Week 3 HW: Lab Automation
Python Script for Opentrons
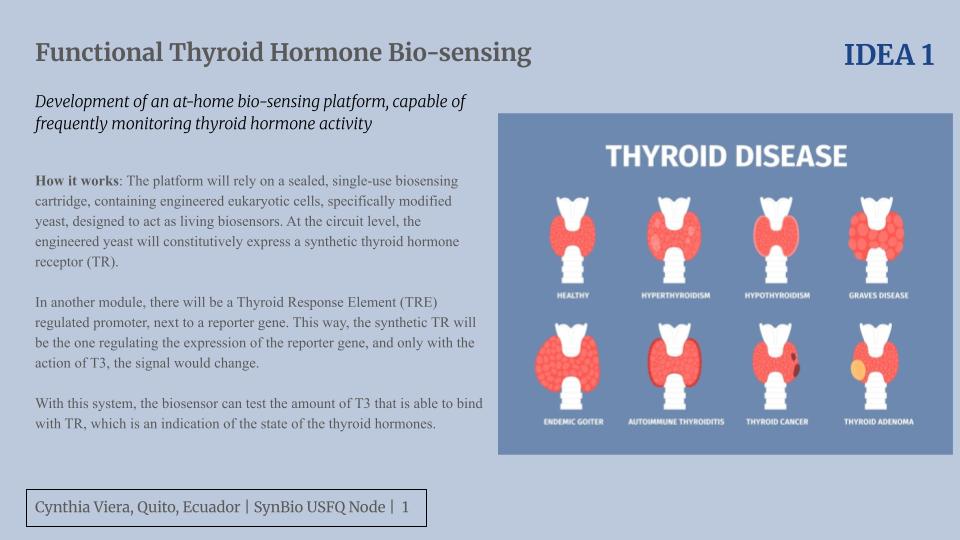
For this assignment, we were expected to create an artwork using OpenTrons. I decided to draw an artistic eye with my name on the bottom. The desgin was created through the GUI page and was edited appropriately to run on Google Colab.
The final design, after passing through the simulator, looks like this!
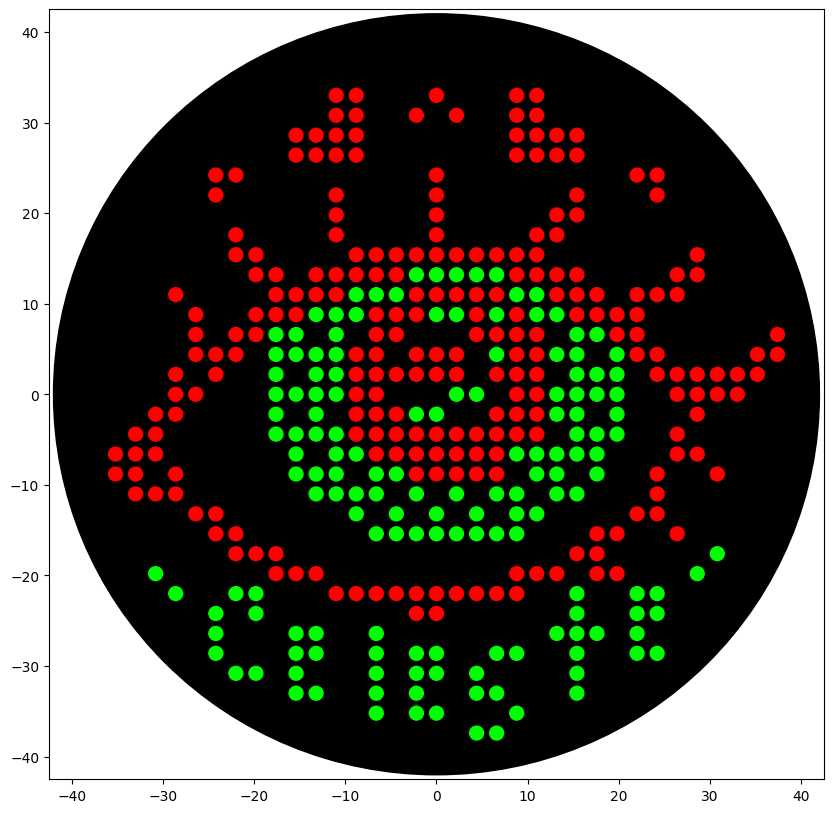

You can find the Colab and the relevant script here, or here: https://colab.research.google.com/drive/1sO_X5QGUP9i2MGskw67G1CD6hE0IIU-E#scrollTo=pczDLwsq64mk&line=1&uniqifier=1
Post-Lab Questions
Question 1
Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is responsible for the most recent pandemic that agitated the world. Its highly contagious nature created the necessity for frequent and efficient diagnostic testing. The standard test used nasopharyngeal swabs processed by RT-qPCR. However, this method relies on specialized equipment and trained medical personnel, resulting in relatively high costs. To address this problem, the paper proposes a strategy using a saliva-based test combined with automation to reduce costs when testing large-scale groups, such as universities.
