Week 03 HW: Lab automation


Assignment: Python Script for Opentrons Artwork — DUE BY YOUR LAB TIME!
0. Your task this week is to Create a Python file to run on an Opentrons liquid handling robot.
1. Review this week’s recitation and this week’s lab for details on the Opentrons and programming it.
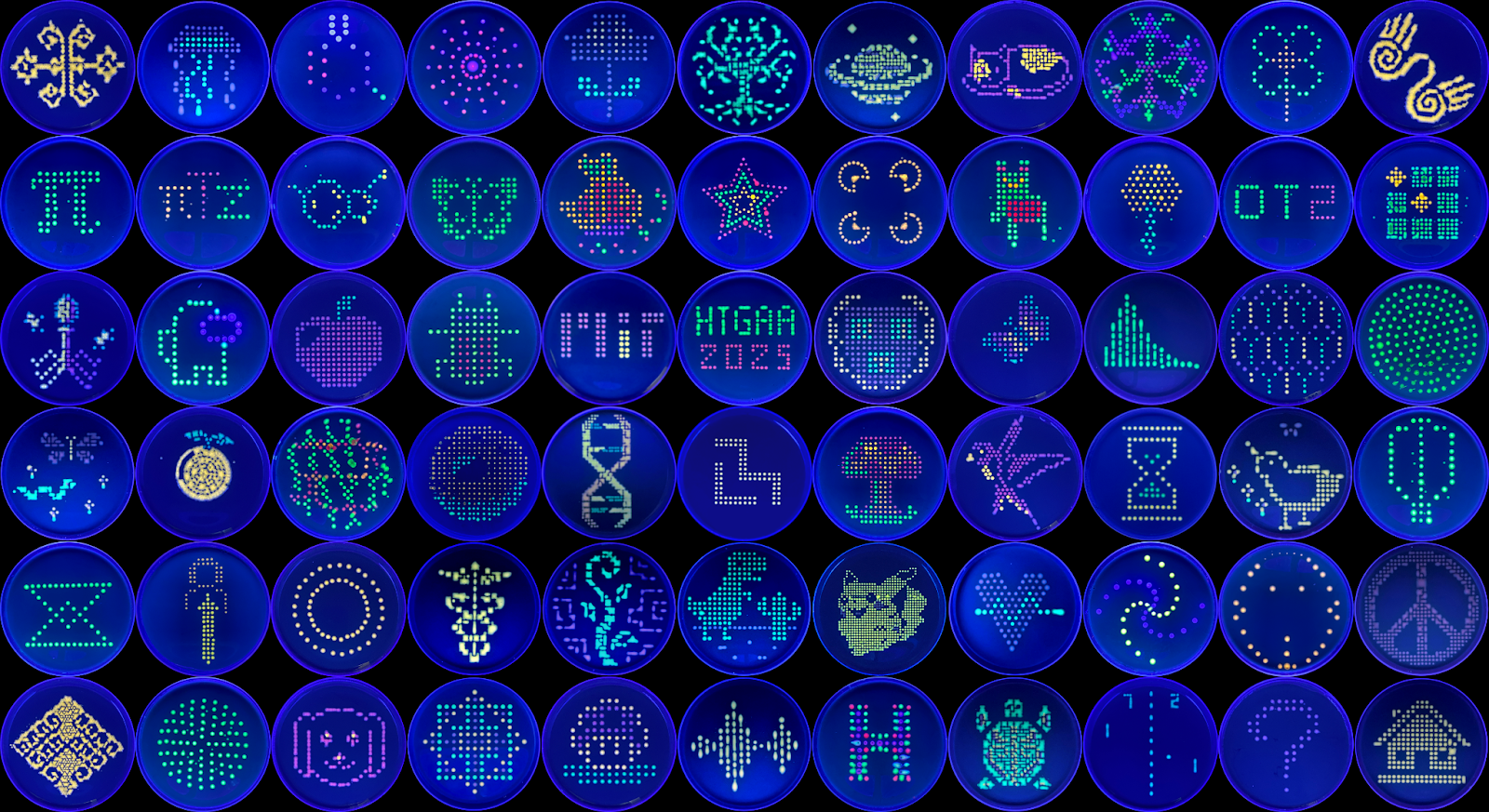
