Week 06 HW: Genetic circuits part I: Assembly Technologies


DNA Assembly
Answer these questions about the protocol in this week’s lab:
1. What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
The Phusion High-Fidelity PCR Master Mix is a mix that has several components that allow the user to add only the DNA template, primers, and water to perform PCR.
Advantages:
- Robust Reactions - Maximal success with minimal optimization.
- Extreme Fidelity - > 50X greater than Taq
- High Speed - Dramatically reduced extension times (10X faster than Pfu)
- High Yield - Increased product yield using minimal amount of enzyme.
- Versatile - Can be used for routine per as well as long or difficult templates.


2. What are some factors that determine primer annealing temperature during PCR?
PCR has three important steps:
1. Denaturation (95 °C) → the DNA separates into two strands.
2. Annealing (50–65 °C) → the primers attach to the DNA.
3. Extension (~72 °C) → the polymerase copies the DNA.
The annealing temperature is the temperature adequate for the primer to bind to the DNA.
If the temperature is not adequate, the primer might bind in the wrong place or not bind to the DNA.
Factors that determine the annealing temperature
1. Melting temperature (Tm)
The Tm is the temperature at which half of the primers are attached to the DNA and the other half are not.
If the Tm is high, you need a higher annealing temperature.
2. Content of GC
The DNA bases are bound by hydrogen bonds
A–T → 2 bonds (weak)
G–C → 3 bonds (strong)
Primers with more GC will have a Tm higher
Primers with fewer GC will have a Tm lower
3. Primer length
Long primer → Stronger bond → Higher Tm
Short primer → weakest bond → lowest Tm
4. Salt Concentration (Na⁺ or Mg²⁺)
Salts stabilize the interaction between DNA and primer.
More salt → the negative charges of the DNA are neutralized → the primer binds more easily.
