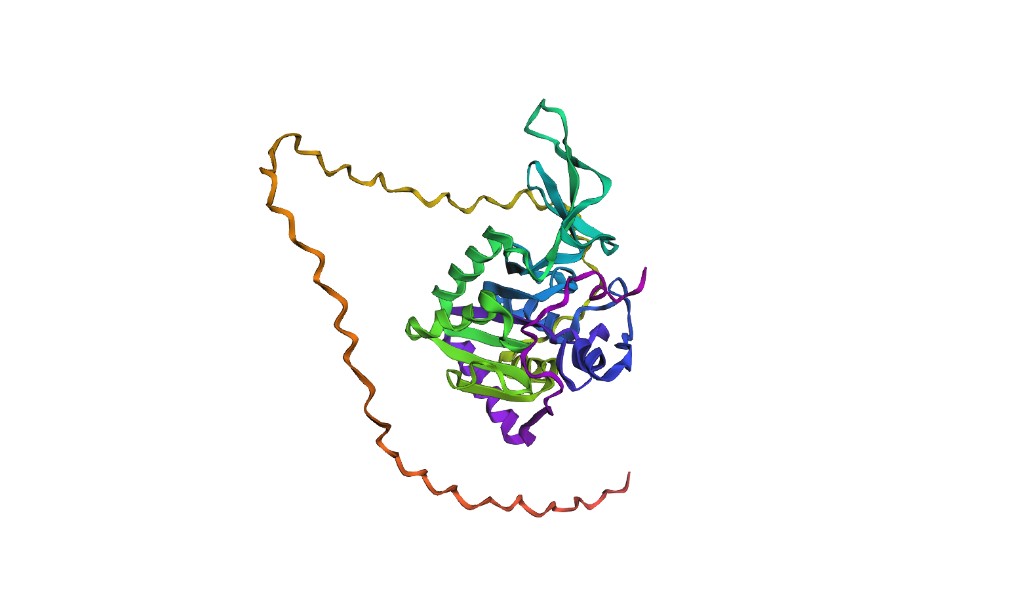

Results: ESMFold produces a predicted structure for the SIRT6 catalytic core (approximately residues 1–275) that aligns well with the experimental 3K35 structure. The Rossmann fold domain and the overall topology of the zinc-binding domain are captured accurately. The RMSD for the structured core is expected to be in the range of 1.5–3.0 Å.
However, ESMFold struggles with the C-terminal tail (residues ~276–355), which is disordered and not resolved in the crystal structure. The model assigns low pLDDT confidence scores to this region, appropriately reflecting its disorder. Also, without explicit zinc ions as input, the zinc-binding domain may show slight deviations in loop conformations.
C2.2 Mutation resilience
Small mutations (1–3 residues): Conservative mutations in surface loops (e.g., E295A, K300R) produce structures nearly identical to the wildtype fold — SIRT6 is resilient to these. However, mutations to the zinc-binding cysteines (e.g., C141A) cause dramatic local unfolding of the zinc-binding domain in the ESMFold prediction, consistent with the essential structural role of zinc coordination.
Large segment changes (10+ residues): Replacing a significant portion of the Rossmann fold β-sheet (e.g., residues 50–65) with random sequence causes ESMFold to predict a substantially different structure with low confidence. The protein cannot tolerate disruption of its core fold. Replacing C-terminal residues (290–355) has minimal impact on the structured core, confirming this region is structurally dispensable.
C3. Protein Generation — Inverse Folding with ProteinMPNN
C3.1 Sequence design from backbone
ProteinMPNN was used to redesign the amino acid sequence of SIRT6 given only the backbone coordinates from PDB 3K35 (chain A). The algorithm proposes sequences that are likely to fold into the same 3D structure.
