Week 2 HW: DNA READ WRITE AND EDIT
Part 1: Benchling & In-silico Gel Art
Rectriction enzymes used:
- EcoRI
- HindIII
- BamHI
- KpnI
- EcoRV
- SacI
- SalI
I create a pattern that simulates angel feathers in benchling. I use the tool https://rcdonovan.com/gel-art to guide me c:
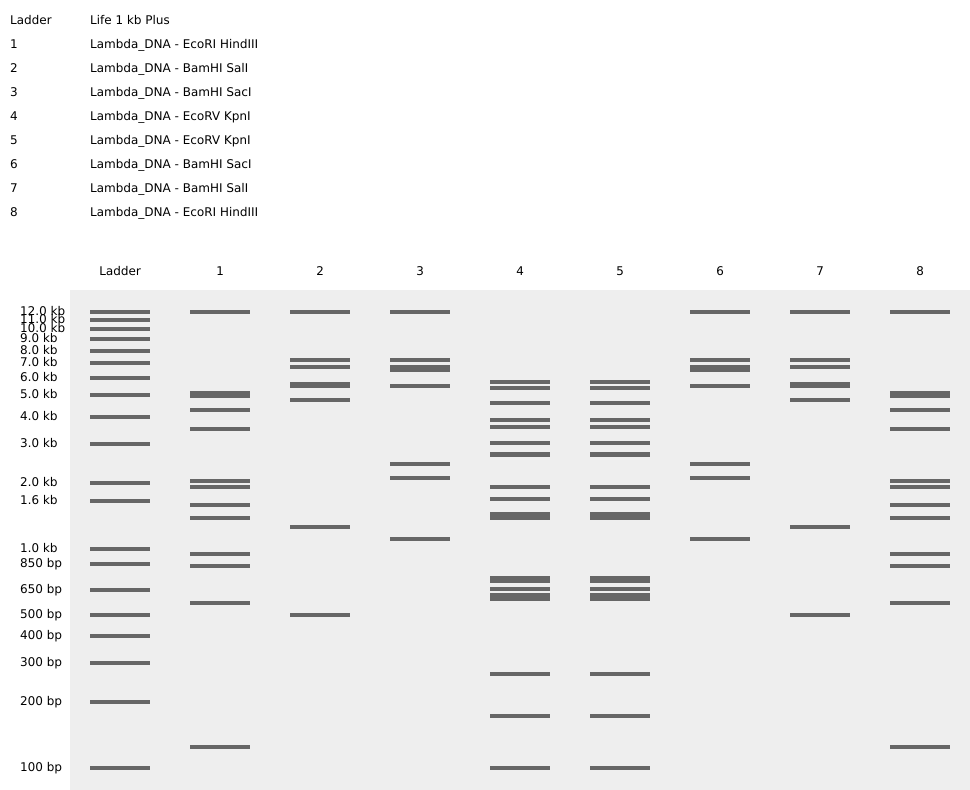

Part 3: DNA Design Challenge
3.1. Choose your protein = Ferrochelatase
Recently I have been reading about therapeutics that target mitochondria. Specifically in the paper of Corson Lab they mention about this protein that when is inhibited causes mitochondrial dysfunction and this is benefitial for anti-angiogenesis purposes in retinal pathologies.
>sp|P22830|HEMH_HUMAN Ferrochelatase, mitochondrial OS=Homo sapiens OX=9606 GN=FECH PE=1 SV=2 MRSLGANMAAALRAAGVLLRDPLASSSWRVCQPWRWKSGAAAAAVTTETAQHAQGAKPQV QPQKRKPKTGILMLNMGGPETLGDVHDFLLRLFLDRDLMTLPIQNKLAPFIAKRRTPKIQ EQYRRIGGGSPIKIWTSKQGEGMVKLLDELSPNTAPHKYYIGFRYVHPLTEEAIEEMERD GLERAIAFTQYPQYSCSTTGSSLNAIYRYYNQVGRKPTMKWSTIDRWPTHHLLIQCFADH ILKELDHFPLEKRSEVVILFSAHSLPMSVVNRGDPYPQEVSATVQKVMERLEYCNPYRLV WQSKVGPMPWLGPQTDESIKGLCERGRKNILLVPIAFTSDHIETLYELDIEYSQVLAKEC GVENIRRAESLNGNPLFSKALADLVHSHIQSNELCSKQLTLSCPLCVNPVCRETKSFFTS QQL
3.2. Reverse Translate
Using Online tools: bioinformatic.com I determined the nucleotide sequence that correponds to Ferrochelatase.