More salt → increases Tm
Less salt → decreases Tm
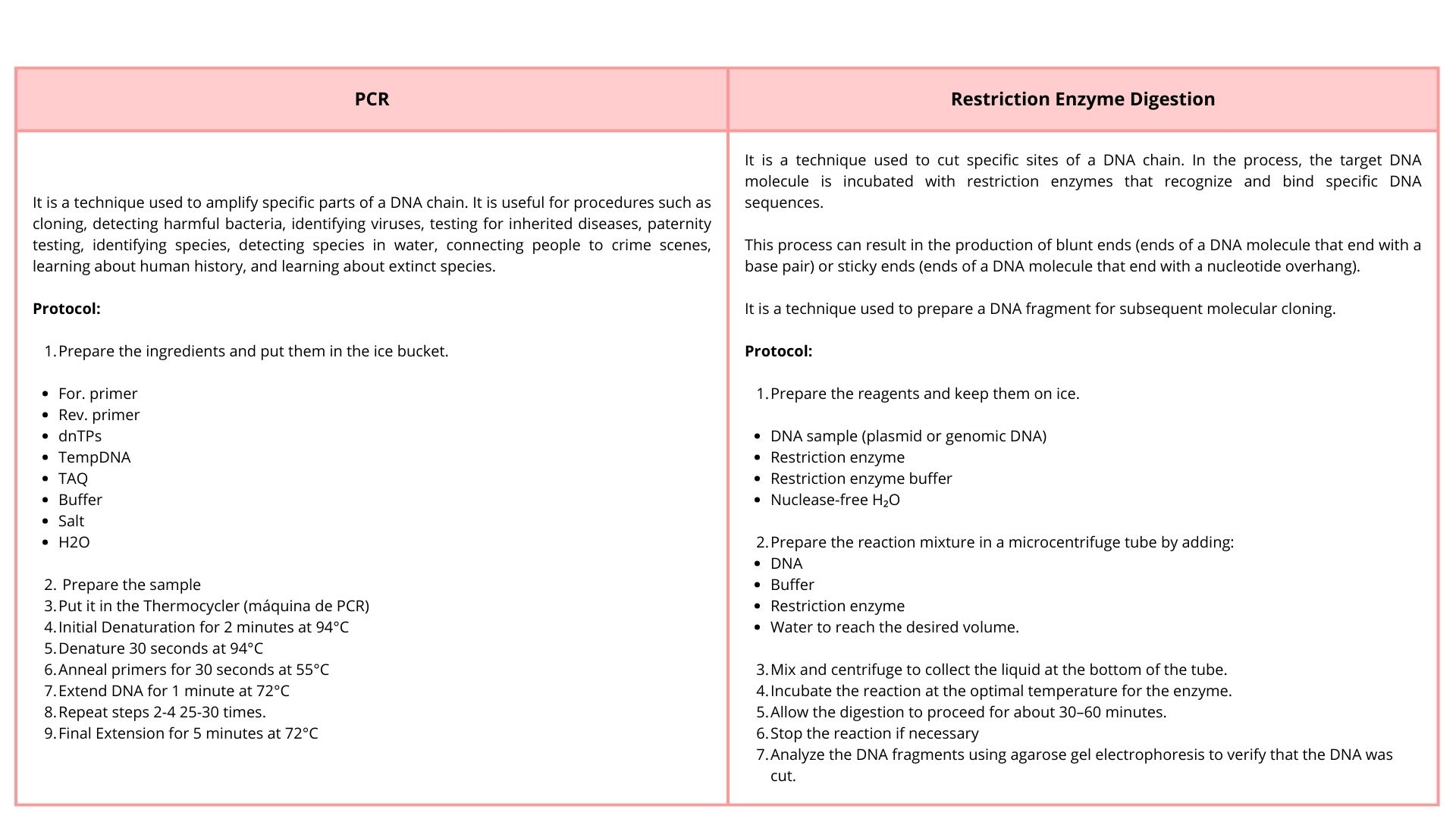
3. There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.

4. How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
Gibson cloning is a technique used to assemble multiple linear DNA fragments, since this technique does not require specific restriction sites and leaves no scar between joined fragments.
Researchers must ensure that the DNA fragments that they want to join are compatible, because adjacent segments should have identical sequences at the ends, approximately 30 bp that match the ends of adjacent fragments
5. How does the plasmid DNA enter the E. coli cells during transformation?
The process is known as bacterial transformation. In this technique, scientists use cells that have been treated with calcium chloride to allow plasmid DNA to attach to the cell membrane. These cells are called competent cells. A brief heat shock then creates temporary pores in the membrane, allowing the plasmid DNA to enter the cell. Another method is electroporation, which uses an electrical field to increase the permeability of the cell membrane, allowing plasmids to enter the cell. After transformation, the bacteria can be cultivated on agar plates.
