Week 09 HW: Cell-free systems

General homework questions
1. Explain the main advantages of cell-free protein synthesis over traditional in vivo methods, specifically in terms of flexibility and control over experimental variables. Name at least two cases where cell-free expression is more beneficial than cell production.
Cell-free protein synthesis is usually better than traditional in vivo expression systems because it is an open system. It allows direct tuning of the reaction conditions, monitoring the process, and adding components like labels, chaperones, cofactors, or non-natural amino acids much more easily than in living cells.
Control and flexibility: This system avoids the limits of cell growth and cellular metabolism; consequently, its resources can be devoted to making the target protein. Meaning that, it is not constrained by cell viability, membrane transport, or homeostasis, so you can rapidly change variables such as buffer composition, template type, ion concentrations, folding aids, and amino acid content. That makes optimization faster and experimental control much tighter than in vivo expression, where many internal cellular processes are hard to isolate or adjust.
When is it especially beneficial?
Cell-free expressions are especially useful for:
- Toxic proteins, because there are no living host cells to damage
- Difficult-to-express proteins, specifically including some proteins that are unstable or rapidly degraded in cells
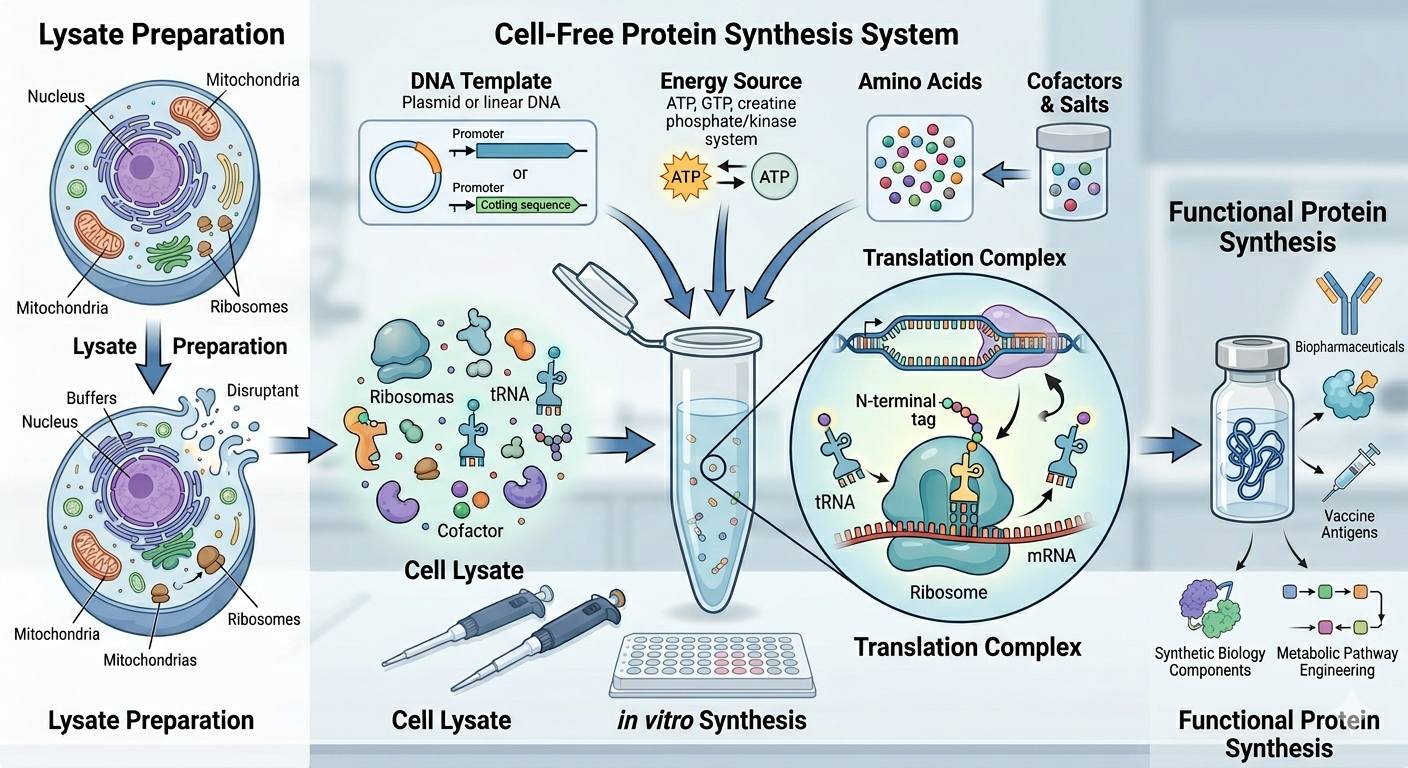
2. Describe the main components of a cell-free expression system and explain the role of each component.
Cell extract: This is the biological “machinery” of the system. It supplies ribosomes, tRNAs, initiation/elongation/termination factors, and other enzymes needed for transcription and translation.
