Week 11 HW: Bioproduction & Cloud Labs
Week 11 — Bioproduction & Cloud Labs
Constantin Convalexius · Lifefabs Node
Part A — Pixel Artwork
I placed two anchor designs early in the top-left corner: a Lifefabs logo and an MIT logo. Both were eventually overwritten as more people filled the canvas, which was actually the interesting part.
What I liked most was the emergence. Many uncoordinated people created patterns that still made sense in the end. It felt like swarm intelligence, like ants in a colony. The art and the experiment were almost the same thing: local actions becoming a collective pattern.
For next year, I would improve the anti-bot / anti-script rules. I tested the system and found that scripted placement was possible. Either rate-limit per user, add a CAPTCHA, or make bots an official part of the challenge.
Part B — Cell-Free Reagents
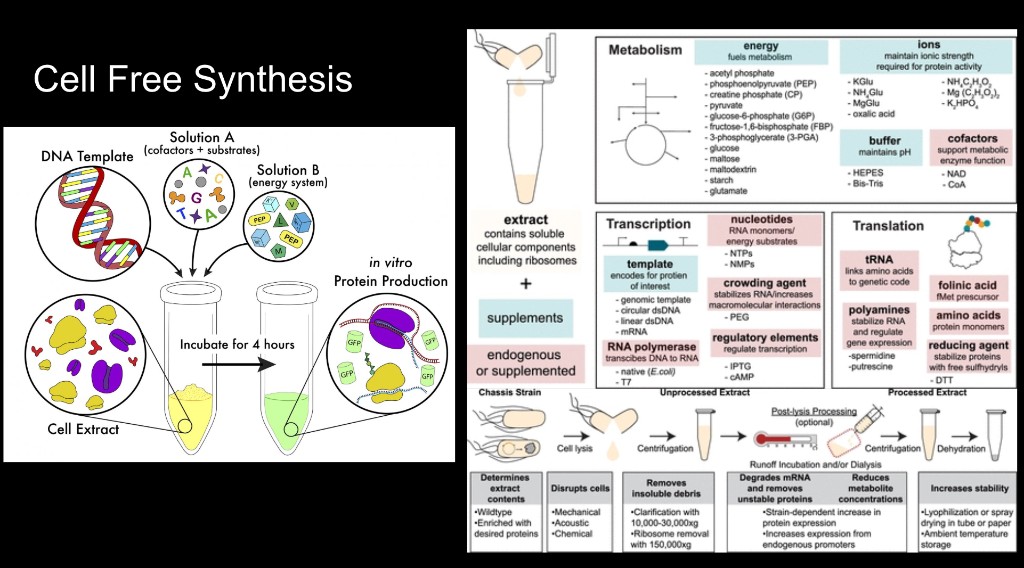

Q1. Component Roles
E. coli BL21 (DE3) Star lysate with T7 RNAP: the engine of the reaction. It provides ribosomes, tRNAs, translation factors, chaperones, metabolic enzymes, and T7 RNA polymerase.
Potassium glutamate: sets ionic strength and helps ribosomes stay stable. Glutamate is gentler than chloride for transcription and translation.
HEPES-KOH pH 7.5: keeps the pH stable during the reaction without strongly chelating Mg2+.
Magnesium glutamate: the most important ion in the mix. Mg2+ stabilizes rRNA, ribosome assembly, polymerases, and ATP-dependent enzymes.
Potassium phosphate monobasic + dibasic: buffer the reaction and support phosphate chemistry, but too much free phosphate can bind Mg2+ and hurt translation.
Ribose + glucose: feed metabolism in the lysate so ATP can be regenerated slowly over many hours.
AMP, CMP, GMP, UMP: nucleotide monophosphates. Lysate kinases can phosphorylate them into NTPs when needed.
Guanine: can be salvaged into GMP, helping refill the GTP pool.
17 amino acid mix + tyrosine + cysteine: amino acids for translation. Tyrosine and cysteine are separated because they are less stable in mixed stocks.
Nicotinamide: inhibits NAD-consuming enzymes so NAD+ remains available for metabolism.
Nuclease-free water: fills the reaction volume without adding nucleases that would degrade DNA or RNA.
Q2. 1-Hour PEP-NTP Mix vs 20-Hour NMP-Ribose-Glucose Mix