6. Describe another assembly method in detail (such as Golden Gate Assembly)
- Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
- Model this assembly method with Benchling or Asimov Kernel!
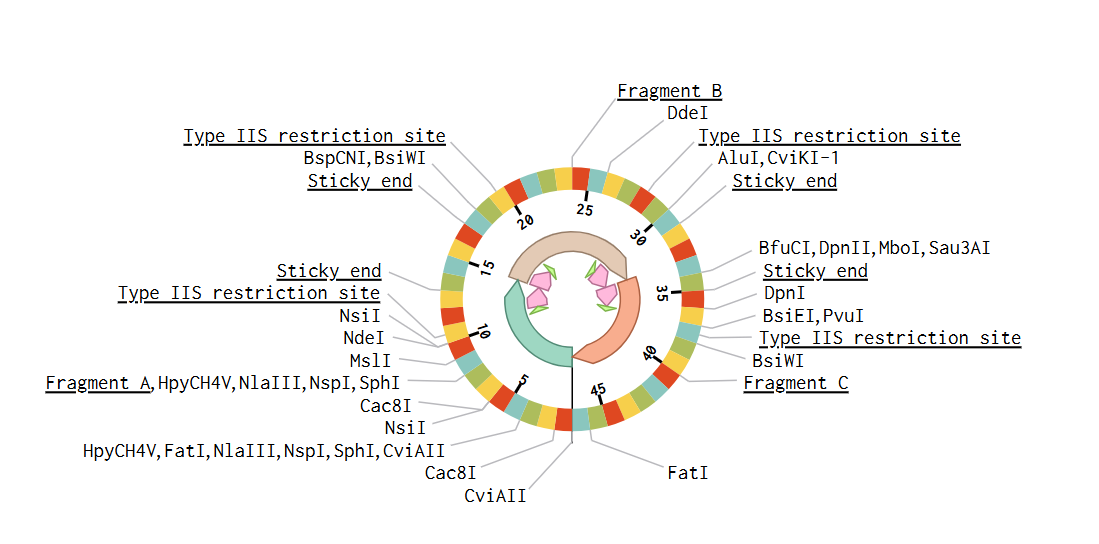
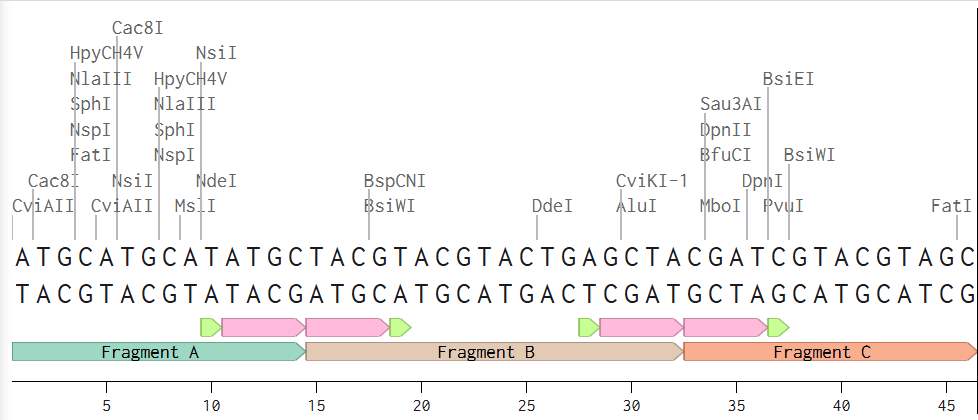
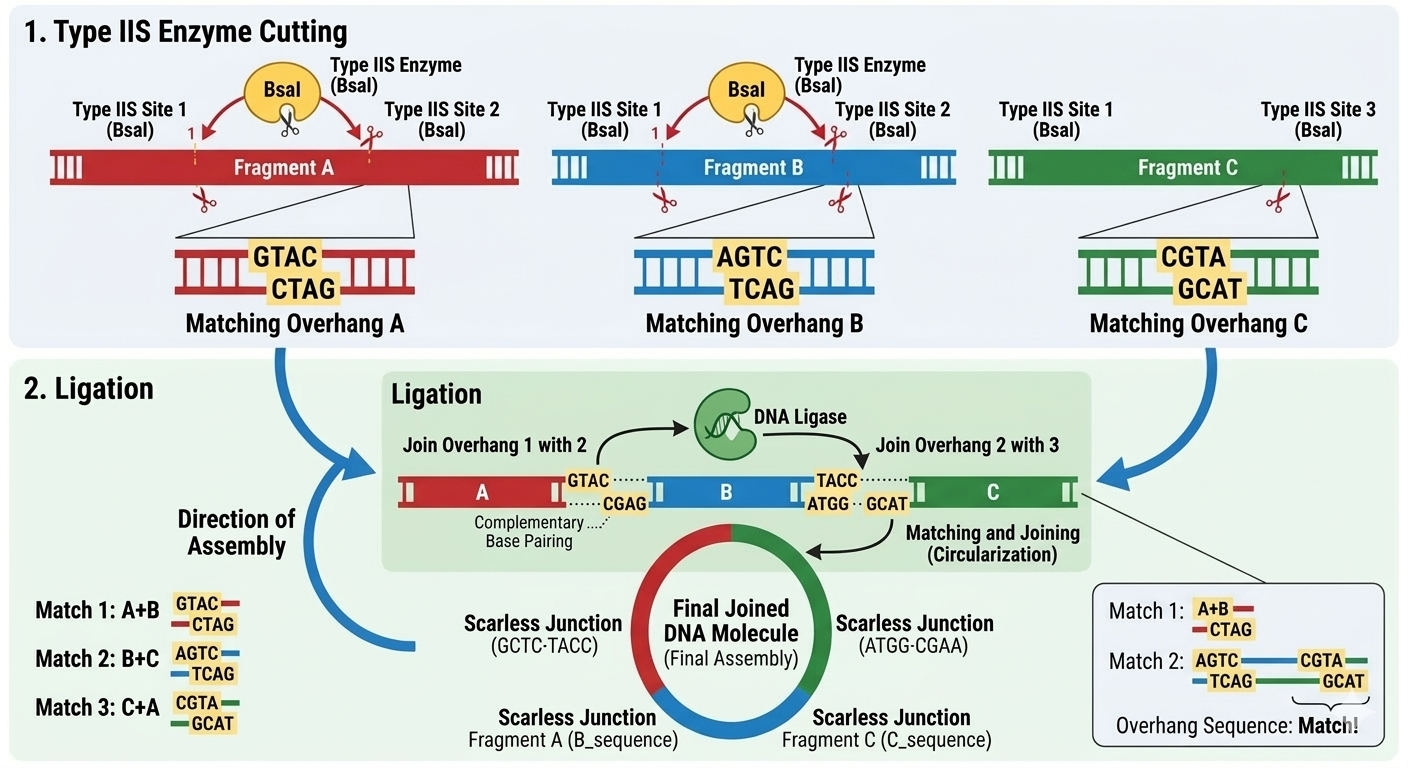
Golden Gate Assembly is a molecular cloning method that allows the assembly of multiple DNA fragments in a single reaction. This method uses Type IIS restriction enzyme digestion, which cuts DNA outside of its recognition sequence, generating specific overhangs. These overhangs are designed with the objective that adjacent DNA fragments have complementary ends that can join in the correct order. DNA ligase joins the fragments to generate a single DNA molecule.
The principal advantage of this method is that the correct order of DNA fragments is ensured by designing specific complementary overhangs that only allow adjacent fragments to ligate together.