Template DNA or mRNA: This provides the instructions for the protein you want to make.
Energy system: The reaction includes ATP/GTP and often an energy-regeneration module, because protein synthesis consumes a lot of energy.
Amino acids: These are the raw building blocks that get assembled into the target protein.
Salts and cofactors: Magnesium, potassium, and other cofactors help maintain ribosome structure and support enzymatic activity throughout the reaction.
Buffer system: The buffer keeps the pH and ionic conditions in a range during the process.
3. Why is energy provision regeneration critical in cell-free systems? Describe a method you could use to ensure continuous ATP supply in your cell-free experiment.
Energy provision regeneration is required in cell-free systems because the whole process of protein synthesis requires a large amount of energy; the reaction will stop quickly if ATP is not continuously replenished.
Efficient regeneration also helps avoid the buildup of inhibitory byproducts, which can further suppress protein production.
A method used to maintain ATP is an acetyl phosphate/acetate kinase system, which can regenerate ATP continuously while also helping reduce phosphate accumulation in some formats.
4. Compare prokaryotic versus eukaryotic cell-free expression systems. Choose a protein to produce in each system and explain why.
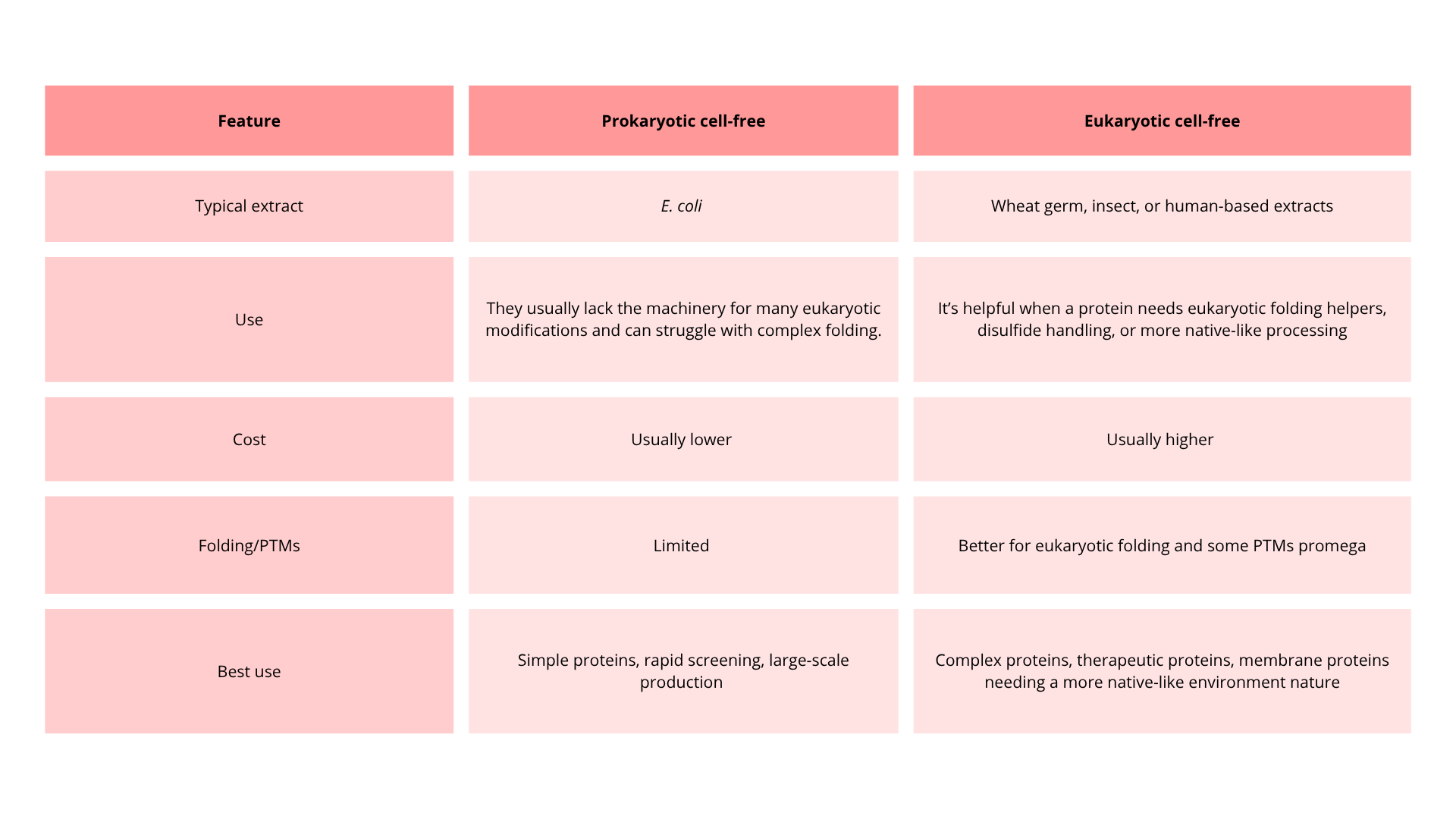

Consequently, A prokaryotic cell-free system is a strong choice for sfGFP or another easy-to-express bacterial reporter protein because these proteins are compact, do not require elaborate post-translational processing, and can be produced efficiently in an E. coli extract.
A eukaryotic cell-free system might be more complex. Still, it is a better choice for human insulin, specifically proinsulin, because insulin production benefits from a more eukaryotic-like folding environment and processing workflow.
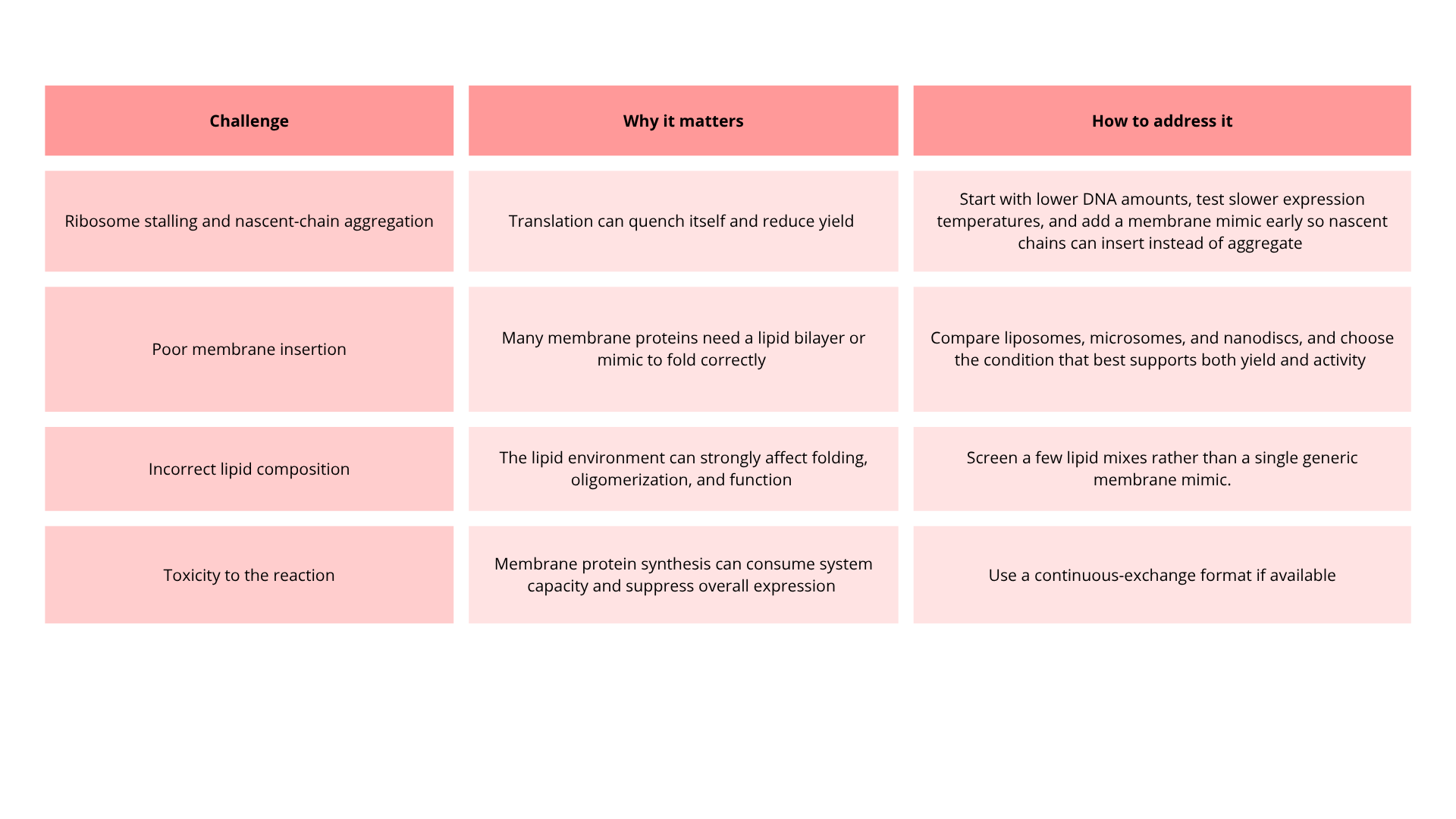
5. How would you design a cell-free experiment to optimize the expression of a membrane protein? Discuss the challenges and how you would address them in your setup.