atgcgcagcctgggcgcgaacatggcggcggcgctgcgcgcggcgggcgtgctgctgcgc gatccgctggcgagcagcagctggcgcgtgtgccagccgtggcgctggaaaagcggcgcg gcggcggcggcggtgaccaccgaaaccgcgcagcatgcgcagggcgcgaaaccgcaggtg cagccgcagaaacgcaaaccgaaaaccggcattctgatgctgaacatgggcggcccggaa accctgggcgatgtgcatgattttctgctgcgcctgtttctggatcgcgatctgatgacc ctgccgattcagaacaaactggcgccgtttattgcgaaacgccgcaccccgaaaattcag gaacagtatcgccgcattggcggcggcagcccgattaaaatttggaccagcaaacagggc gaaggcatggtgaaactgctggatgaactgagcccgaacaccgcgccgcataaatattat attggctttcgctatgtgcatccgctgaccgaagaagcgattgaagaaatggaacgcgat ggcctggaacgcgcgattgcgtttacccagtatccgcagtatagctgcagcaccaccggc agcagcctgaacgcgatttatcgctattataaccaggtgggccgcaaaccgaccatgaaa tggagcaccattgatcgctggccgacccatcatctgctgattcagtgctttgcggatcat attctgaaagaactggatcattttccgctggaaaaacgcagcgaagtggtgattctgttt agcgcgcatagcctgccgatgagcgtggtgaaccgcggcgatccgtatccgcaggaagtg agcgcgaccgtgcagaaagtgatggaacgcctggaatattgcaacccgtatcgcctggtg tggcagagcaaagtgggcccgatgccgtggctgggcccgcagaccgatgaaagcattaaa ggcctgtgcgaacgcggccgcaaaaacattctgctggtgccgattgcgtttaccagcgat catattgaaaccctgtatgaactggatattgaatatagccaggtgctggcgaaagaatgc ggcgtggaaaacattcgccgcgcggaaagcctgaacggcaacccgctgtttagcaaagcg ctggcggatctggtgcatagccatattcagagcaacgaactgtgcagcaaacagctgacc ctgagctgcccgctgtgcgtgaacccggtgtgccgcgaaaccaaaagcttttttaccagc cagcagctg
3.3. Codon optimization
Codon optimization is essential because of the degeneracy of the genetic code. Since there are multiple ways to get to some amino acids, different organisms use different codons for this purpose to translate more efficiently in their context. So, if we do not do codon optimization before a synthetic biologic application, the organism will double its efforts trying to read a code it is not familiar with.
Rather than produce this protein in mass, I want to study it without using animal models to extract it. For that reason, I would express it in yeast (Saccharomyces cerevisiae), as it is a eukaryotic model that allows better 3D structure formation compared to E. coli, and post-translational changes in protein, while it is also reproducible and cheap.
For this reason, using online tools:IDT i got the follow codon optimization.
ATG CGT AGT CTA GGT GCT AAC ATG GCA GCA GCA TTG CGT GCG GCA GGG GTG TTG TTG AGA GAT CCA CTG GCC TCC AGC AGT TGG CGT GTT TGT CAG CCG TGG AGA TGG AAA TCA GGA GCA GCA GCC GCA GCG GTT ACA ACG GAA ACT GCT CAA CAT GCA CAA GGC GCC AAG CCT CAG GTT CAG CCT CAA AAA AGA AAG CCT AAA ACA GGG ATT CTG ATG TTA AAC ATG GGT GGA CCT GAA ACC TTG GGA GAC GTC CAT GAT TTC TTA TTA AGA TTA TTT TTG GAC AGG GAT CTT ATG ACC TTA CCC ATT CAA AAT AAA CTG GCC CCT TTT ATT GCC AAA AGG AGA ACT CCG AAA ATT CAG GAA CAA TAT CGT AGG ATT GGC GGC GGA TCT CCC ATT AAG ATT TGG ACT AGC AAA CAA GGA GAG GGA ATG GTT AAG TTG CTA GAC GAG TTG TCT CCA AAT ACT GCA CCC CAC AAA TAC TAT ATT GGT TTT AGG TAT GTT CAT CCA CTA ACT GAA GAA GCT ATA GAA GAA ATG GAA AGG GAC GGC TTG GAG AGA GCT ATA GCC TTT ACT CAA TAC CCA CAA TAC TCC TGT TCT ACA ACA GGT TCT TCC CTT AAT GCT ATT TAT AGA TAC TAC AAT CAA GTT GGT AGA