The 1-hour PEP mix is a fast burst system. It already contains NTPs and uses PEP for ATP regeneration, but every PEP-to-ATP cycle releases inorganic phosphate. Phosphate binds free Mg2+, and when free Mg2+ drops too low, ribosomes fall apart and translation shuts down.
The 20-hour NMP-ribose-glucose mix is slower but more stable. Instead of front-loading energy, it uses glucose and ribose metabolism to regenerate ATP gradually. That avoids the phosphate/Mg2+ crash and keeps the reaction alive longer.
For a 36-hour artwork reaction, I would push the same logic further: stronger buffering, enough potassium for ionic strength, enough Mg2+ to survive chelation, and enough amino acids so translation does not run out of monomers.
Bonus — Transcription Without GMP
The lysate can use nucleotide salvage. Guanine can be converted into GMP through salvage enzymes, then phosphorylated to GDP and GTP by lysate kinases. It is slower than adding GMP directly, but it can still support transcription and translation.
Part C — Master Mix Design
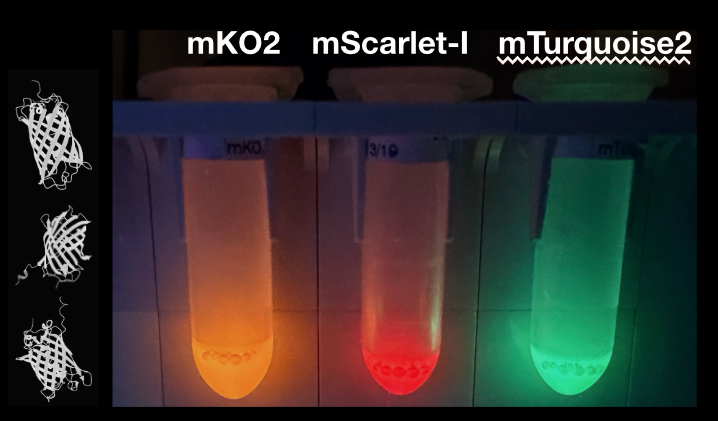


Q1. Fluorescent Protein Properties
sfGFP: fast folding and fast maturation. It is the safest positive control for CFPS because it usually expresses well and becomes fluorescent quickly.
mRFP1: slower maturation and oxygen-dependent chromophore formation. It needs more time and good oxygen access to become bright.
mKO2: orange fluorescent protein with relatively fast maturation, but still oxygen-dependent. pH and folding quality affect the readout.
mTurquoise2: bright cyan fluorescent protein with high quantum yield. It is useful when you want strong signal with less maturation delay than many red proteins.
mScarlet-I: bright red fluorescent protein with good maturation and pH stability. It is one of the better red options for CFPS.
Electra2: far-red protein with slower maturation and less CFPS optimization history. It may need longer incubation and better oxygen handling.
Q2. Hypothesis
For mRFP1, I would increase Mg2+ slightly and reduce reaction volume. More Mg2+ should keep ribosomes stable longer, while smaller volume increases air-water surface area and helps oxygen reach the reaction. The expected effect is stronger red fluorescence after 36 hours because mRFP1 needs both protein production and chromophore oxidation.
Q3. Master Mix Composition
To complete after receiving the assigned well / fluorescent protein instructions. The design will be submitted through the CFPS composition tool.
Q4. Data Analysis
To complete when the fluorescence data returns. I would compare fluorescence endpoint values across reagent compositions and normalize by the expected protein color channel.
Part D — Build-A-Cloud-Lab
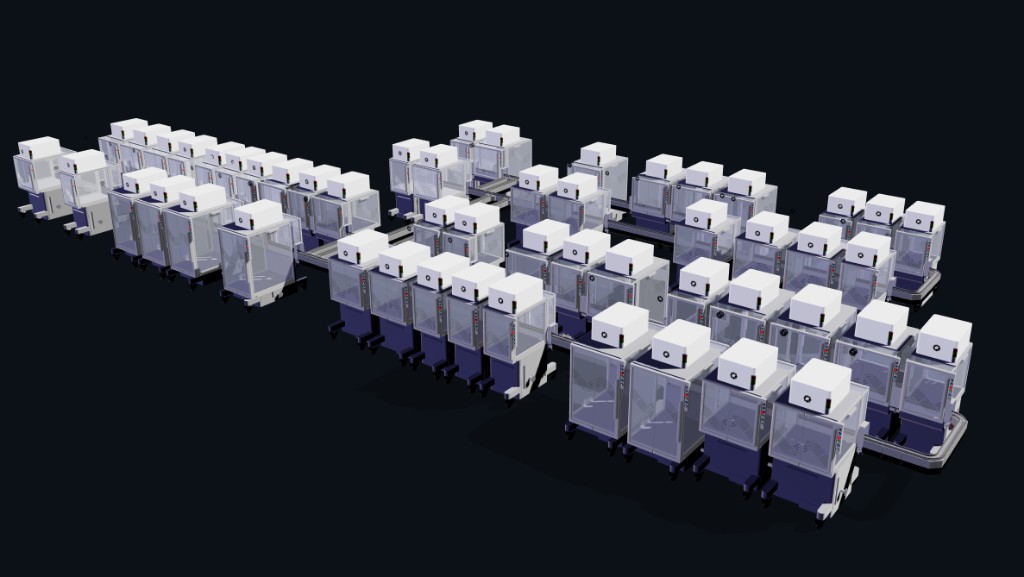


This is my cloud lab rendering from the RAC simulation tool. I like how the carts look modular but still coordinated, like a physical version of the cell-free artwork experiment: many small units, each simple alone, becoming powerful as a network.
Sources
- FPbase entries for sfGFP, mRFP1, mKO2, mTurquoise2, mScarlet-I, and Electra2.
- HTGAA Week 11 cloud lab recitation slides.
- bioRxiv 2026.02.05.703998 — GPT-5-driven CFPS optimization.
- bioRxiv 2025.08.01.668204 — low-cost CFPS reagent design.