I believe it is a good approach to develop an experiment where I can run a small factorial screen that varies three things:
- The DNA template amount
- The membrane-mimetic environment
- The reaction conditions, such as temperature and incubation time.
The main goal is to find the condition that gives the best balance between total yield and correctly folded, membrane-inserted protein, because membrane proteins often stall translation and aggregate before they fold properly.
Set up
Use a cell-free transcription/translation system, then compare a modest matrix of conditions in parallel. I suggest that a practical starting design is:
- 2 to 3 DNA concentrations
- 3 membrane environments: no membrane mimic, liposomes or microsomes, and nanodiscs
- 2 to 3 temperatures
- 2 harvest times
That provides a general screen while still testing the major variables known to affect membrane protein synthesis in cell-free systems.
It is necessary to track both expression and function, not just total protein.

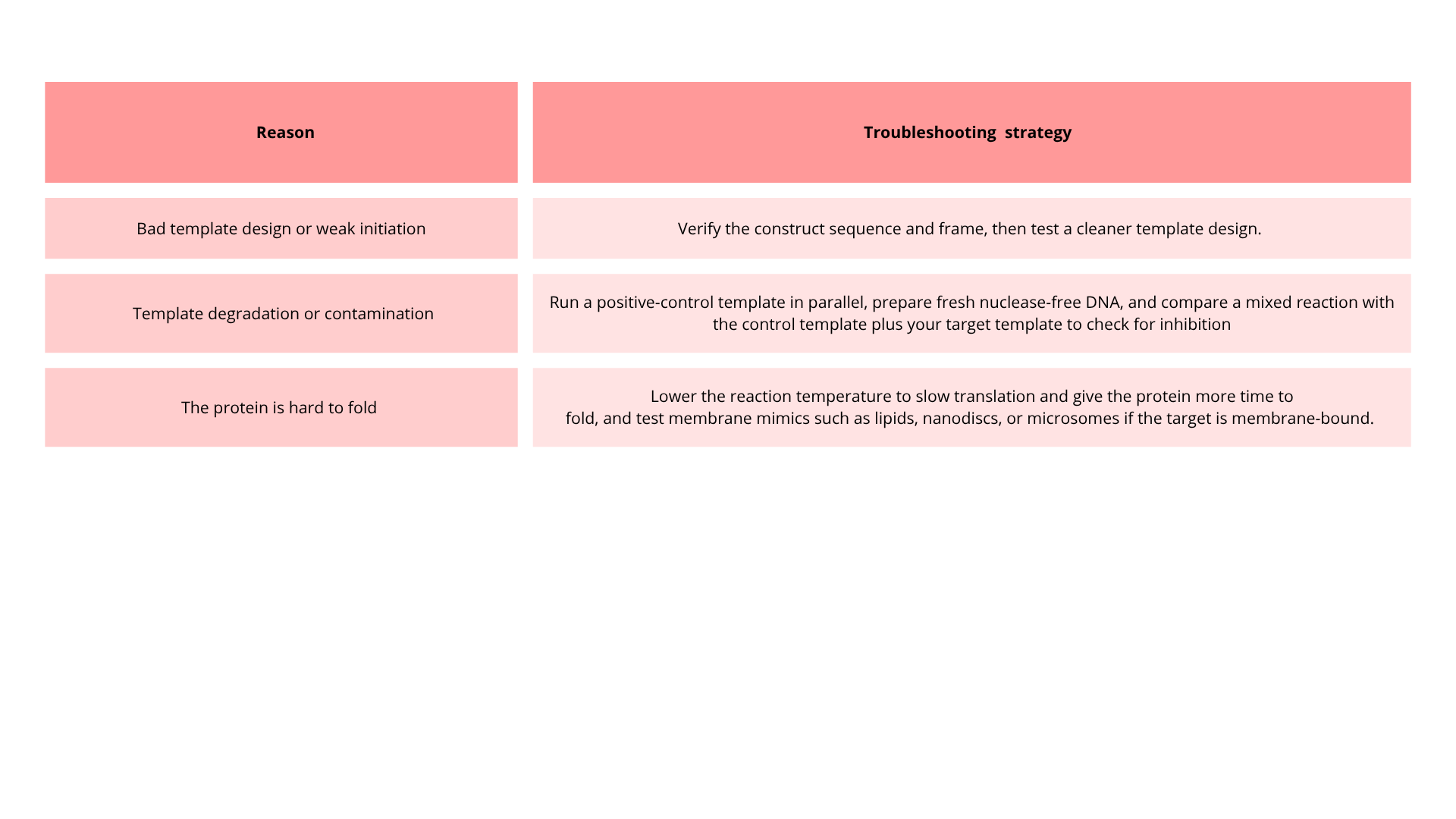
6. Imagine you observe a low yield of your target protein in a cell-free system. Describe three possible reasons for this and suggest a troubleshooting strategy for each.

Homework question from Kate Adamala
Design an example of a useful synthetic minimal cell as follows:
1. Pick a function and describe it.
a. What would your synthetic cell do? What is the input and what is the output?
Synthetic minimal cell design: OPRM1 A118G biosensor
My synthetic minimal cell would be a liposome-encapsulated cell-free biosensor designed to detect the A118G polymorphism in the OPRM1 gene. The function of the synthetic cell is to identify a specific single-nucleotide polymorphism in a DNA sample. The input would be a DNA sample containing the target sequence, and the output would be a measurable fluorescent signal, such as GFP expression, only when the G allele is present.
Input: DNA containing the OPRM1 A118G sequence.
Output: Fluorescence or another visible signal indicating the presence of the G allele.
b. Could this function be realized by cell-free Tx/Tl alone, without encapsulation?
