Week 7 HW: Gene Synthesis & Genome Engineering
🧬 Week 7: Gene Synthesis & Genome Engineering
Part 1. Intracellular Artificial Neural Networks (IANNs)
Question 1: Advantages of IANNs over Traditional Boolean Genetic Circuits
Traditional genetic circuits implement Boolean logic — genes are essentially ON or OFF, and circuits are built by wiring together AND, OR, and NOT gates. While powerful for simple decisions, this approach has fundamental limitations that IANNs overcome:
| Feature | Boolean Genetic Circuits | IANNs (Perceptron-Based) |
|---|---|---|
| Signal type | Digital / binary (ON or OFF) | Analog / continuous (graded output across a spectrum of input concentrations) |
| Multi-input integration | Requires cascading multiple logic gates, which becomes unwieldy with many inputs | A single perceptron neuron inherently integrates many weighted inputs with a tunable threshold — elegant and modular |
| Noise tolerance | Limited — molecular noise can cause erratic switching near the threshold. Trade-offs between amplitude and frequency detection | Analog signal processing naturally handles noisy biological environments. Feed-forward architectures can simultaneously filter both amplitude and frequency noise |
| Adaptability | Static once designed — changing function requires rewiring gates | Different computational functions can be implemented by tuning weights and thresholds in the same circuit topology (no rewiring needed) |
| Computational power | Cannot solve nonlinearly separable problems (like XOR) without complex multi-layer gate cascades | Multilayer perceptrons can learn curved, complex decision boundaries — solving XOR and beyond with fewer components |
| Programmability | Each function requires a unique circuit topology | One framework can encode minimum, maximum, average, soft majority, analog-to-digital conversion, and ternary switches — all from the same basic architecture |
| Dynamic range | Information compressed into binary states — fine-grained signal information is lost | High output dynamic range preserves continuous signal information with high computational precision |
In summary, IANNs exploit the inherently analog nature of biology (continuous protein concentrations, graded promoter responses) rather than fighting it. A single perceptron equation replaces layers of logic gates, making circuits simpler to design, more robust to noise, and far more flexible in the computations they can perform.
Question 2: Application: Multi-Biomarker Cancer Diagnostic IANN
System overview
An IANN-based diagnostic circuit inside engineered immune cells (e.g., CAR-T cells) that detects circulating tumor markers and classifies cancer risk by integrating multiple biomarker signals simultaneously — something a Boolean circuit would struggle to do with graded, noisy biological inputs.
Input/output behavior
Inputs (analog biomarker signals):
| Input | Biomarker | Sensing mechanism |
|---|---|---|
| X₁ | Cancer antigen CA-125 concentration | Engineered protein-binding domain converts extracellular CA-125 to intracellular transcription factor activity |
| X₂ | Metastasis-associated microRNA (miR-373) abundance | Complementary RNA binding sequences regulate an internal reporter |
| X₃ | Phosphorylated tyrosine kinase activity | Synthetic phospho-responsive protein interaction triggers gene expression |
Computation: The perceptron computes a weighted sum: if (w₁·X₁ + w₂·X₂ + w₃·X₃ − threshold) exceeds a decision boundary, classify as “high cancer risk.” Unlike a Boolean AND gate that requires all markers above a sharp threshold, the IANN performs soft classification — one strong marker plus two moderate ones can still trigger a positive result, better reflecting clinical reality.
Output: Graded GFP fluorescence proportional to cancer risk score (low = healthy, high = danger). Above a critical threshold, a second output activates: synthesis of a therapeutic cytokine (e.g., IL-2) to recruit immune cells to the tumor site.
Limitations
Biological noise: Stochastic fluctuations in mRNA/protein levels can cause the perceptron output to oscillate around the decision boundary, leading to false positives/negatives. Robust threshold setting and temporal integration would be needed.
Weight tuning: The weights w₁, w₂, w₃ must be calibrated to clinically relevant biomarker ranges. Weights optimized for one patient population may not generalize to others.