2. Generate an artistic design using the GUI at opentrons-art.rcdonovan.com.
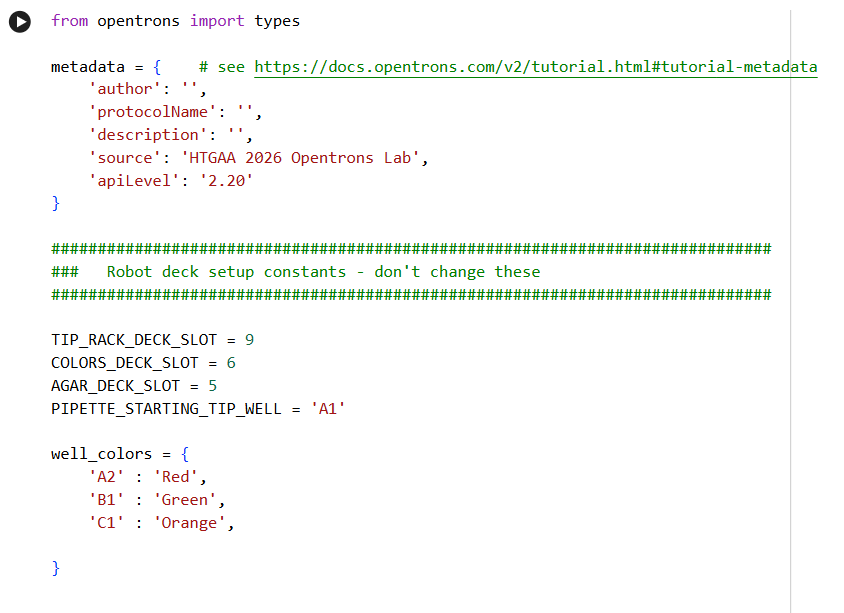
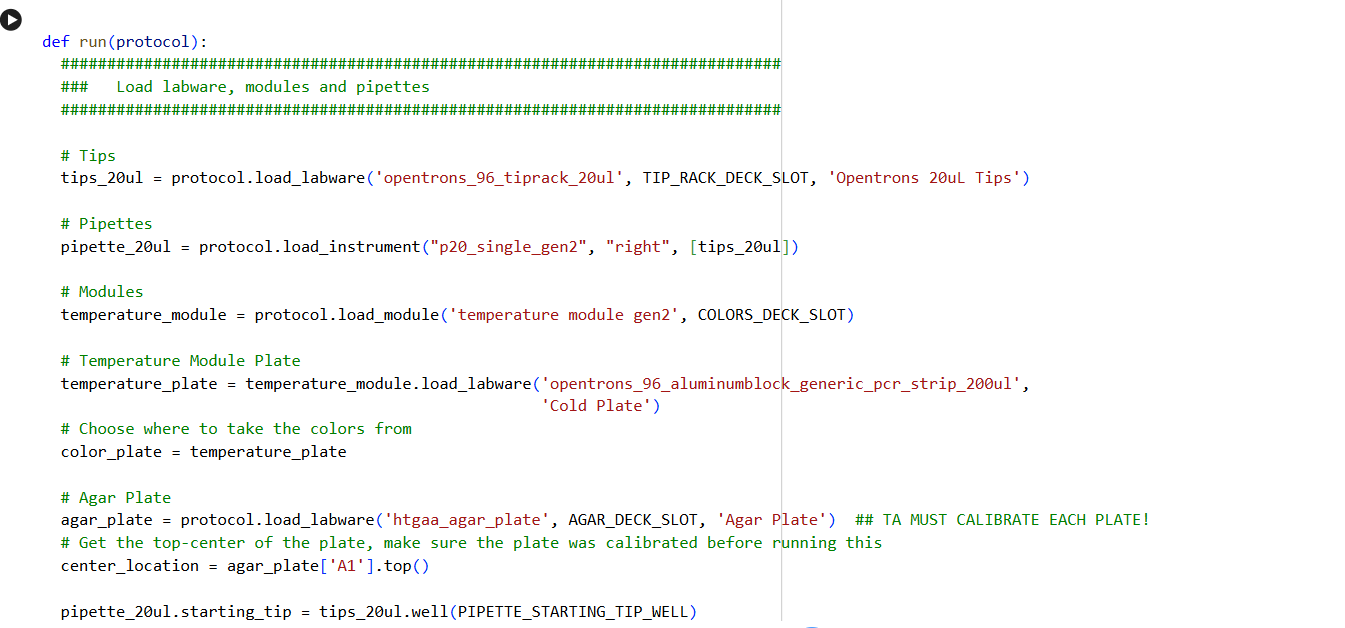
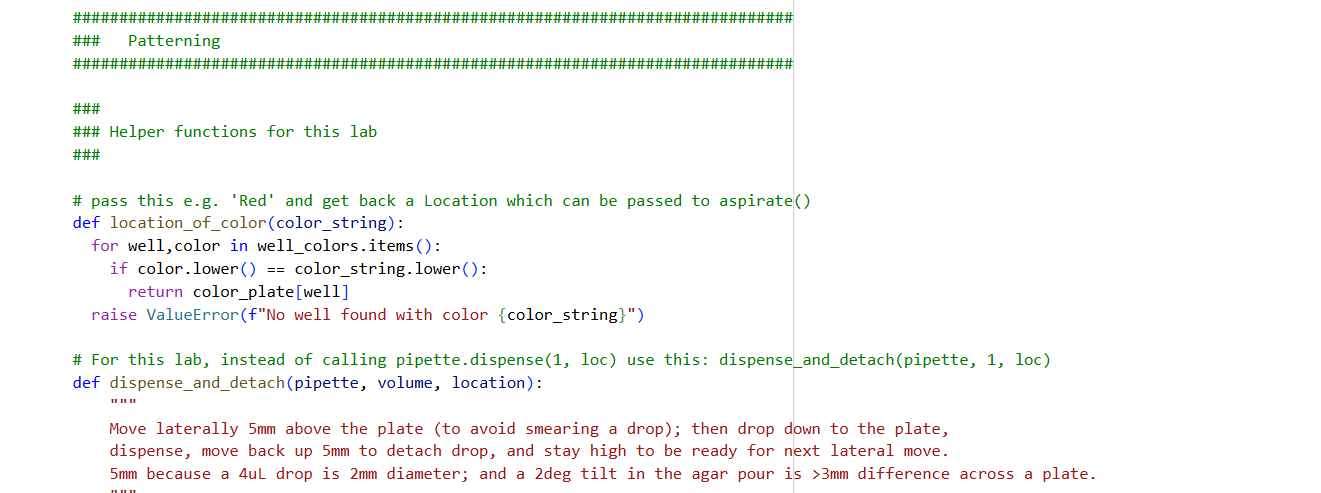
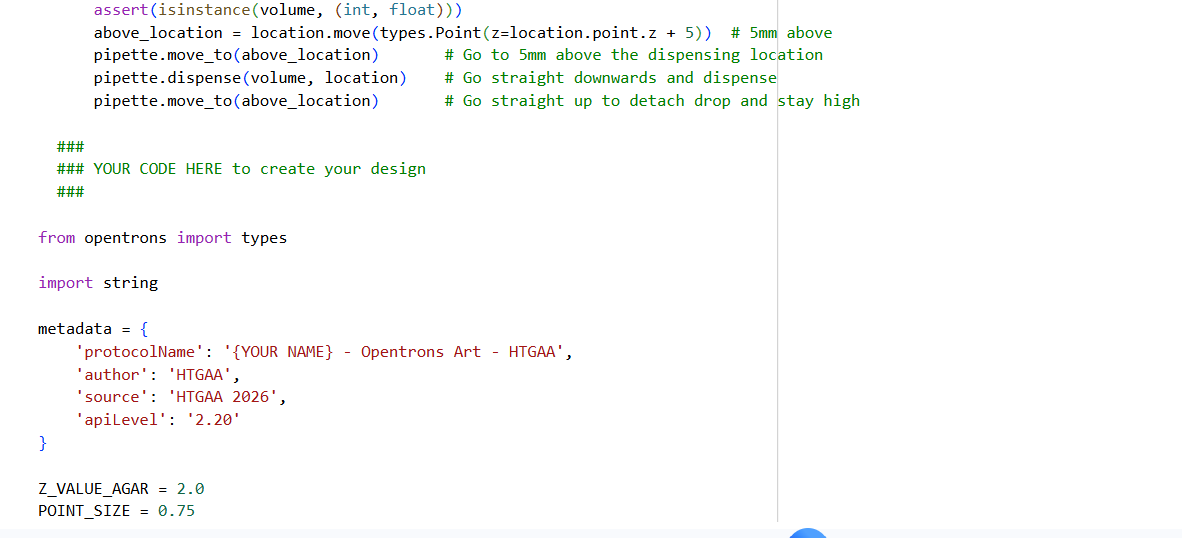
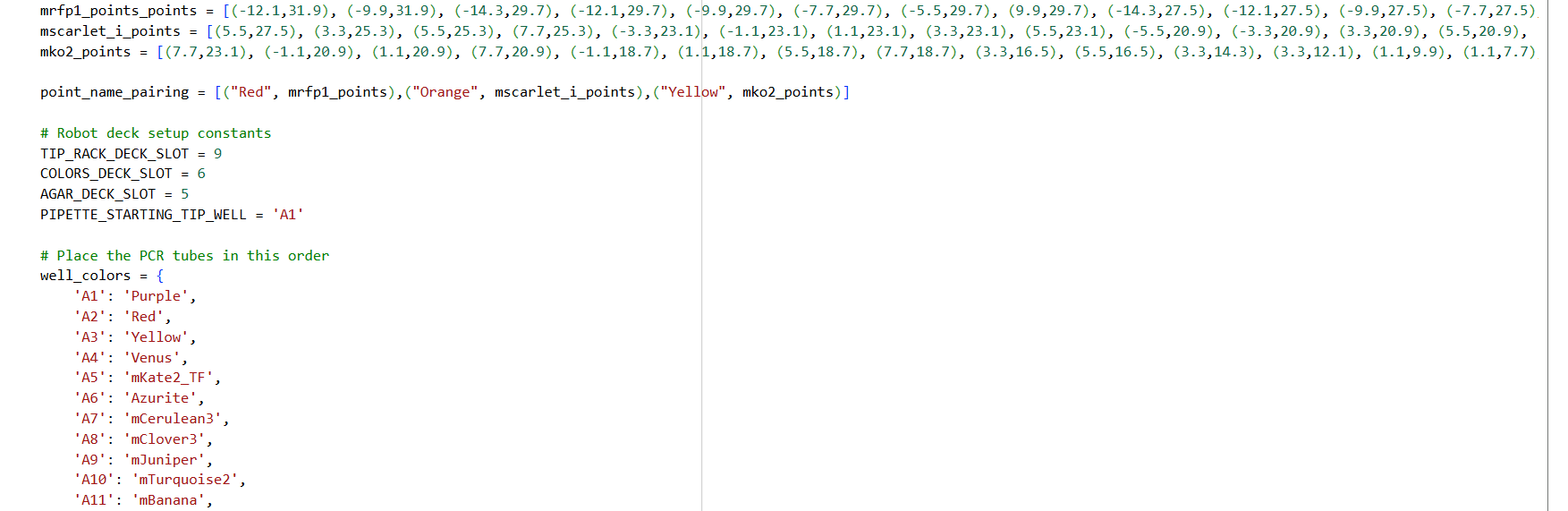
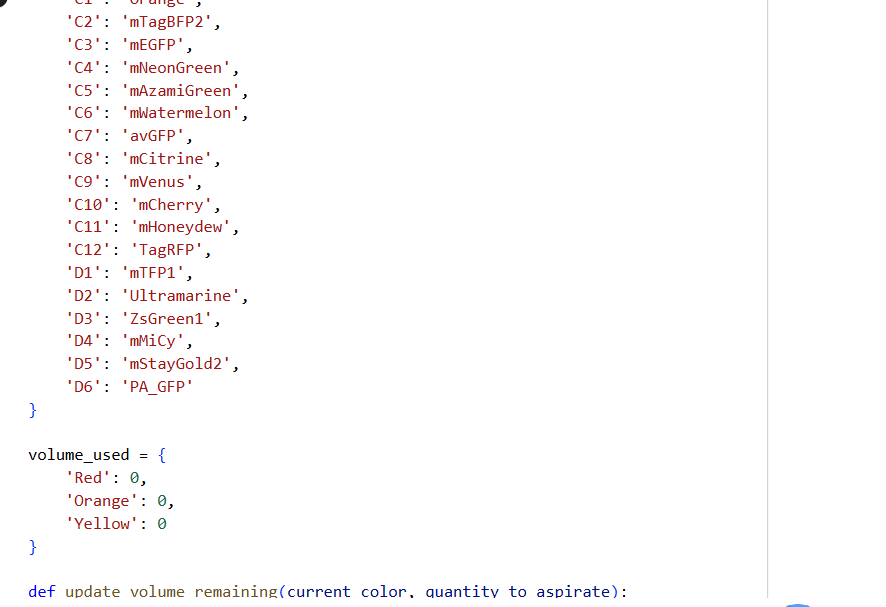