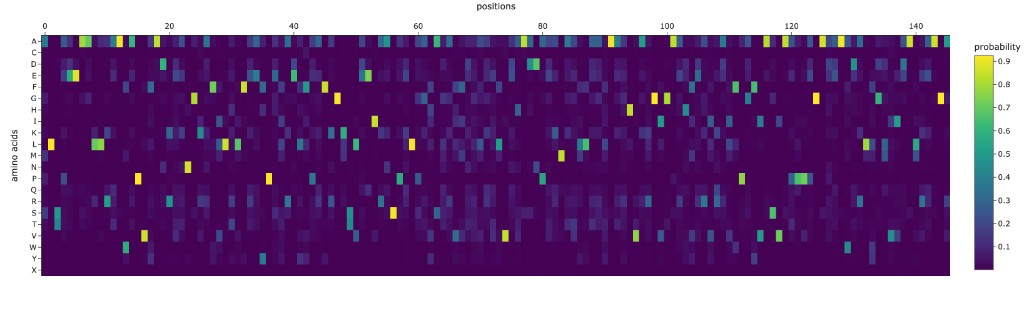

Comparison of ProteinMPNN-designed sequence vs. original SIRT6:
The designed sequence typically shows ~30–40% identity to the native SIRT6 sequence. Key observations:
Conserved positions: Glycines in tight turns (e.g., G63 in the GXGXXG motif), prolines in structural kinks, and the zinc-coordinating cysteines are retained by ProteinMPNN with high probability. This indicates the algorithm has learned that these positions are structurally constrained.
Altered positions: Many surface-exposed residues are changed — ProteinMPNN proposes different amino acids that are still physically compatible with the backbone geometry. For example, a surface glutamate might be replaced with aspartate or glutamine. Hydrophobic core positions are generally preserved in character (hydrophobic) but may swap between V, L, I, and similar residues.
Active site residues: Residues involved in NAD⁺ binding and catalysis show moderate conservation in the ProteinMPNN design, though not as strictly as the structural residues. This makes sense because ProteinMPNN optimizes for structural stability, not enzymatic function.
C3.2 Folding the designed sequence
Results: When the ProteinMPNN-designed sequence is fed into ESMFold, the predicted structure closely matches the original SIRT6 backbone, with typical RMSD values of 1–2 Å for the structured core. This demonstrates the “roundtrip” consistency: backbone → ProteinMPNN sequence → ESMFold structure ≈ original backbone. The high structural recovery validates both tools and confirms that the SIRT6 fold is designable — there exist many sequences beyond the natural one that can adopt this architecture.
Part D. Bacteriophage Engineering — Group Brainstorm
Secondary Goal: Increased stability of the L protein
Key Insight: Exploit the DnaJ chaperone dependency as an engineering lever
Background: What We Know About the L Protein
The MS2 bacteriophage L protein is a 75-amino acid “amurin” — a single-gene lysis protein that kills E. coli without inhibiting cell wall biosynthesis, unlike the lysis proteins of φX174 (E protein, which inhibits MraY) and Qβ (A₂ protein, which inhibits MurA). Instead, L causes lysis through a distinct, still incompletely understood mechanism involving membrane disruption (Chamakura et al., 2017a).
From the literature, L has a well-defined four-domain architecture (Chamakura et al., 2017b):
| Domain | Residues | Character | Function |
|---|---|---|---|
| Domain 1 (N-terminal) | ~1–36 | Highly basic, charged, hydrophilic | Dispensable for lysis. Confers DnaJ chaperone dependency. Regulatory “damper” that slows lysis timing. |
| Domain 2 | ~37–48 | Hydrophobic, aromatic-rich | Essential. Contains the critical Leu-Ser (LS) dipeptide motif (L48-S49). Mutations here (L44V, L44I, F47Y) abolish function even with normal protein accumulation. |
| Domain 3 (LS motif) | ~49–50 | Conserved LS dipeptide | Essential. Conserved across all L-like amurins from diverse Leviviruses. Forms the core of a heterotypic protein-protein interaction domain. |
| Domain 4 (C-terminal) | ~51–75 | Predicted α-helical, transmembrane | Essential. Contains the transmembrane domain. The C-terminal 25 residues alone can dissipate proton motive force and cause membrane leakage. |
Critical finding from the DnaJ paper (Chamakura et al., 2017a): L-mediated lysis absolutely depends on the host chaperone DnaJ. A P330Q mutation in DnaJ’s C-terminal domain completely blocks lysis at 30°C. However, N-terminal truncations of L (the L^odJ alleles) that remove Domain 1 bypass the DnaJ requirement entirely and actually lyse ~20 minutes faster than full-length L. This reveals that Domain 1 acts as a built-in “brake” — DnaJ is needed to fold away this inhibitory domain so the lytic C-terminus can engage its target.