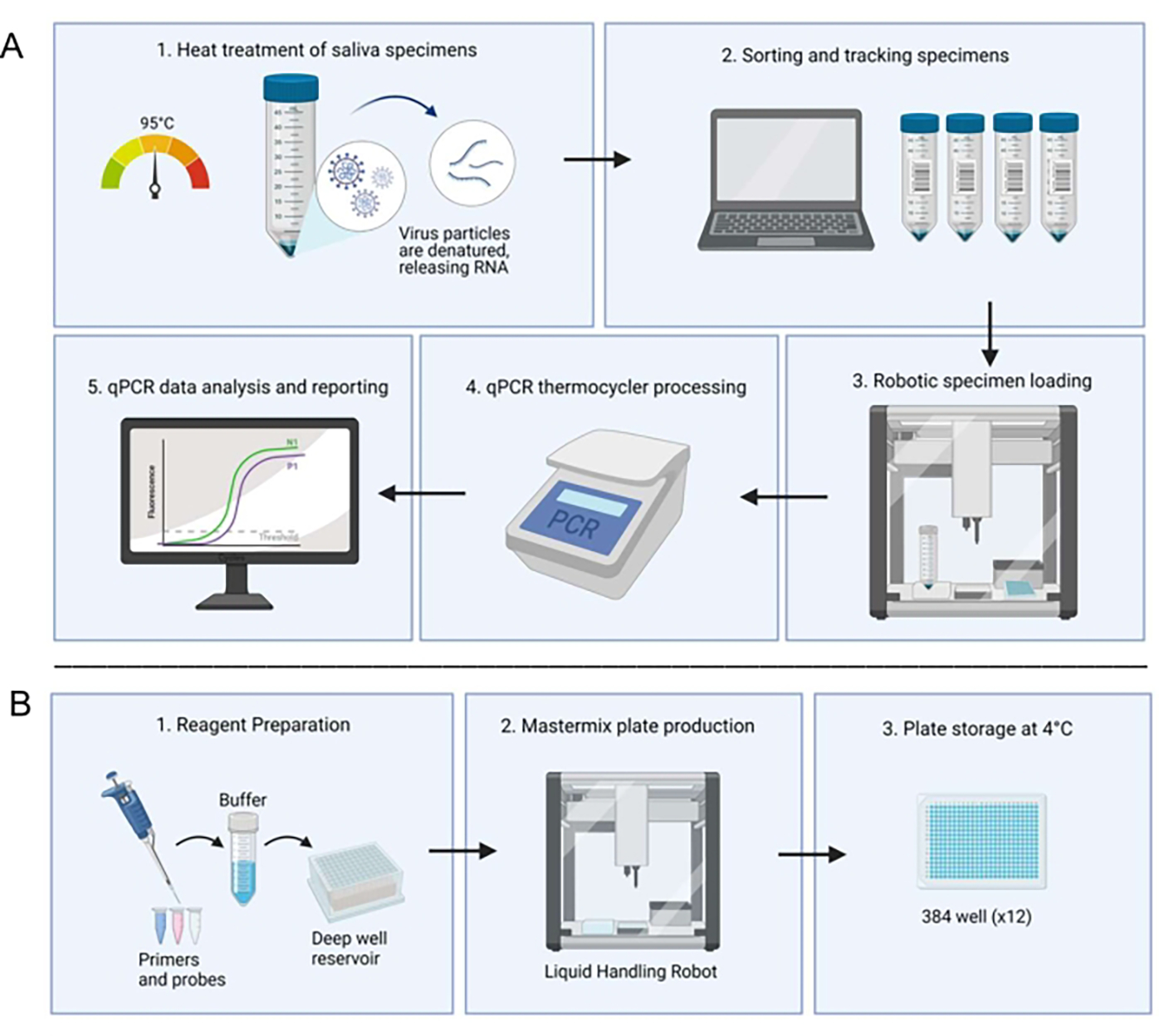
 Figure 1. Laboratory workflow utilizing the saliva-based RT-qPCR diagnostic system.