My assembly:
Assignment: Asimov Kernel
1. Create a Repository for your work
2. Create a blank Notebook entry to document the homework and save it to that Repository
3. Explore the devices in the Bacterial Demos Repo to understand how the parts work together by running the Simulator on various examples, following the instructions for the simulator found in the “Info” panel (click the “i” icon on the right to open the Info panel)
4. Create a blank Construct and save it to your Repository
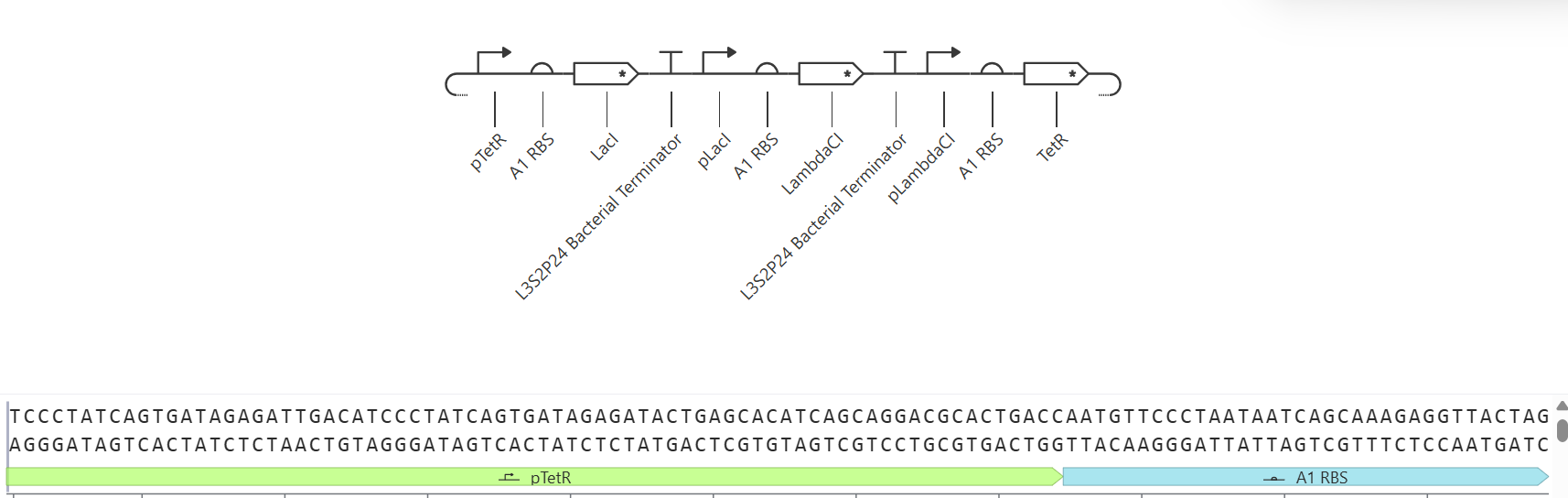
5. Recreate the Repressilator in that empty Construct by using parts from the Characterized Bacterial Parts repository
6. Search the parts using the Search function in the right menu
7. Drag and drop the parts into the Construct
8. Confirm it works as expected by running the Simulator (“play” button) and compare your results with the Repressilator Construct found in the Bacterial Demos repository