This function could be performed by cell-free Tx/Tl. However, a cell-free system inside a vesicle gives compartmentalization, protection of the reaction, and better control over the sensing environment.
c. Could this function be realized by a genetically modified natural cell?
Yes, this function could also be engineered in a genetically modified natural cell, such as E. coli or yeast. However, a synthetic minimal cell is better for a biosensor because it avoids the complexity of living metabolism and cell growth
d. Describe the desired outcome of your synthetic cell operation.
The desired outcome is a vesicle that can be used as a tiny diagnostic device: when it encounters the correct DNA sequence, it produces a fluorescent signal that can be measured externally.
2. Design all components that would need to be part of your synthetic cell.
a. What would be the membrane made of?
The membrane would be a lipid bilayer liposome made from phospholipids
b. What would you encapsulate inside? Enzymes, small molecules.
Inside the synthetic cell I would encapsulate:
- A cell-free Tx/Tl system from E. coli
- A DNA recognition module specific for the A118G polymorphism
- A reporter gene such as gfp
c. Which organism your Tx/Tl system will come from? Is bacterial OK, or do you need a mammalian system for some reason? (hint: for example, if you want to use small molecule modulated promotors, like Tet-ON, you need mammalian)
I would use a bacterial cell-free system, especially from E. coli, because it is well characterized, easy to use, and suitable for a minimal synthetic cell.
d. How will your synthetic cell communicate with the environment? (hint: are substrates permeable? or do you need to express the membrane channel?)
- Allowing the DNA sample or small molecules to enter during assay setup
- Possibly using a membrane pore to permit small-molecule exchange
- Releasing a fluorescent signal that can be detected outside the vesicle
3. Experimental details
a. List all lipids and genes. (bonus: find the specific genes; for example, instead of just saying “small molecule membrane channel” pick the actual gene.)
- gfp for fluorescence reportin
- hla if a membrane pore is needed
- A synthetic DNA-sensing module designed to distinguish the A and G alleles of OPRM1
b. How will you measure the function of your system?