Figure 1. Laboratory workflow utilizing the saliva-based RT-qPCR diagnostic system.
Before starting the procedure, the Opentrons liquid-handling robot is used to prepare a master mix and distribute it into the wells. The protocol (Figure 1) begins with the collection of saliva samples, which can be performed by each individual without the need for specialized personnel. The recollected samples are then heat-treated at 95°C for 30 minutes. Next, the liquid-handling robot loads the samples into duplicate wells, which already have the master mix, and then they are processed in a qPCR thermocycler. The results are generated by an automated computer system and verified by a technician. If any result is unclear, the sample is rerun. It is also important to note that the technician manually loads the control samples. Through this automation process, the overall processing time is reduced.
A significant result from this paper is that the new strategy achieved 90% sensitivity and 98.9% specificity. Additionally, results from three participants did not initially agree between saliva and nasopharyngeal samples. The saliva tests were positive, and when these individuals were retested two days later, the results remained positive, which may suggest earlier detection using this method. In conclusion, the variation between automated and manual saliva sample processing was not statistically significant. Therefore, the use of automation, which reduces processing time compared to clinical laboratories, is very useful for large-scale surveillance and research.
Question 2
Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more. You may reference this week’s recitation slide deck for lab automation details.
Concise Automation-Focused Description For my final project, I want to develop an optimized, yeast-based functional thyroid hormone (T3) biosensor. I will integrate automation tools throughout the workflow to enable reproducibility, scalability, and systematic circuit optimization. The 4-step plan is as follows:
Step 1: Automated DNA Assembly
A small combinatorial library of genetic circuits will be designed, incorporating variants of Thyroid Response Elements (TREs), diverse promoter strengths, and alternative reporter genes such as GFP and luciferase. An automated liquid handler, Opentrons specifically, will be employed to execute automated plasmid assembly reactions, prepare transformation mixes, and transformed yeast in a 96-well format.
Step 2: Automated Strain Screening
Following transformation, the robot will inoculate yeast colonies into 96-well plates, standardize culture volumes, and normalize cell density through optical density (OD) adjustments. Experimental replicates will then be prepared.
Step 3: Automated Hormone Dose–Response Calibration
To ensure biosensor sensitivity, we will use the liquid handler to generate serial dilutions of T3. After which, precise hormone concentrations will be dispensed across the wells, and synchronized treatments will be initiated to ensure the generation of reproducible dose-response curves.
Step 4: High-Throughput Readout
We can do fluorescence measurements using a plate reader. If available, we can integrate with a cloud laboratory platform (e.g., Ginkgo Nebula) to centralize our data, perform remote execution of standardized protocols, or quickly iterate design-build-test-learn cycles.
Final Project Ideas
References
Ham, R. E., Smothers, A. R., King, K. L., Napolitano, J. M., Swann, T. J., Pekarek, L. G., Blenner, M. A., & Dean, D. (2022). Efficient SARS-CoV-2 Quantitative Reverse Transcriptase PCR Saliva Diagnostic Strategy utilizing Open-Source Pipetting Robots. Journal of visualized experiments : JoVE, (180), 10.3791/63395. https://doi.org/10.3791/63395