10. Document all of this work in your Notebook entry - you can copy the glyph image and the simulator graphs, and paste them into your Notebook
11. Build three of your own Constructs using the parts in the Characterized Bacterials Parts Repo
12. Explain in the Notebook Entry how you think each of the Constructs should function
13. Run the simulator and share your results in the Notebook Entry. If the results don’t match your expectations, speculate on why and see if you can adjust the simulator settings to get the expected outcome
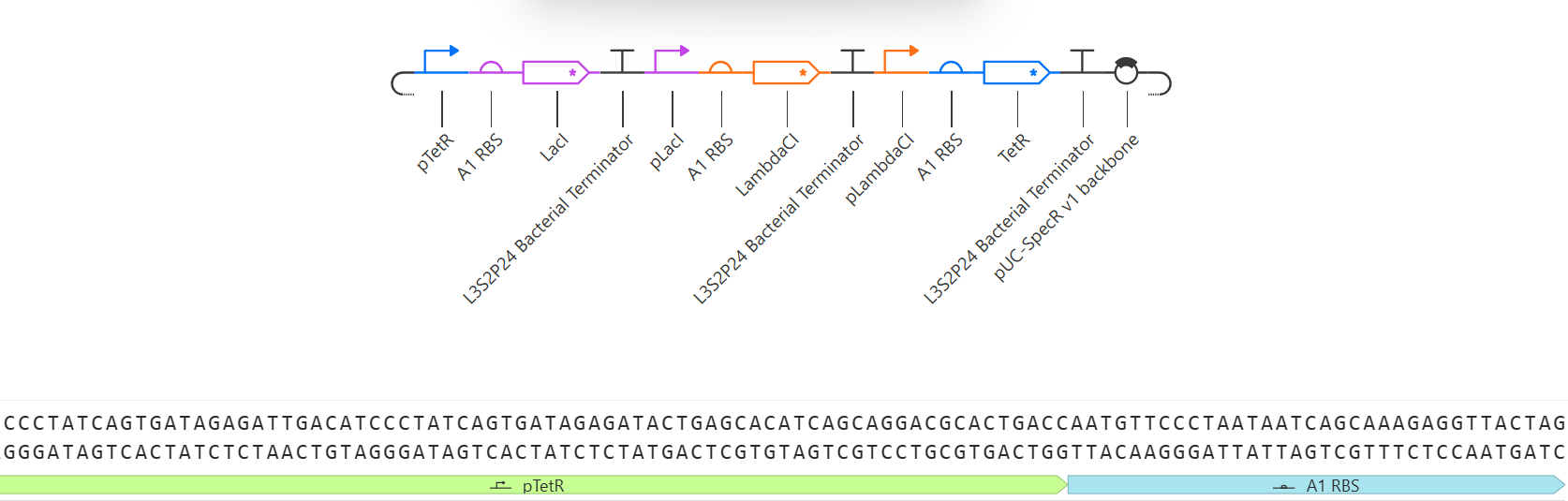
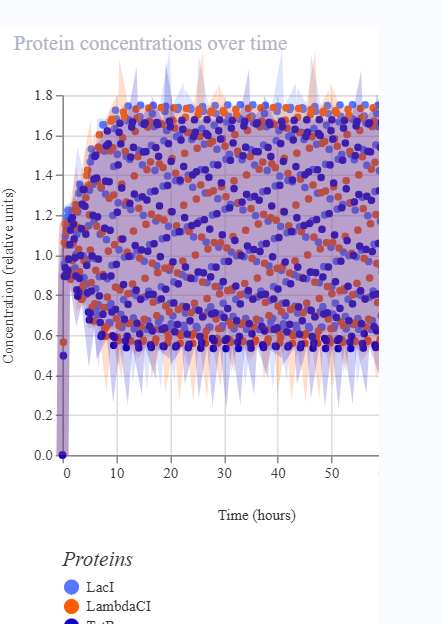
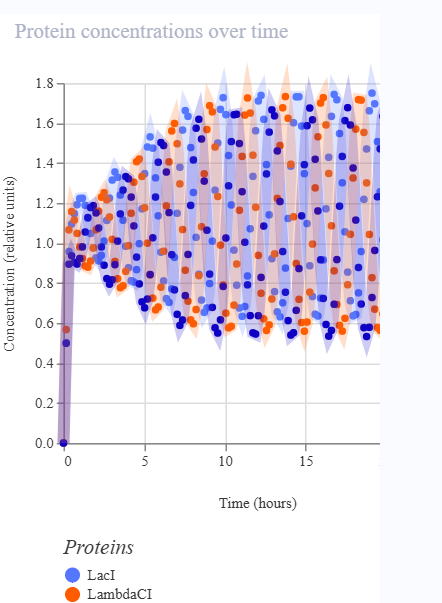
The repressilator consists of three transcription units arranged in a cyclic inhibitory network. Each gene produces a repressor that inhibits the next gene in the circuit. Simulation results show oscillatory behavior in protein concentrations over time, confirming the expected dynamic behavior of the repressilator.
