Week 04 – Protein Analysis
Protein Analysis and Molecular Foundations
I used Gemini AI to answer questions 1-11. The prompt was the question itself.
1. Amino Acids in 500g of Meat
To find the total number of molecules, we estimate the protein content of meat (roughly 25%) and use Avogadro’s number.
- Protein Mass: 500g multiplied by 0.25 equals 125g of protein.
- Moles of Amino Acids: Using an average mass of 100 Da (100 g/mol), we calculate 125g / 100 g/mol = 1.25 moles.
- Total Molecules: 1.25 x 6.022 x 1023 = **7.5 x 1023 molecules**.
2. Why Humans Eat Beef but Don’t Become Cows
When you consume meat, your digestive enzymes break down the animal proteins into individual amino acids. These building blocks are then transported to your cells, where your own ribosomes reassemble them into human proteins based on the specific instructions in your DNA.
3. Why There Are Only 20 Natural Amino Acids
While hundreds of amino acids exist chemically, life settled on a standard set of 20 because they provide a “functional toolkit” diverse enough (acidic, basic, polar, and hydrophobic) to build almost any protein structure. This set likely became “frozen” early in evolution—changing the code later would have caused lethal mutations across all existing proteins.
4. Non-Natural Amino Acids (ncAAs)
Scientists can create and incorporate non-natural amino acids using expanded genetic codes.
- Design Concept: An amino acid with a cyano-group (C≡N) side chain. This can be used as a sensitive infrared probe to measure local electric fields or pH changes within a protein’s active site.
5. Pre-biotic Origins
Before enzymes existed, amino acids formed via abiotic synthesis.
- Chemical Evolution: The Miller-Urey experiment showed that lightning-like sparks in a reducing atmosphere (containing methane, ammonia, hydrogen, and water) could create glycine and alanine.
- Astrobiology: Amino acids have been found on meteorites, suggesting these building blocks can form in space before life begins.
6. Handedness of D-amino Acid Helices
Natural L-amino acids form right-handed alpha-helices. If you utilize D-amino acids (the mirror image), the steric constraints are reversed, resulting in a left-handed helix.
7. Discovering Additional Helices
Beyond the standard alpha-helix, proteins occasionally use:
- 3-10 helix: A tighter, more elongated helix.
- Pi-helix: A wider, shorter helix often found near functional active sites.
- Polyproline II (PP-II) helix: A left-handed, extended helix common in collagen.
8. Why Most Helices are Right-Handed
This is determined by the chirality of L-amino acids. In a right-handed helix, the side chains point outward, minimizing steric clashes. In a left-handed helix made of L-amino acids, the side chains would crash into the protein backbone, making the structure energetically unstable.
9. Why Beta-sheets Tend to Aggregate
Beta-sheets have “sticky edges”—exposed hydrogen-bond donors and acceptors that are not “satisfied” or covered. If these edges aren’t capped by another part of the protein, they will seek out other beta-sheets to bond with, leading to uncontrolled stacking.
10. Driving Forces for Beta-sheet Aggregation
- Hydrogen Bonding: Inter-strand hydrogen bonds act like molecular “Velcro.”
- Hydrophobic Effect: Many beta-sheets have hydrophobic faces. To escape the water, these faces stack together, burying the non-polar side chains in a dry core.
11. Amyloids: Disease and Materials
- Amyloid Diseases: In diseases like Alzheimer’s or Parkinson’s, proteins misfold into highly stable, insoluble “cross-beta” structures that the body cannot easily clear.
- Materials Science: Because amyloid beta-sheets are incredibly strong and stable, they are being used to design nanofibers, conductive wires, and biocompatible scaffolds for tissue engineering.
Protein Analysis: SOSTDC1 (NP_056167)
GDP-fucose protein O-fucosyltransferase 1 & Tooth Regeneration
I selected the protein SOSTDC1. This protein is known to block the growth of a second set of adult teeth. Currently, there are clinical trials in Japan where researchers are blocking this protein using antibodies to stimulate tooth regrowth. I am investigating if there are other structural ways to manipulate it.
1. Sequence and Composition
Amino Acid Sequence:
MEKLAPTHWPPEKRVAYCFEVAAQRSPDKKTCPMKEGNPFGPFWDQFHVSFNKSELFTGISFSASYREQWSQRFSPKEHPVLALPGAPAQFPVLEEHRPLQKYMVWSDEMVKTGEAQIHAHLVRPYVGIHLRIGSDWKNACAMLKDGTAGSHFMASPQCVGYSRSTAAPLTMTMCLPDLKEIQRAVKLWVRSLDAQSVYVATDSESYVPELQQLFKGKVKVVSLKPEVAQVDLYILGQADHFIGNCVSSFTAFVKRERDLQGRPSSFFGMDRPPKLRDEF
- Length: 280 amino acids.