AAG CCT ACG ATG AAA TGG TCA ACG ATT GAC AGA TGG CCC ACC CAT CAT TTA TTG ATA CAA TGT TTT GCT GAT CAC ATT TTA AAG GAA CTT GAT CAC TTT CCT CTA GAG AAA AGA TCA GAA GTG GTA ATA TTA TTC TCA GCA CAT TCT TTA CCA ATG TCA GTA GTG AAT AGA GGA GAT CCC TAT CCT CAA GAA GTC TCT GCG ACT GTT CAA AAG GTT ATG GAA AGG CTA GAG TAC TGT AAC CCT TAT AGG CTT GTC TGG CAA AGC AAA GTA GGA CCA ATG CCA TGG TTG GGT CCA CAG ACT GAT GAA TCC ATA AAG GGT TTG TGT GAA AGA GGT AGA AAA AAT ATT TTG TTA GTA CCG ATC GCA TTT ACC TCA GAC CAT ATT GAA ACC CTT TAC GAA TTG GAT ATT GAA TAC TCT CAA GTT CTA GCG AAA GAA TGC GGT GTT GAA AAC ATA AGA CGT GCT GAA TCC TTG AAT GGT AAT CCC CTG TTT AGT AAG GCT TTA GCG GAT TTG GTG CAC TCC CAT ATA CAG TCC AAT GAG CTG TGC TCA AAA CAA TTA ACC CTA AGC TGT CCT TTG TGC GTA AAT CCT GTG TGT CGT GAA ACC AAG TCT TTC TTC ACA TCT CAA CAA CTA
3.4. Technologies to produce this protein : Cell-dependent method in yeast
1. Clone the gene
I have to insert the codon-optimized DNA into a yeast expression vector: a plasmid. This plasmid must have an appropiate promoter and a selectable marker as an antibiotic resistance gene. Also I would put an affinity tag to purifiy it later easily. To do this I can use technologies as PCR or Gibson assembly.
2. Transform the cells
I introduce thr plasmid into the yeast cells through electroporation (creating pores in cells thanks to a a pulse of electricity)
3. Selecciont of transformed yeast
I make use of the selectable marker to allow only the transform yeast to grow
4. Transcription and translation
Yeast will transcribe the DNA into mRNA and then translate it
5. Harvest and purify the protein
I break (lyse) the yeast cells when the biomass increased, and then purify the protein through affinity chromatography thanks to the affinity tag.
Part 4: Prepare a Twist DNA Synthesis Order
Insert
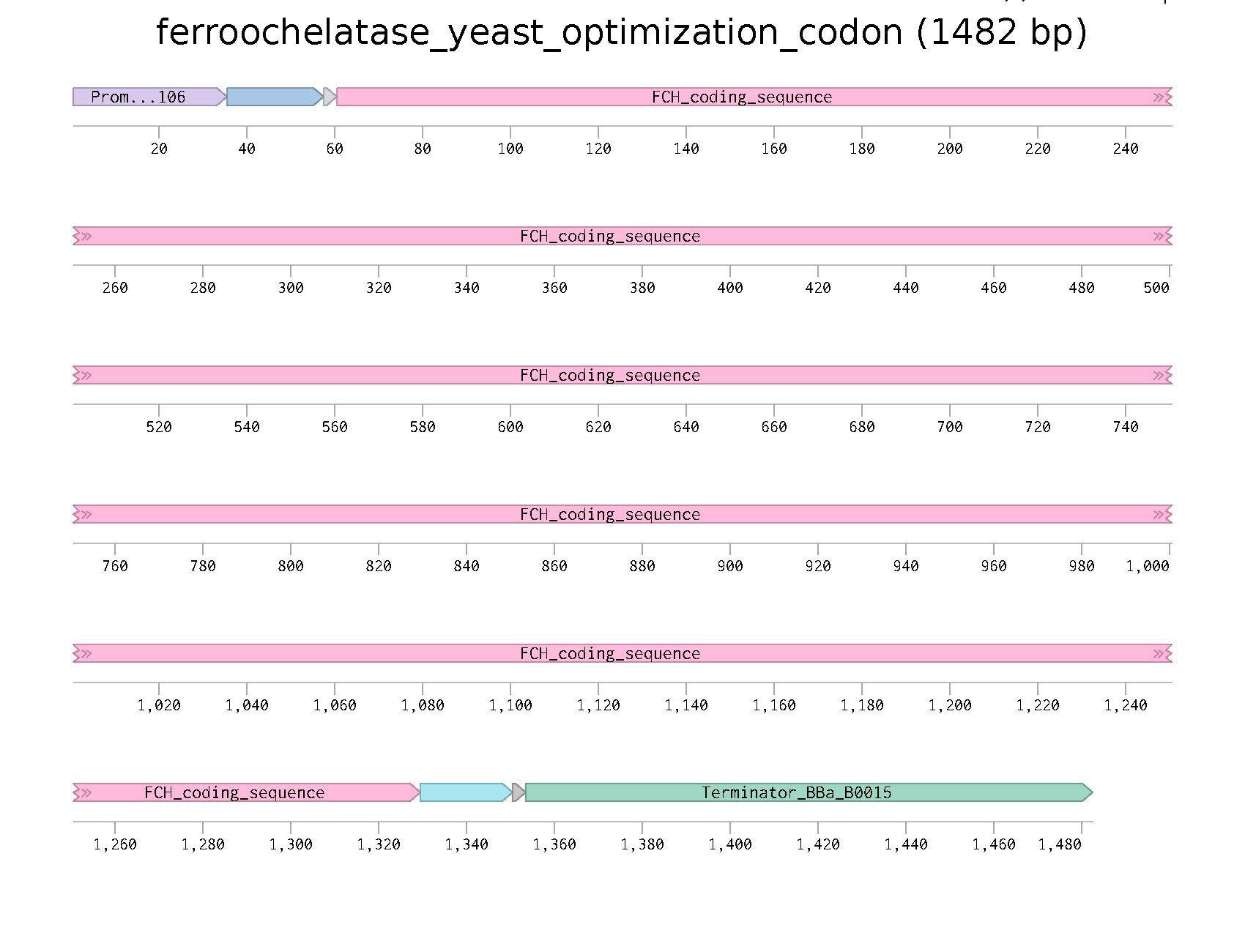

Parts:
- Promoter
- RBS
- Start Codon
- COding sequence
- 7x His Tag
- Stop Codon
- Terminator
Online tools used: Benchling
Plasmid


Vector: pTwist Amp High Copy
Online tools used: Benchling, Twist BIOSCIENCE
Part 5: DNA Read/Write/Edit
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why?
Mitochondrial DNA from tardigrades, I am curious if the protecting proteins for DNA in radiation come from this source.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
I would use Illumina sequencing due to the fidelity of this short fragment sequencing platform.
- Is your method first-, second- or third-generation or other? How so?
My method is second generation because it allows sequencing in parallel huge genomes, but it uses short fragments, so it is second.
- What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps.
My input would be extracted DNA from tardigrades. I would prepare it through DNA extraction kits.
- What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?
The essential steps of my technology are basically to create these clusters of amplification where they read multiple parts/fragments of DNA, and thanks to fluorescent signals all this is recorded.