Original construct
My copy
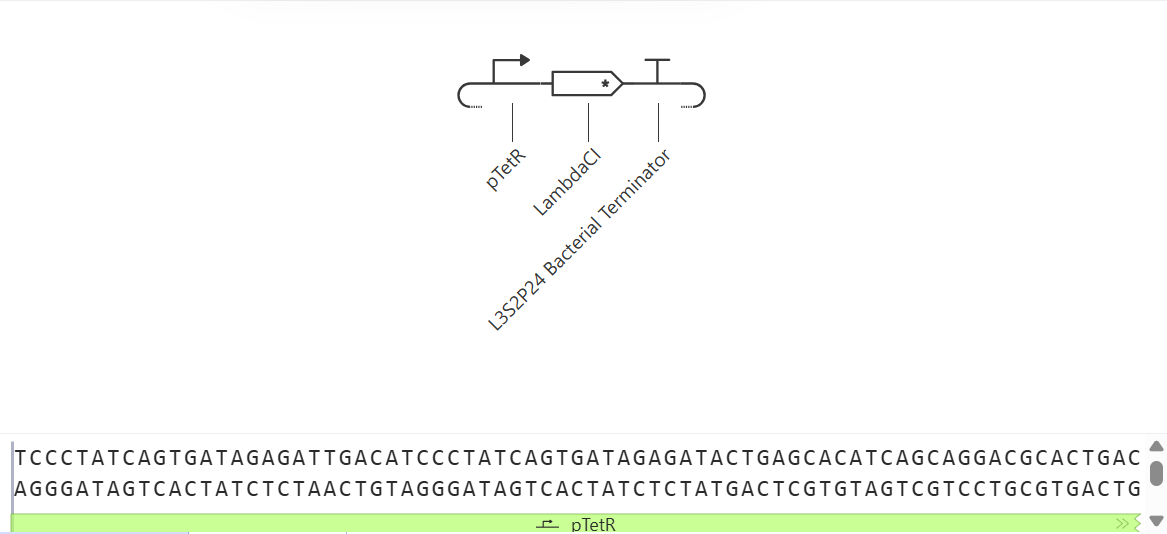
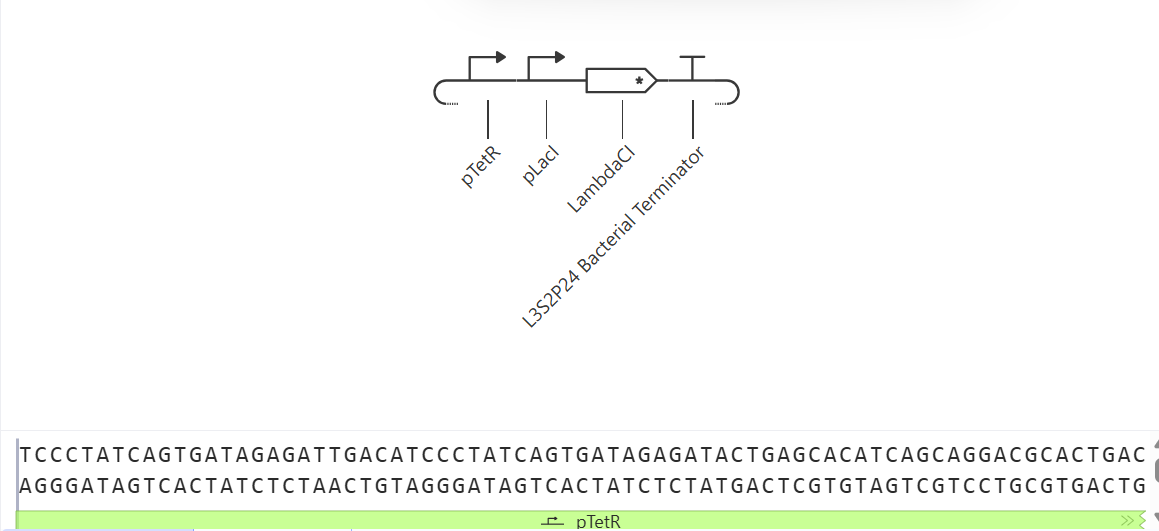
1. Construct A118G Prototype
- Activatable Promoter: Represents the region that detects the A118G polymorphism of the MPR1 gene.
- CDS Placeholder (LambdaCl): Simulates the reporter protein (equivalent to GFP) that indicates the presence of the SNP.
- Terminator: Terminates the transcription of the construct.
- When the A118G sequence is present, the activatable promoter is activated.
- This activation triggers the expression of the CDS placeholder (LambdaCl).
- The terminator ensures that transcription is completed correctly.
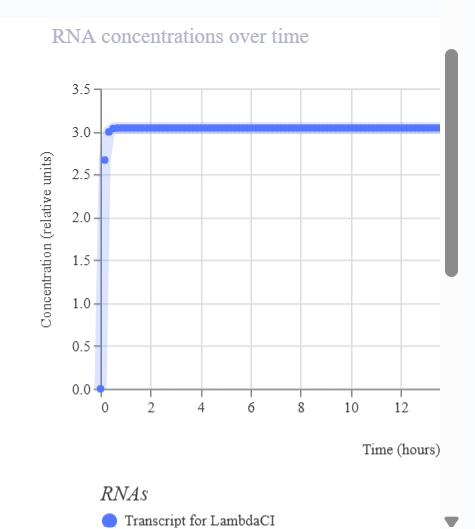
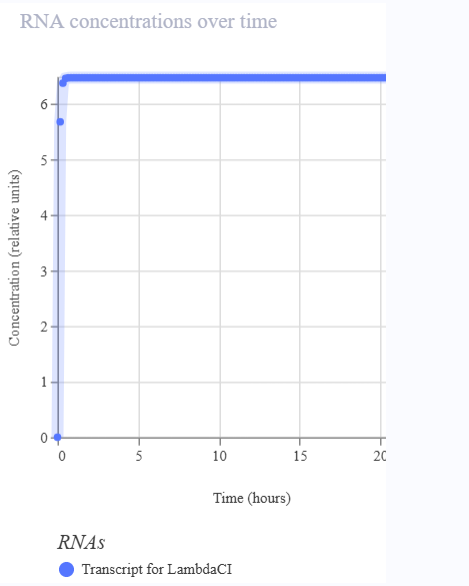
- The simulation illustrates the dynamics of the reporter protein as a function of promoter activation.
