Week 7 — Genetic Circuits Part II: Neuromorphic Circuits
This week covers neuromorphic genetic circuits, showing how engineered gene networks can implement neural-network “perceptron”-like computation and learning.
Lecture (Tues, Mar 17)
Genetic Circuits Part II: Neuromorphic Circuits
(▶️Recording)
Ron Weiss
Recitation (Wed, Mar 18)
Neuromorphic circuits & Biomaterials
(▶️Recording |
💻Slides)
Evan Holbrook, Ren Ramlan
Lab (Thurs-Fri, Mar 19 - 20)
Homework — DUE BY Mar 31 2PM ET
Assignment Part 1: Intracellular Artificial Neural Networks (IANNs)
Assignees for this section
| MIT/Harvard students | Required |
| Committed Listeners | Required |
- What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions?
- Describe a useful application for an IANN; include a detailed description of input/output behavior, as well as any limitations an IANN might face to achieve your goal.
- Below is a diagram depicting an intracellular single-layer perceptron where the X1 input is DNA encoding for the Csy4 endoribonuclease and the X2 input is DNA encoding for a fluorescent protein output whose mRNA is regulated by Csy4. Tx: transcription; Tl: translation.
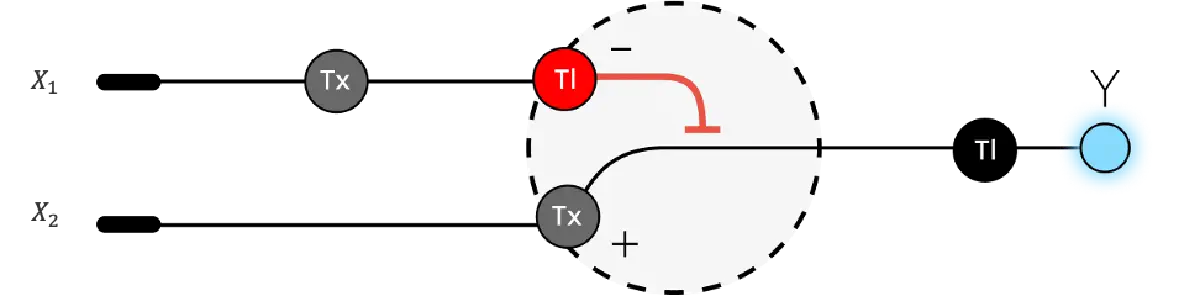
 Draw a diagram for an intracellular multilayer perceptron where layer 1 outputs an endoribonuclease that regulates a fluorescent protein output in layer 2.
Draw a diagram for an intracellular multilayer perceptron where layer 1 outputs an endoribonuclease that regulates a fluorescent protein output in layer 2.
Assignment Part 2: Fungal Materials
Assignees for this section
| MIT/Harvard students | Required |
| Committed Listeners | Required |
- What are some examples of existing fungal materials and what are they used for? What are their advantages and disadvantages over traditional counterparts?
- What might you want to genetically engineer fungi to do and why? What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
Assignment Part 3: First DNA Twist Order
Assignees for this section
| MIT/Harvard students | Required |
| Committed Listeners | Required |
- Review the Individual Final Project documentation guidelines.
- Submit this Google Form with your draft Aim 1, final project summary, HTGAA industry council selections, and shared folder for DNA designs. DUE MARCH 20 FOR MIT/HARVARD/WELLESLEY STUDENTS
- Review Part 3: DNA Design Challenge of the week 2 homework. Design at least 1 insert sequence and place it into the Benchling/Kernel/Other folder you shared in the Google Form above. Document the backbone vector it will be synthesized in on your website.