Metabolic burden: Expressing multiple sensor proteins, the computational circuitry, and therapeutic outputs creates significant metabolic load on the host cell, potentially affecting viability and computational fidelity.
Scalability: Expanding to 10+ biomarkers requires wider input layers or multilayer architectures, increasing complexity and potential crosstalk between genetic components.
Leakiness: Genetic components are never perfectly switch-like — leaky transcription and variable Hill coefficients introduce nonlinearities not perfectly captured by the idealized perceptron model.
Question 3: Multilayer Perceptron Circuit Architecture
Multilayer Perceptron Diagram
Below is my perceptron diagram for the intracellular multilayer perceptron. Layer 1 integrates two transcription factor inputs and produces the endoribonuclease Csy4. Layer 2 uses Csy4 to regulate GFP output through cleavage of the GFP mRNA.
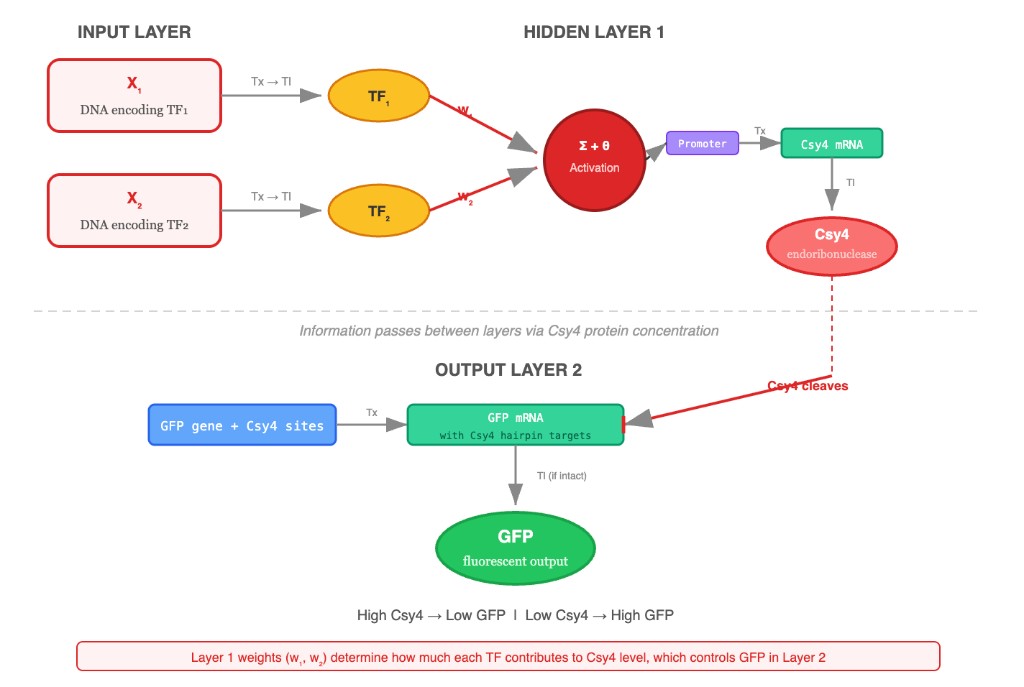
How information flows
Layer 1 (Hidden Layer): Two upstream input signals (e.g., promoter activities driven by small molecules like IPTG and aTc) are integrated through weighted regulation. The “weights” are implemented biologically as promoter strengths and ribosome binding site (RBS) efficiencies — stronger promoters or optimized RBS sequences correspond to higher weights (w₁, w₂). The summed transcriptional output drives expression of the endoribonuclease Csy4. The bias term (θ) corresponds to basal promoter leakiness.