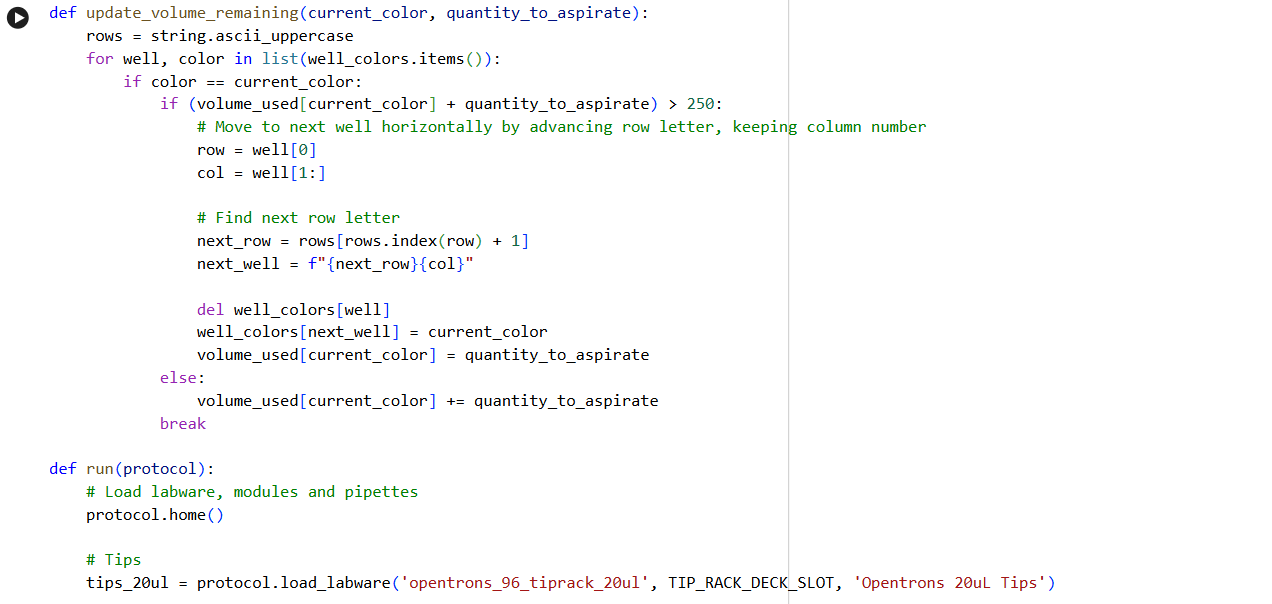
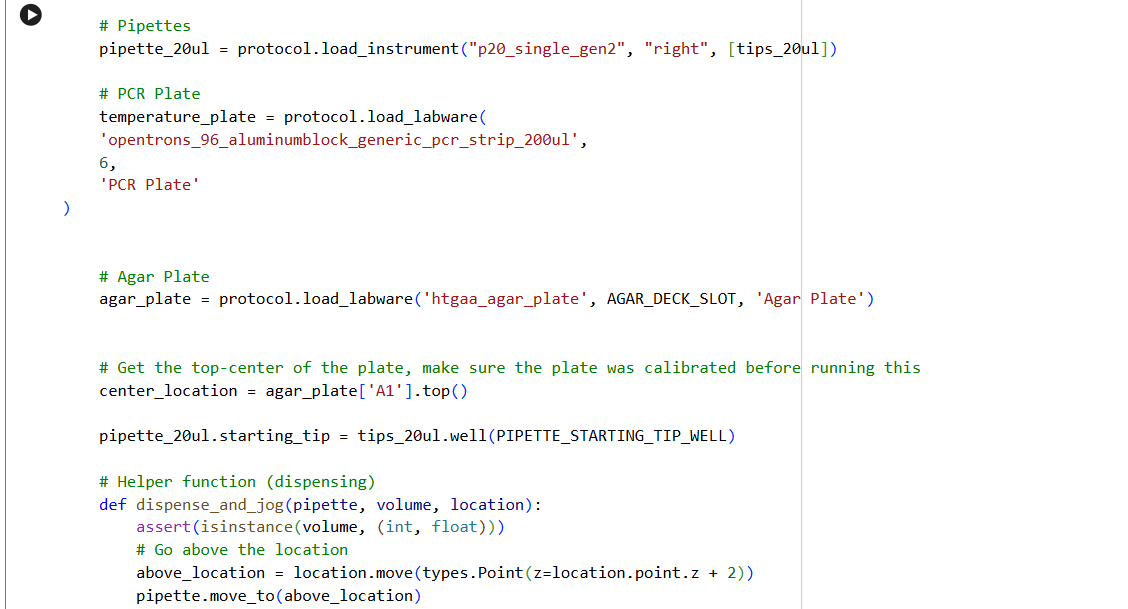
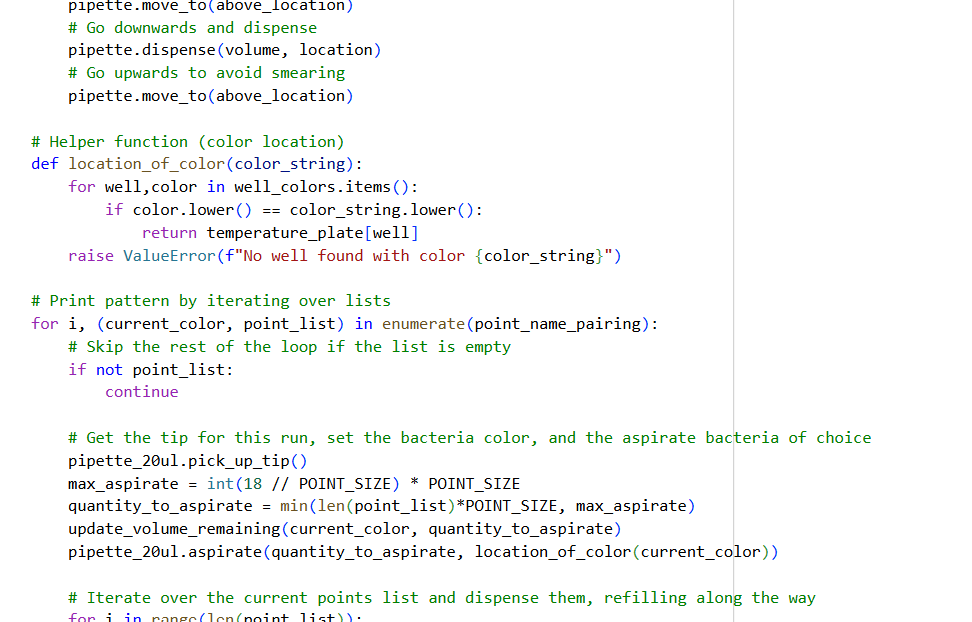
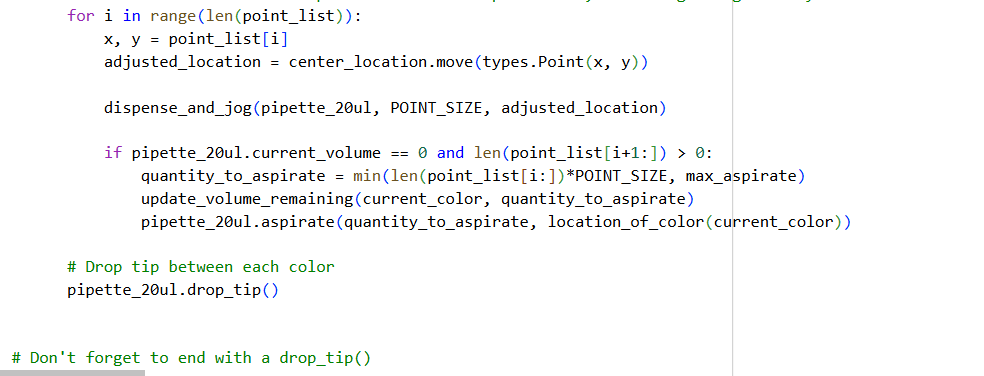
- Using the coordinates from the GUI, follow the instructions in the HTGAA26 Opentrons Colab to write your own Python script which draws your design using the Opentrons.