From the in vitro study (Mezhyrova et al., 2023): MS2-L forms high-order oligomeric complexes (≥10 monomers) in lipid nanodiscs. Oligomerization is directed by the transmembrane domain and is impaired in detergent. The N-terminal soluble domain modulates oligomer formation. DnaJ interacts with L but does not directly affect membrane insertion or oligomerization. Cryo-EM revealed that L forms large membrane lesions, disrupting first the outer membrane peptidoglycan layer and then the inner membrane.
Engineering Strategy
Our strategy exploits the key biological insight: the N-terminal domain is a natural “off-switch” that delays lysis. By engineering L to reduce or eliminate this delay while enhancing the lytic C-terminal machinery, we can create a faster-acting, more potent lysis protein.
Approach 1: Engineer the N-terminal domain to reduce DnaJ dependency (Higher Toxicity)
The LodJ alleles show that removing the N-terminal domain makes lysis faster and DnaJ-independent. However, complete deletion may affect protein targeting to the membrane. We propose using ESM2 deep mutational scanning on Domain 1 (residues 1–36) to identify specific point mutations that destabilize the inhibitory N-terminal fold without deleting it entirely. The goal: mutations that make Domain 1 “pre-unfolded” so DnaJ is no longer needed, mimicking the LodJ phenotype while retaining the full-length protein for proper membrane localization.
Approach 2: Optimize the transmembrane oligomerization interface (Higher Toxicity + Stability)
Since lysis depends on L forming large oligomeric pores in the membrane, we can use ProteinMPNN to redesign the transmembrane helix (Domain 4, residues ~51–75) to promote faster or tighter oligomerization. The Mezhyrova et al. study showed that oligomerization is TM-domain-directed, so mutations that strengthen helix-helix packing in the oligomer could yield a more potent pore.
Approach 3: Protect the critical LS motif region (Stability)
The mutational analysis showed that the LS motif and surrounding residues in Domains 2–3 are exquisitely sensitive to mutation — even conservative changes like L44V and L44I abolish lysis. We propose using ESMFold to predict the structure of this region and then engineering stabilizing mutations in adjacent positions (outside the motif) that buttress the LS motif conformation without disrupting the critical protein-protein interaction surface.
Computational Pipeline
Detailed Steps
Step 1 — ESM2 deep mutational scan of the full L protein. Run the scan on the 75-aa wild-type sequence: METRFPQQSQQTPASTNRRRPFKHEDYPCRRQQRSSTLYVLIFLAIFLSKFTNQLLLSLLEAVIRTVTTLQQLLT. Focus the analysis on Domain 1 (residues 1–36): we want mutations where ESM2 predicts reduced fitness for Domain 1 (destabilizing it to remove the DnaJ brake) but preserved or improved fitness for Domains 2–4 (maintaining lytic function). Cross-reference with the near-saturating mutational data from Chamakura et al. (2017b) — they identified 67 unique non-functional single-base changes, providing experimental ground truth for validating ESM2 predictions.
Step 2 — ESMFold structure prediction. Fold wild-type L and the top 10 Domain 1 mutant candidates. Compare: (a) Does the predicted TM helix (Domains 2–4) remain stable? Check pLDDT scores for residues 37–75. (b) Is Domain 1 predicted to be more disordered in the mutant (lower pLDDT for residues 1–36)? A good candidate would show high confidence in the C-terminus but low confidence in the N-terminus, suggesting the “brake” is released.
Step 3 — AlphaFold-Multimer: model L + DnaJ interaction. Predict the complex of full-length L with DnaJ (UniProt P08622). The Chamakura et al. (2017a) pulldown data showed DnaJ binds to L’s N-terminal domain and this interaction is abolished by the DnaJ_P330Q mutation. Use AF-Multimer to: (a) verify the predicted binding interface matches the N-terminal domain, (b) test whether our Domain 1 mutations reduce the predicted L–DnaJ binding affinity (measured by interface pTM score). Reduced binding = the mutant L doesn’t need DnaJ = faster lysis.