- What is the output of your chosen sequencing technology?
The output will be the fluorescent signals that can be translated into DNA sequences, a process called “Base calling”, and at the end I get a FASTQ document that also tells me the quality of sequencing.
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why?
I recently read about DNA origami scaffolds used for delivery of epitopes as a form of new HIV vaccine. I would love to synthesize it because it prevents any off-target immune system effect in complex diseases.
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
Method: phosphoramidite chemistry. It is the most common method used, very standard and optimized.
- What are the essential steps of your chosen sequencing methods?
It is basically chemical synthesis, through the cycle: Deblocking, Coupling, Capping, and Oxidation; it adds one nucleotide at a time.
- What are the limitations of your method?
It is not 100% efficient. For long scaffolds, it should be synthesized in truncated fragments and then joined together. Environmentally, it is not as friendly as it requires a lot of organic solvents. So it would not be good for scaling up, which is essential for vaccines.
5.1 DNA Edit
(i) What DNA would you want to edit and why?
I would love to edit genetic risk factors so people are less prone to develop some diseases. For example, the allele APOE4 is known for its predisposition to cause Alzheimer, a serious neurodegenerative disease which has no definitive cure yet. So I would propose to edit the DNA from gametes to avoid people having this predisposition.
(ii) What technology or technologies would you use to perform these DNA edits and why?
It has to be very precise, so I would use CRISPR-CAS9, as it nowadays is one of the more precise ways of editing DNA.
- How does your technology of choice edit DNA? What are the essential steps?
Basically it is made of two parts: a protein CAS9 that can cut the DNA and a guide RNA that specifically guides you to the place where it has to be cut. If you also insert a template DNA, the cell can use this to repair the cut DNA and do a precise modification.
- What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
I would design a guide RNA that specifically targets APOE4 and a DNA template with the APOE2 sequence for repair. The inputs include the guide RNA, Cas9 protein, the repair template, and the cells to be edited: gametes. Additional reagents like buffers may also be needed.
- What are the limitations of your editing methods (if any) in terms of efficiency or precision?
CRISPR-CAS9 may have off-target effects if the guide RNA is not perfectly specific. It is somewhat “aggressive” because it breaks the DNA double strand, so efficiency and precision can vary depending on the cell type and delivery method. Previous trials in murine cells may not reveal all secondary effects. Finally, editing in gametes raises a lot of ethical issues that must be considered.