- You may use AI assistance for this coding — Google Gemini is integrated into Colab (see the stylized star bottom center); it will do a good job writing functional Python, while you probably need to take charge of the art concept.
- If you’re a proficient programmer and you’d rather code something mathematical or algorithmic instead of using your GUI coordinates, you may do that instead.
4. If you use AI to help complete this homework or lab, document how you used AI and which models made contributions.
5. Sign up for a robot time slot if you are at MIT/Harvard/Wellesley or at a Node offering Opentrons automation. The Python script you created will be run on the robot to produce your work of art!
6. Submit your Python file via this form.
✨✨My code✨✨
https://colab.research.google.com/drive/1rOfQVambbO3m8ZcjPQd7lQDcfa-qjr_j?usp=sharing
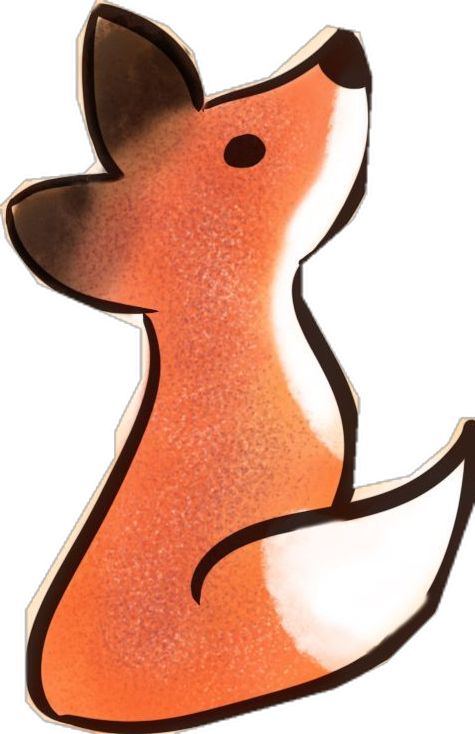
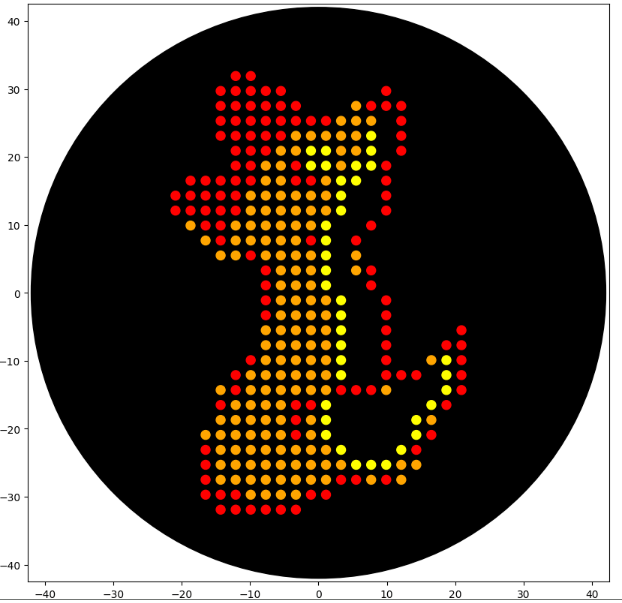
My inspiration was this image of the little fox from the story of the little prince.