- Most Frequent Amino Acid: Alanine (A) is the most frequent. (Note: The previous count of 292 was likely a typo as it exceeded the total protein length).
2. Family and Domain Classification
SOSTDC1 belongs to several recognized protein families and domains:
| Database | Classification / ID | Description |
|---|---|---|
| FunFam | 2.10.90.10:FF:000019 | Sclerostin domain-containing protein 1 |
| Gene3D | 2.10.90.10 | Cystine-knot cytokines |
| InterPro | IPR008835 | Sclerostin/SOSTDC1 |
| PANTHER | PTHR14903 | Sclerostin-related |
| Pfam | PF05463 | Sclerostin |
3. Structural Analysis (RCSB PDB)
The structure was solved in 2017 with a high-quality resolution of 2.09 Å.

- Molecules: Aside from the protein, the structure contains water molecules.
- Classification: It belongs to the Transferase structural family.
- Secondary Structure: The protein contains a significant number of helices.
- Surface Topology: Visualization of the protein surface reveals “holes” or pockets that could potentially be targeted by small molecules.

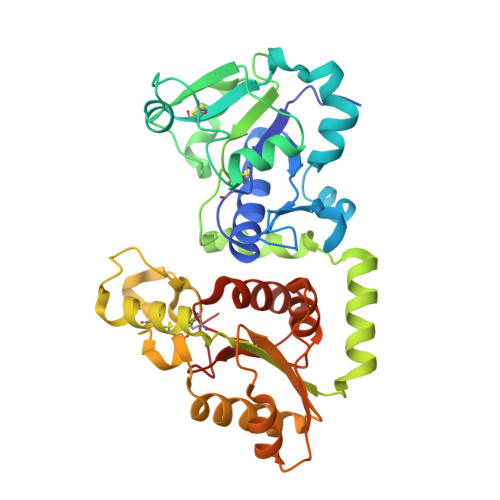
4. 3D Visualization
Below is the 3D representation showing the complex folding and side-chain distributions.
Observations:
- Hydrophobic vs. Hydrophilic: Hydrophobic residues tend to be buried within the core to stabilize the fold, while hydrophilic residues are primarily distributed on the surface to interact with the aqueous environment.
- Targeting: The presence of surface “holes” suggests possibilities for small-molecule inhibitors as an alternative to antibody-based therapies.
Group Project Re-Skinning MS2 for Cancer Targeting
While my current research focuses on the L protein for bacterial lysis, the same protein engineering principles can be used to “re-skin” the MS2 Coat Protein (CP) to target cancer cells. By modifying the FG loop of the coat protein, we can turn the phage into a targeted delivery vehicle.
A. Targeting Peptide: The RGD Motif
To target cancer cells (specifically those overexpressing integrins, like breast or lung cancer), we can “plug” the following peptide sequence into the MS2 coat protein:
Targeting Sequence: Arg-Gly-Asp (RGD)
B. Computational Engineering Workflow
To re-engineer the phage surface, I would follow these steps:
- Scaffold Modeling: Utilize the MS2 coat protein structure (PDB: 2MS2) as the base scaffold.
- Peptide Grafting: Insert the RGD motif into the solvent-exposed FG loop (between residues 70-80).
- Linker Optimization: Add flexible linkers (e.g.,
Gly-Gly-Gly-Ser) to ensure the peptide can reach and bind the tumor receptors. - Structural Validation: Use tools like the Nuclera system (Stage 4) to ensure the mutations do not prevent the 180 coat proteins from assembling into a stable icosahedral shell.
C. Comparison of Engineering Goals
| Feature | Phage Therapy (Bacteria) | Phage Nanomedicine (Cancer) |
|---|---|---|
| Primary Target | E. coli F-pilus | Cancer markers (Integrins, HER2) |
| Protein Focus | Lysis Protein (L) | Coat Protein (CP) |
| Desired Outcome | Bacterial Lysis | Cell-specific drug delivery |
| Key Challenge | DnaJ-independence | Evading human immune clearance |
D. The “Trojan Horse” Strategy
By re-skinning the outside to find the cancer and replacing the internal viral RNA with a suicide gene or chemotherapeutic cargo, the MS2 phage acts as a programmable nanobot. This represents the intersection of the HTGAA phage project and state-of-the-art oncology.