- The line representing LambdaCl rises when the promoter is activated, indicating the presence of the A118G SNP.
- If the promoter is not activated (SNP absent), the line remains low or at zero.
- There are no oscillations like those seen in the repressilator; the signal functions as an ON/OFF switch, reflecting the detection of the SNP.
- The construct effectively simulates a genetic biosensor: the presence of the A118G SNP translates into an observable signal (a rise in LambdaCl).
- It serves to visualize and document the biosensor's operation within the simulator, even though it does not utilize the actual GFP protein.
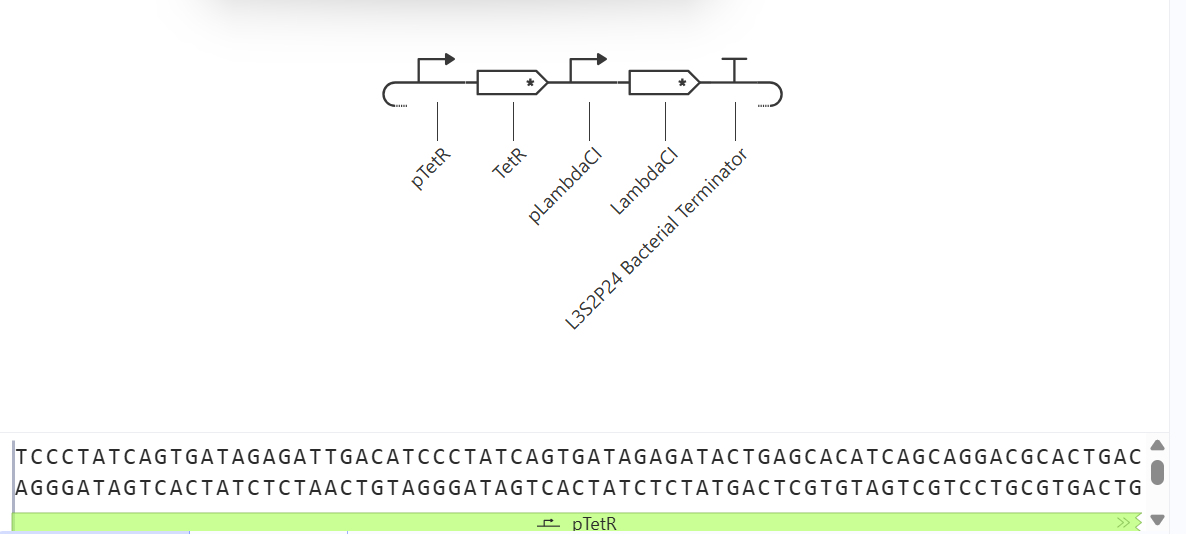
2. Biosensor A118G Amplification
- Activatable promoter: Represents the region that detects the A118G polymorphism of the MPR1 gene.
- TetR CDS placeholder: Functions as an intermediate repressor to simulate signal amplification within the biosensor.
- Reporter promoter: Controlled by TetR, it regulates the expression of the reporter.
- LambdaCl CDS placeholder: Simulates the reporter protein (equivalent to GFP).
- Terminator: Terminates the transcription of the construct.
- When the A118G sequence is present, the activatable promoter becomes active.
- This produces TetR, which regulates the reporter promoter.
- The regulation of the reporter (LambdaCl) simulates signal amplification, increasing the reporter's expression when the SNP is present.
- The terminator ensures that transcription is completed correctly.
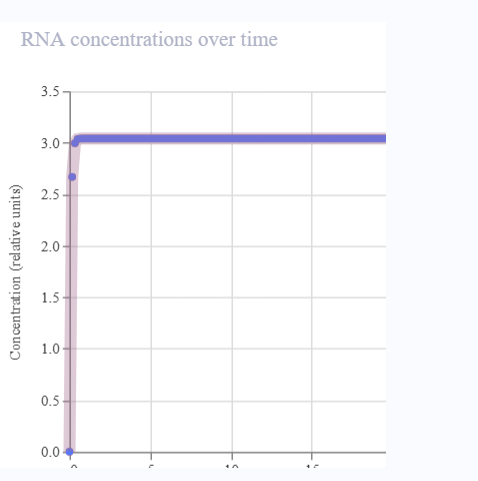
- LambdaCl levels rise when the activatable promoter detects the SNP.
- The signal follows an ON/OFF pattern, representing the activation of the biosensor.
- The construct simulates signal amplification logic using an intermediate repressor.
- Although the graph does not differ significantly from that of the simple construct, the circuit illustrates how the signal could be amplified in a real-world design.
3. A118G AND-type Biosensor
- Activatable Promoter 1: Represents the detection of the A118G polymorphism of the MPR1 gene.