Post-Lab Questions — DUE BY START OF FEB 24 LECTURE
One of the great parts about having an automated robot is being able to precisely mix, deposit, and run reactions without much intervention, and design and deploy experiments remotely.
For this week, we’d like for you to do the following:
1. Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
Title: Automation of protein crystallization scaleup via Opentrons 2 liquid handling
Publication: SLAS Technology, 2025
This article is impressive because it explains that protein crystallization is an important process; however, there are several variables that are difficult to control to execute this process, especially on a small scale. When humans develop this process, it might be difficult, exhausting, and the outcomes may be inaccurate.
Consequently, as a new way to resolve this problem, scientists have developed a new approach optimizing protein crystallization trials at the multi-microliter scale with the Opentrons-2 liquid handling robot.
Scientists explain that although there are different robots on the market whose objective is to improve this process, these robots are expensive and their programming is exclusive. On the other hand, we have Opentrons, which is a robot with automation for several purposes, and it can be programmed using Python.
With Python scripts, scientists compare the efficacy and accuracy of the process developed by Opentrons OT-2 vs the manual method.
The materials they used included:
- Opentrons OT-2
- Crystallization plates (sitting drop 24-well): for forming the protein crystals.
- Protein solutions: the proteins they wanted to crystallize.
- Precipitating and buffer solutions: substances that help the crystals form.
Steps
- Plate Preparation
- The wells of the crystallization plate were placed in specific positions on the robot deck
- Robot Programming
- Python was used to instruct the robot on how to move liquids
- Pick up protein from one container.
- Pick up buffer/precipitate from another container.
- Mix small drops into the wells of the plate.
- Running Assays
- The robot performed all pipetting automatically, drop by drop.
Some plates contained different combinations of protein and precipitate to test various conditions simultaneously.
They illustrate the process with the following images:
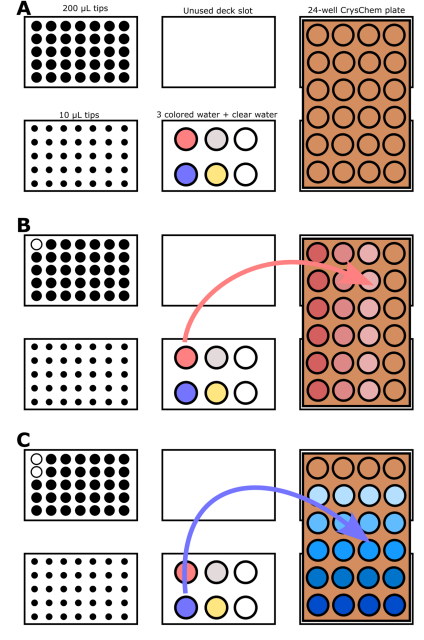
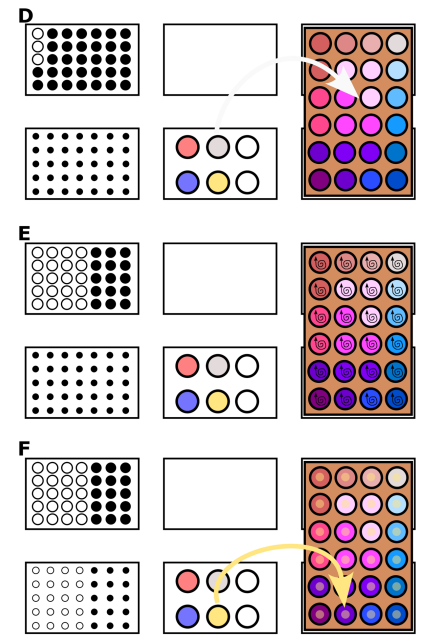
2. Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more. You may reference this week’s recitation slide deck for lab automation details. While your description/project idea doesn’t need to be set in stone, we would like to see core details of what you would automate. This is due at the start of lecture and does not need to be tested on the Opentrons yet.