Layer 2 (Output Layer): A constitutive promoter drives GFP mRNA that contains a Csy4 recognition hairpin (28-nt sequence) in its 5’ UTR. When Csy4 is present, it specifically cleaves this hairpin, destabilizing the GFP mRNA and reducing fluorescent output. This creates an inhibitory (inverting) connection — shown with a blunt-end bar in the diagram:
- High Layer 1 activation → high Csy4 concentration → extensive GFP mRNA cleavage → low GFP fluorescence
- Low Layer 1 activation → low Csy4 concentration → intact GFP mRNA → high GFP fluorescence
Key advantage over single-layer: The hidden layer performs a weighted, nonlinear transformation of the raw inputs before passing the result to the output layer. This enables the circuit to compute functions (like XOR) that a single-layer perceptron fundamentally cannot.
Part 2. Fungal Materials
Question 1: Existing Fungal Materials: Examples, Uses, Advantages & Disadvantages
| Material | Example Companies | Application | Replaces |
|---|---|---|---|
| Mycelium packaging | Ecovative Design | Protective packaging for electronics, shipping cushioning | Polystyrene (Styrofoam) |
| Mycelium leather | MycoWorks (Reishi™), Bolt Threads (Mylo™) | Fashion, handbags, shoes, accessories | Animal leather, PU leather |
| Mycelium insulation | Mykor (MykoFoam) | Thermal & acoustic insulation for buildings | Polystyrene foam, fiberglass |
| Acoustic panels | Mogu, BuzziSpace | Sound absorption in offices, studios | Synthetic acoustic foam |
| Mycoprotein (food) | Quorn (Marlow Foods) | Meat-alternative protein products | Animal meat |
| Biomedical scaffolds | Research stage | Tissue engineering, wound healing | Synthetic polymer scaffolds |
Advantages over traditional counterparts
Biodegradable and compostable: Mycelium materials decompose in weeks and enrich soil, versus Styrofoam’s 500+ year persistence in landfills.
Grows on waste: Mycelium feeds on abundant agricultural waste (hemp hurd, straw, sawdust), turning waste streams into valuable materials — a circular economy in action.
Carbon-neutral production: No petroleum extraction or energy-intensive manufacturing. The fungus metabolizes substrate carbohydrates at ambient temperature.
Customizable properties: By varying fungal species, substrate, and growth conditions, material properties (density, strength, flexibility) can be tuned for specific applications.
Fire resistance: Mycelium composites are naturally self-extinguishing with low smoke production — superior to many synthetic foams.
Fast production: Packaging materials grow in 5–10 days; leather in 4–9 days.
Disadvantages
Moisture sensitivity: Mycelium composites can absorb >200% of their weight in water, causing mechanical weakening. This limits outdoor applications.
Low mechanical strength: Compressive strength of 0.05–0.18 MPa restricts use to non-structural, non-load-bearing applications.
Cost: Currently more expensive than petroleum-based alternatives (polystyrene costs a few cents per cubic foot).
Scalability: Manufacturing processes are still being standardized. Batch-to-batch consistency remains a challenge.
Short lifespan for packaging: The very biodegradability that makes mycelium appealing also limits shelf life for long-distance shipping.
Consumer perception: “Grown” materials can face skepticism about durability and quality compared to established synthetics.
Question 2: Genetic Engineering of Fungi & Advantages Over Bacteria
What would I engineer fungi to do?
1. Enhanced material properties: Engineer mycelium to produce hydrophobic surface compounds (reducing water absorption from >200% to <30%), express structural proteins that reinforce hyphal networks (increasing compressive strength 10-fold), or deposit mineral compounds during growth for fire resistance. This would address the main disadvantages of current mycelium materials.
2. Bioremediation of plastics: Fungi naturally secrete powerful extracellular enzymes (laccases, peroxidases, hydrolases) that can break polymer bonds. Engineering Aspergillus or Penicillium species to overexpress plastic-degrading enzymes and broaden their substrate specificity could accelerate degradation of polyethylene, polystyrene, and polyurethane from months to days.