Fluorescence intensity Comparison of samples with A allele, G allele, and no-template control
Homework question from Peter Nguyen
Freeze-dried cell-free systems can be incorporated into all kinds of materials as biological sensors or as inducible enzymes to modify the material itself or the surrounding environment. Choose one application field — Architecture, Textiles/Fashion, or Robotics — and propose an application using cell-free systems that are functionally integrated into the material. Answer each of these key questions for your proposal pitch:
1. Write a one-sentence summary pitch sentence describing your concept.
A cell-free biosensor band for astronauts that detects toxic environmental exposure and produces an immediate visible signal
2. How will the idea work, in more detail? Write 3-4 sentences or more.
It will be a flexible wearable band or patch designed for astronauts or people working in extreme environments. The material would contain cell-free biosensors that become active when a small amount of water or a sample is added, and they would detect dangerous environmental toxins such as heavy metals or other harmful chemicals. When the target toxin is present, the cell-free system would express a reporter such as GFP, creating an easy-to-read signal.
3. What societal challenge or market need will this address?
This idea addresses the need for portable, low-maintenance safety monitoring in places where conventional sensors may be expensive or difficult to use. It could be valuable for astronauts, space missions, emergency responders, industrial workers, or anyone exposed to toxic environments.
4. How do you envision addressing the limitation of cell-free reactions (e.g., activation with water, stability, one-time use)?
The main limitation of cell-free systems is that they typically require water to initiate and can lose activity over time. I would solve this by freeze-drying the biosensor inside the textile or material, so it stays stable during storage and activates only when needed. To make it reusable or longer-lasting, the band could use replaceable sensing patches.
Homework question from Ally Huang
Freeze-dried cell-free reactions have great potential in space, where resources are constrained. As described in my talk, the Genes in Space competition challenges students to consider how biotechnology, including cell-free reactions, can be used to solve biological problems encountered in space. While the competition is limited to only high school students, your assignment will be to develop your own mock Genes in Space proposal to practice thinking about biotech applications in space!
For this particular assignment, your proposal is required to incorporate the BioBits® cell-free protein expression system, but you may also use the other tools in the Genes in Space toolkit (the miniPCR® thermal cycler and the P51 Molecular Fluorescence Viewer). For more inspiration, check out https://www.genesinspace.org/ .
1. Provide background information that describes the space biology question or challenge you propose to address. Explain why this topic is significant for humanity, relevant for space exploration, and scientifically interesting. (Maximum 100 words)
In space exploration, microgravity affects cellular function, reducing proliferation and altering metabolism. Understanding how microgravity impacts cell growth is essential for astronaut health and for maintaining effective medications and microbial cultures. BioBits® lets us test protein activity without using live cells. By measuring fluorescence under microgravity and UV radiation, we can see how these conditions affect cell growth. This experiment will show how space affects metabolism and growth in a simple, visual way.
2. Name the molecular or genetic target that you propose to study. Examples of molecular targets include individual genes and proteins, DNA and RNA sequences, or broader -omics approaches. (Maximum 30 words)
GFP protein expressed in the BioBits® system, used as a visual indicator of metabolic activity to measure how simulated cell growth changes under microgravity and UV radiation.
3. Describe how your molecular or genetic target relates to the space biology question or challenge your proposal addresses. (Maximum 100 words)
GFP fluorescence reflects the metabolic activity of the BioBits® system, serving as an indicator of simulated cell growth: higher fluorescence indicates faster growth, while lower fluorescence indicates slower growth. By comparing fluorescence under microgravity and UV radiation, we can assess how these space conditions affect growth rates. This information is relevant for predicting effects on human cells, microbial cultures, and protein stability in space. Using GFP as an indicator allows quick, safe, and portable measurements, providing practical insights for protecting astronaut health and supporting biotechnological applications during long space missions.
4. Clearly state your hypothesis or research goal and explain the reasoning behind it. (Maximum 150 words)
Hypothesis:
Microgravity reduces the simulated cellular growth rate compared to UV radiation, with GFP fluorescence serving as a reliable indicator of metabolic activity.
Research goal:
To evaluate how microgravity and UV radiation affect simulated cell growth using BioBits®. This will help determine whether microgravity slows critical metabolic processes, which has direct implications for astronaut health, drug stability, and microbial culture productivity. We expect lower fluorescence under microgravity, indicating reduced activity, while UV exposure may produce a different pattern. This experiment will generate preliminary data to inform strategies that mitigate microgravity effects, such as optimizing antioxidants, nutrients, or cellular support systems for long-term missions.