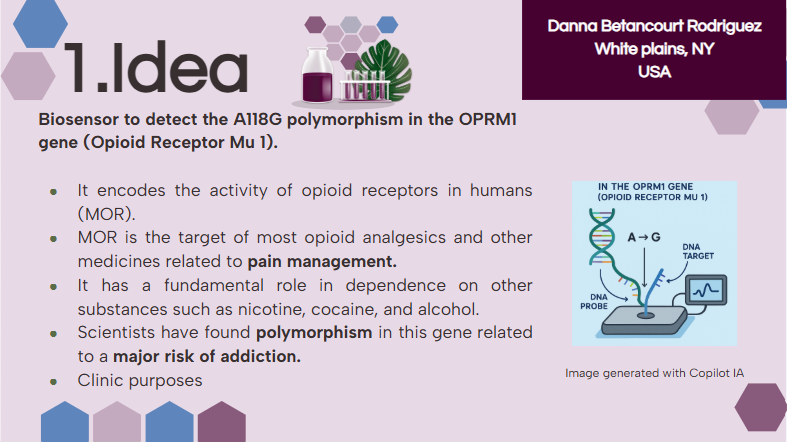
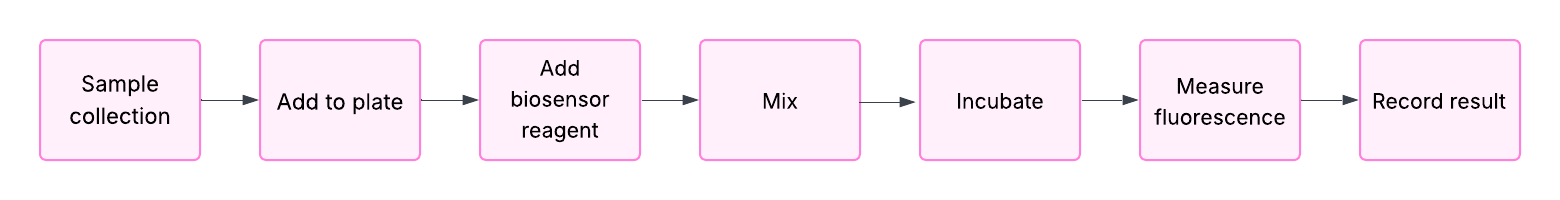
For my project, I want to develop a biosensor. I plan to automate sample preparation and measurement using a liquid-handling robot (like Opentrons) and a plate reader (like PHERAstar).
This automation will be used only during laboratory development to ensure reproducible and accurate results. However, this automation will not be included in the final product, because the main goal of my biosensor is that it will be used in ambulatory settings.

Final Project Ideas — DUE BY START OF FEB 24 LECTURE
As explained in this week’s recitation, add 1-3 slides in your Node’s section of this slide deck with 3 ideas you have for an Individual Final Project. Be sure to put your name, city, and country on your slide!
References
- DeRoo, Jacob B., et al. “Automation of Protein Crystallization Scaleup via Opentrons-2 Liquid Handling.” SLAS Technology, vol. 32, June 2025, p. 100268, pmc.ncbi.nlm.nih.gov/articles/PMC12229254/, https://doi.org/10.1016/j.slast.2025.100268. Accessed 12 Dec. 2025.
- Taqi, M. M., Faisal, M., & Zaman, H. (2019). OPRM1 A118G polymorphisms and its role in opioid addiction: Implication on severity and treatment approaches. Pharmacogenomics and Personalized Medicine, Volume 12, 361–368. https://doi.org/10.2147/pgpm.s198654
- Tasoulis, T., & Isbister, G. (2017). A review and database of snake venom proteomes. Toxins, 9(9), 290. https://doi.org/10.3390/toxins9090290
- Alonso, L. L., Slagboom, J., Casewell, N. R., Samanipour, S., & Kool, J. (2025). Categorization and Characterization of Snake Venom Variability through Intact Toxin Analysis by Mass Spectrometry. Journal of Proteome Research, 24(3), 1329–1341. https://doi.org/10.1021/acs.jproteome.4c00923