3. Complex protein production: Engineer filamentous fungi to produce recombinant spider silk (284.9 kDa, similar to natural), collagen for medical implants (avoiding BSE/prion risk from animal sources), or therapeutic antibodies. Fungi can properly fold these complex proteins in ways bacteria cannot.
4. Living materials with embedded function: Engineer mycelium to produce pigments, antimicrobial compounds, or even conductive polymers as it grows — creating materials with built-in functionality (self-coloring, self-sterilizing, or electrically conductive composites).
Advantages of synthetic biology in fungi vs. bacteria
| Advantage | Why It Matters |
|---|---|
| Eukaryotic post-translational modifications | Fungi perform glycosylation, disulfide bond formation, and proteolytic processing — essential for producing functional therapeutic proteins. Bacteria like E. coli largely lack these systems, so recombinant eukaryotic proteins often misfold or aggregate into inclusion bodies. |
| Protein secretion | Fungi naturally secrete proteins into the culture medium, enabling direct harvesting without cell lysis. Bacteria typically trap heterologous proteins inside the cell, requiring expensive extraction and refolding. |
| Cellular compartmentalization | Fungal cells have organelles (ER, Golgi, mitochondria, peroxisomes) that maintain distinct chemical environments. Competing metabolic pathways can be physically separated — impossible in bacteria’s single cytoplasm. |
| Growth on cheap waste substrates | Fungi efficiently degrade cellulose, hemicellulose, and lignin from agricultural waste. Most bacteria require purified media. This dramatically reduces production costs. |
| 3D structure formation | Mycelium naturally forms interconnected 3D hyphal networks that can be shaped into materials. Bacteria form flat biofilms at best — they cannot create self-supporting 3D structures. |
| Rich secondary metabolism | A single fungal species can produce 100+ different secondary metabolites (antibiotics, immunosuppressants, pigments). Whole-genome sequencing reveals tens of thousands of unexplored biosynthetic gene clusters in fungi — far more than in bacteria. |
| Larger genome capacity | Fungal genomes can accommodate larger, more complex transgene inserts. Better codon adaptation for expressing eukaryotic proteins. More similar to animal/plant cells for heterologous expression. |
Part 3. First DNA Twist Order — EZH2 Y726D Expression Cassette
Step 1 — Choose Your Protein
| Property | Details |
|---|---|
| Protein | EZH2 (Enhancer of Zeste Homolog 2) |
| Organism | Homo sapiens (Human) |
| UniProt ID | Q15910 |
| Function | Catalytic subunit of Polycomb Repressive Complex 2 (PRC2). Trimethylates histone H3 at lysine 27 (H3K27me3), a key repressive epigenetic mark involved in gene silencing and cell fate decisions. |
| Length | 746 amino acids |
| Mutation | Y726D — Tyrosine → Aspartic acid at position 726. This mutation in the SET domain active site abolishes methyltransferase activity while preserving the protein’s structural scaffold. |
| Rationale | By expressing catalytically dead EZH2, we can study non-canonical (methylation-independent) functions of PRC2 in partial reprogramming contexts — e.g., chromatin compaction, transcription factor recruitment, and RNA binding. |
Step 2 — Amino Acid Sequence (EZH2 Y726D)
Full-length EZH2 protein sequence from UniProt Q15910, with the Y726D mutation (position 726: Y → D, highlighted below):
The mutation site is in the SET domain catalytic pocket. Wild-type has Y726 (Tyrosine), which participates in SAM cofactor positioning. Replacing it with D (Aspartic acid) disrupts methyltransferase activity while maintaining overall protein folding.