- Activatable Promoter 2: Represents an additional input signal (e.g., another marker or condition).
- LambdaCl CDS Placeholder: Simulates the reporter protein (equivalent to GFP).
- Terminator: Terminates the transcription of the construct.
- Expression of the reporter (LambdaCl) occurs only if both promoters are active.
- This simulates AND logic; both conditions must be met for a signal to be generated.
- The terminator ensures that transcription is completed correctly.
- LambdaCl levels rise only when both promoters are activated simultaneously.
- If only one is activated → the line remains low → no signal is generated.
- It functions as a highly specific switch: ON only when both conditions are met.
- This construct simulates a more specific genetic biosensor, capable of filtering out false signals.
- It allows for the demonstration of AND logic within Asimov, even though the promoters and CDS are placeholders.
- It is useful for illustrating how a more robust biosensor could be designed to detect the A118G SNP in conjunction with another input.
References
- Addgene. “Addgene: What Is Polymerase Chain Reaction (PCR).” Addgene.org, 2019, www.addgene.org/protocols/pcr/.
- ---. “Bacterial Transformation.” Addgene, 13 Nov. 2017, www.addgene.org/protocols/bacterial-transformation/.
- ---. “Molecular Biology Protocol - Restriction Digest of Plasmid DNA.” Addgene.org, 11 Oct. 2016, www.addgene.org/protocols/restriction-digest/.
- addgene. “Addgene: Gibson Assembly Protocol.” Www.addgene.org, www.addgene.org/protocols/gibson-assembly/.
- “Buffers for Biochemical Reactions.” Www.promega.com, www.promega.com/resources/guides/lab-equipment-and-supplies/buffers-for-biochemical-reactions/.
- “Deoxynucleotide Triphosphates (DNTP): Definition & Overview.” Www.excedr.com, www.excedr.com/resources/deoxynucleotide-triphosphates-dntp.
- “DNTPs (Deoxynucleotide Triphosphates).” Promega.com, 2026, www.promega.com/products/pcr/taq-polymerase/deoxynucleotide-triphosphates-dntps/?catNum=U1205. Accessed 22 Mar. 2026.
- “DNTPs: Structure, Role & Applications.” Baseclick GmbH, 15 Jan. 2025, www.baseclick.eu/science/glossar/dntps/.
- Excedr. “What Is the Role of MgCl2 in PCR Amplification Reactions?” Www.excedr.com, 19 Apr. 2022, www.excedr.com/resources/what-is-the-role-of-mgcl2-in-pcr.
- National Human Genome Research institute. “Reacción En Cadena de La Polimerasa (PCR) | NHGRI.” Genome.gov, 2024, www.genome.gov/es/genetics-glossary/Reacci%C3%B3n-en-cadena-de-la-polimerasa-PCR.
- New England Biolabs. “PhusionTM High-Fidelity PCR Master Mix with HF Buffer.” Neb.com, 2026, www.neb.com/en-us/products/m0531-phusion-high-fidelity-pcr-master-mix-with-hf-buffer?srsltid=AfmBOordry1wI142MJKbWwHlBq8WS18SoRym6puZTlpVRlkvbUOZgHqf. Accessed 22 Mar. 2026.
- ---. Neb.com, 2016, www.neb.com/en-us/applications/cloning-and-synthetic-biology/dna-preparation/restriction-enzyme-digestion?srsltid=AfmBOopkvxDAKNCNwCGI1NxjmySwglebU_iEos8Cy0py_cihV-FqNn1N. Accessed 22 Mar. 2026.
- “PCR Cycling Parameters—Six Key Considerations for Success - US.” Www.thermofisher.com, www.thermofisher.com/us/en/home/life-science/cloning/cloning-learning-center/invitrogen-school-of-molecular-biology/pcr-education/pcr-reagents-enzymes/pcr-cycling-considerations.html.
- “Phusion High-Fidelity PCR Master Mix with HF Buffer.” Www.thermofisher.com, www.thermofisher.com/order/catalog/product/F531L.
- “Restriction Enzymes Digestion | GenScript.” Genscript.com, 2026, www.genscript.com/what-is-restriction-digestion.html?__cf_chl_tk=crrewo623z2zjnCx9kOFdTqz0j9BoDdXu3XELXiUgz0-1773342635-1.0.1.1-gQT1okoakeY4JnNUIaPNcP6YhydQbvodS2WN._vGsHI. Accessed 22 Mar. 2026.
- SnapGene. “Golden Gate Assembly.” Www.snapgene.com, www.snapgene.com/guides/golden-gate-assembly.
- “The Plasmid Cloning Cycle - Snapgene.” Www.snapgene.com, www.snapgene.com/guides/the-plasmid-cloning-cycle.
- University of Utah. “PCR.” Utah.edu, 2000, learn.genetics.utah.edu/content/labs/pcr/.