5. Outline your experimental plan - identify the sample(s) you will test in your experiment, including any necessary controls, the type of data or measurements that will be collected, etc. (Maximum 100 words)
Two BioBits® systems with GFP will be set up: one simulating microgravity and one exposed to UV radiation. Controls include a system without microgravity or UV, and one with standard GFP for calibration. Fluorescence will be measured every 10 minutes for 1 hour using the P51 Molecular Fluorescence Viewer. Fluorescence intensity will be compared between conditions to infer relative growth rates. Additional experiments may include antioxidants or nutrients to test mitigation strategies. Data will be recorded quantitatively and graphed to analyze patterns in metabolic activity and simulated growth under space conditions
Homework Part B: Individual Final Project
Project slide:
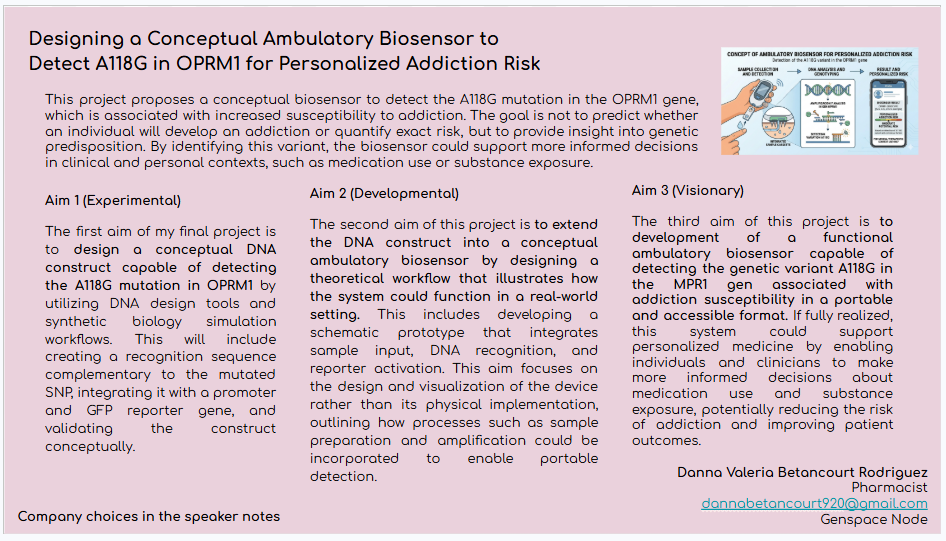


References
- Baldo, Thaisa A., et al. “Wearable and Biodegradable Sensors for Clinical and Environmental Applications.” ACS Applied Electronic Materials, 10 Dec. 2020, https://doi.org/10.1021/acsaelm.0c00735.
- ---. “Wearable and Biodegradable Sensors for Clinical and Environmental Applications.” ACS Applied Electronic Materials, 10 Dec. 2020, https://doi.org/10.1021/acsaelm.0c00735.
- Banks, Alice M., et al. “Key Reaction Components Affect the Kinetics and Performance Robustness of Cell-Free Protein Synthesis Reactions.” Computational and Structural Biotechnology Journal, vol. 20, 1 Jan. 2022, pp. 218–229, www.sciencedirect.com/science/article/pii/S2001037021005213, https://doi.org/10.1016/j.csbj.2021.12.013.
- ---. “Key Reaction Components Affect the Kinetics and Performance Robustness of Cell-Free Protein Synthesis Reactions.” Computational and Structural Biotechnology Journal, vol. 20, 1 Jan. 2022, pp. 218–229, www.sciencedirect.com/science/article/pii/S2001037021005213, https://doi.org/10.1016/j.csbj.2021.12.013.
- Borhani, Shayan G, et al. “On-Demand Insulin Manufacturing Using Cell-Free Systems with an “On-Column” Conversion Approach.” New Biotechnology, 1 June 2025, https://doi.org/10.1016/j.nbt.2025.06.002. Accessed 12 Sept. 2025.
- ---. “On-Demand Insulin Manufacturing Using Cell-Free Systems with an “On-Column” Conversion Approach.” New Biotechnology, 1 June 2025, https://doi.org/10.1016/j.nbt.2025.06.002. Accessed 12 Sept. 2025.