Step 3 — Reverse Translation & Codon Optimization
The EZH2 Y726D protein sequence was reverse-translated to DNA and codon-optimized for human (mammalian) expression using the most frequently used codons from the Homo sapiens codon usage table (Kazusa database). Key optimization metrics:
| Metric | Value |
|---|---|
| CDS length | 2,238 bp (746 codons) |
| GC content | 65.6% |
| Internal stop codons | 0 (verified) |
| Codon Adaptation Index (CAI) | ~1.0 (all most-frequent human codons) |
| Rare codons eliminated | Yes — no codons below 10% usage frequency |
| Back-translation verified | ✓ DNA translates back to exact Y726D protein |
Step 4 — Expression Cassette Design
The insert is designed for cloning into the pTwist CMV mammalian expression backbone from Twist Bioscience. Since the backbone already provides the CMV promoter and bGH polyA terminator, the insert contains:
(in backbone)
GCCACC
(Start)
2,238 bp (746 aa)
CAC×6
(Stop)
(in backbone)
◀ 5′ ——— INSERT (2,265 bp, ordered from Twist) ——— 3′ ▶
| Component | Sequence / Source | Length |
|---|---|---|
| Kozak consensus | GCCACC (optimal mammalian ribosome initiation) | 6 bp |
| Start codon | ATG (first codon of CDS) | 3 bp |
| EZH2 Y726D CDS | Human codon-optimized, Y726D mutation at bp 2173–2175 | 2,238 bp |
| 6×His tag | CACCACCACCACCACCAC (C-terminal, for Ni-NTA purification) | 18 bp |
| Stop codon | TAA (universal stop, preferred in mammals) | 3 bp |
| Total insert length | 2,265 bp |
Step 5 — Backbone Vector
| Property | Details |
|---|---|
| Backbone | pTwist CMV (Twist Bioscience mammalian expression vector) |
| Promoter | CMV (Cytomegalovirus) — strong, constitutive mammalian promoter |
| Terminator | bGH polyA signal (bovine Growth Hormone polyadenylation) |
| Selection marker | Ampicillin resistance (AmpR) for bacterial cloning |
| Cloning method | Insert arrives pre-cloned in backbone from Twist |
| Expression host | HEK293T or similar mammalian cell line (transient transfection) |
| Why this vector? | CMV promoter drives high-level expression in mammalian cells. Since we are studying epigenetic reprogramming, the protein must be expressed in a mammalian context where histone modifications are biologically relevant. |
Step 6 — Complete Insert DNA Sequence (2,265 bp)
This is the full insert to be ordered from Twist Bioscience as a clonal gene in pTwist CMV:
Step 7 — Twist Bioscience Order Summary
| Order Parameter | Value |
|---|---|
| Product | Clonal Gene (in vector) |
| Gene name | EZH2_Y726D |
| Insert length | 2,265 bp |
| GC content | 65.6% |
| Backbone vector | pTwist CMV |
| Species optimization | Homo sapiens |
| Verification | Sequence-verified by Twist (Sanger sequencing) |
| Deliverable | Lyophilized plasmid DNA, ready for transformation and transfection |
EZH2_Y726D_protein.fasta and EZH2_Y726D_insert.fasta.Design Rationale Summary
The EZH2 Y726D catalytically dead mutant is the cornerstone construct for our final project on non-canonical PRC2 functions in partial reprogramming. By expressing a version of EZH2 that can still assemble into PRC2 and bind chromatin but cannot methylate H3K27, we can decouple the enzymatic activity from the structural scaffolding role. This allows us to ask: does PRC2 contribute to reprogramming through histone methylation, or through physical chromatin organization and transcription factor sequestration?
The mammalian codon optimization ensures high expression levels in HEK293T or iPSC-derived cells. The C-terminal 6×His tag enables purification and western blot detection without disrupting the N-terminal domains critical for PRC2 complex assembly (EED and SUZ12 binding interfaces).
AI Disclosure
I used Claude (Anthropic) to help with: formatting and structuring this homework page, codon optimization strategy for the EZH2 Y726D construct, expression vector design rationale, biological multilayer perceptron diagram design, and spelling/grammar clean-up throughout the document.
HTGAA Spring 2026 · Week 7 Homework · Gene Synthesis & Genome Engineering · Constantin · Committed Listener