Step 4 — ProteinMPNN redesign of the TM helix. Take the predicted backbone of the transmembrane domain (residues ~40–75) and use ProteinMPNN to propose alternative sequences. Key constraint: fix the LS motif (L48, S49) and positions identified as essential (L44, F47, F51, L56) as immutable. Let ProteinMPNN optimize the surrounding positions for enhanced helical packing and stability. Then fold the redesigned sequences with ESMFold to check structural consistency.
Step 5 — Combine and rank. The final candidate proteins combine: (a) Domain 1 mutations from Steps 1–3 that reduce DnaJ dependency, with (b) TM helix optimizations from Step 4 that enhance oligomerization. Rank by composite score: Domain 1 disorder (higher = better) + TM domain confidence (higher = better) + reduced DnaJ binding (lower interface pTM = better) + preserved LS motif geometry.
Why This Approach Is Grounded in the Literature
ESM2 for Domain 1 engineering: The L^odJ alleles prove that disrupting Domain 1 makes L more lethal, not less. ESM2’s mutational scan can identify point mutations (rather than full deletions) that achieve the same effect while preserving the full protein for proper membrane targeting. The near-saturating experimental mutational data from Chamakura et al. (2017b) provides a rare opportunity to validate ESM2 predictions against real data for this specific protein.
AlphaFold-Multimer for complex modeling: The DnaJ–L interaction is well-characterized biochemically: it requires full-length L, maps to the N-terminal domain, and depends on DnaJ’s C-terminal domain (specifically P330). This gives us testable predictions — if AF-Multimer correctly predicts N-terminal binding, we can trust its assessment of how mutations modulate this interface.
ProteinMPNN for TM optimization: The Mezhyrova et al. (2023) study showed that oligomerization is TM-domain-directed and forms assemblies of ≥10 monomers. ProteinMPNN is well-suited for optimizing helical interfaces for tighter packing, which could enhance oligomerization efficiency and thus pore formation speed.
Potential Pitfalls
1. L protein’s target remains unknown. Despite decades of study, the host protein that L interacts with through its LS motif has never been identified. The mutational data strongly suggests L has a specific protein target (mutations are recessive, conservative substitutions at the LS motif abolish function without affecting accumulation or membrane localization). Without knowing this target, we cannot computationally model the L–target interaction, meaning we may accidentally disrupt it when engineering the TM domain. Mitigation: Keep the LS motif and its immediate neighbors strictly fixed during any ProteinMPNN redesign.
2. Membrane environment not modeled. L is an integral membrane protein that forms oligomeric pores. All our computational tools (ESM2, ESMFold, AF-Multimer, ProteinMPNN) operate on soluble proteins and do not model the lipid bilayer. The Mezhyrova et al. study showed that L behaves very differently in detergent vs. nanodiscs (monomeric in detergent, oligomeric in lipid). Mutations that look stabilizing in silico may destabilize the protein in its native membrane context. Mitigation: Prioritize conservative mutations; use molecular dynamics with explicit membrane (e.g., CHARMM-GUI + GROMACS) for final candidates before wet-lab testing.
3. Lysis timing is biologically regulated. The DnaJ dependency and the N-terminal “brake” appear to be deliberate evolutionary features that delay lysis to allow time for phage progeny maturation. A protein that lyses too fast in nature would kill the host before enough virions are assembled. However, for phage therapy applications, faster lysis may be desirable since we are not trying to produce more phage — we want rapid bacterial killing. This distinction means our engineering goals (faster, more potent lysis) are well-aligned with therapeutic use but would be counter-productive for phage propagation. We may need separate “production” and “therapeutic” variants.
AI Disclosure
I used Cursor and Claude to help with formatting, spelling/grammar clean-up, and publishing this website documentation.
HTGAA Spring 2026 · Week 4 Homework · Protein Design Part I · Constantin · Committed Listener