- Calhoun, Kara A., and James R. Swartz. “Energy Systems for ATP Regeneration in Cell-Free Protein Synthesis Reactions.” In Vitro Transcription and Translation Protocols, 2007, pp. 3–17, https://doi.org/10.1007/978-1-59745-388-2_1.
- “Cell-Free Expression Support—Troubleshooting | Thermo Fisher Scientific - US.” Thermofisher.com, 2025, www.thermofisher.com/ca/en/home/technical-resources/technical-reference-library/protein-expression-support-center/cell-free-expression-support/cell-free-expression-support-troubleshooting.html. Accessed 7 Apr. 2026.
- “Cell-Free Protein Expression System- CUSABIO.” Cusabio.com, 2026, www.cusabio.com/cell-free-expression-system.html. Accessed 7 Apr. 2026.
- “Cell-Free Protein Synthesis Explained | IDT.” Integrated DNA Technologies, 2015, www.idtdna.com/pages/applications/cell-free-protein-synthesis.
- Chauhan, Nidhi, et al. “A Review on Biosensor Approaches for the Detection of Hazardous Elements in Water.” Talanta Open, 1 Aug. 2025, pp. 100536–100536, https://doi.org/10.1016/j.talo.2025.100536.
- Grooms, Kelly. “Moving out of the Cell: Advantages of Cell-Free Protein Expression - Promega Connections.” Promega Connections, 22 Aug. 2016, www.promegaconnections.com/moving-out-of-the-cell-advantages-of-cell-free-protein-expression/.
- ---. “Moving out of the Cell: Advantages of Cell-Free Protein Expression - Promega Connections.” Promega Connections, 22 Aug. 2016, www.promegaconnections.com/moving-out-of-the-cell-advantages-of-cell-free-protein-expression/.
- Kim, D M, and J R Swartz. “Prolonging Cell-Free Protein Synthesis with a Novel ATP Regeneration System.” Biotechnology and Bioengineering, vol. 66, no. 3, 1999, pp. 180–8, pubmed.ncbi.nlm.nih.gov/10577472/.
- ---. “Prolonging Cell-Free Protein Synthesis with a Novel ATP Regeneration System.” Biotechnology and Bioengineering, vol. 66, no. 3, 1999, pp. 180–8, pubmed.ncbi.nlm.nih.gov/10577472/.
- Kim, Ho-Cheol, and Dong-Myung Kim. “Methods for Energizing Cell-Free Protein Synthesis.” Journal of Bioscience and Bioengineering, vol. 108, no. 1, July 2009, pp. 1–4, pubmed.ncbi.nlm.nih.gov/19577183/, https://doi.org/10.1016/j.jbiosc.2009.02.007.
- ---. “Methods for Energizing Cell-Free Protein Synthesis.” Journal of Bioscience and Bioengineering, vol. 108, no. 1, July 2009, pp. 1–4, pubmed.ncbi.nlm.nih.gov/19577183/, https://doi.org/10.1016/j.jbiosc.2009.02.007.
- Meyer, Conary, et al. “Designer Artificial Environments for Membrane Protein Synthesis.” Nature Communications, vol. 16, no. 1, 10 May 2025, www.nature.com/articles/s41467-025-59471-1, https://doi.org/10.1038/s41467-025-59471-1. Accessed 7 Apr. 2026.
- ---. “Designer Artificial Environments for Membrane Protein Synthesis.” Nature Communications, vol. 16, no. 1, 10 May 2025, www.nature.com/articles/s41467-025-59471-1, https://doi.org/10.1038/s41467-025-59471-1. Accessed 7 Apr. 2026.
- Nguyen, Peter Q., et al. “Wearable Materials with Embedded Synthetic Biology Sensors for Biomolecule Detection.” Nature Biotechnology, vol. 39, 28 June 2021, pp. 1–9, www.nature.com/articles/s41587-021-00950-3, https://doi.org/10.1038/s41587-021-00950-3.
- ---. “Wearable Materials with Embedded Synthetic Biology Sensors for Biomolecule Detection.” Nature Biotechnology, vol. 39, 28 June 2021, pp. 1–9, www.nature.com/articles/s41587-021-00950-3, https://doi.org/10.1038/s41587-021-00950-3.
- Schaub, Theresa. “Cell-Free Protein Synthesis: A Faster, More Flexible Alternative to Traditional Expression.” Cosmo Bio USA, 2 Mar. 2026, www.cosmobiousa.com/blog/why-cell-free-protein-expression-can-be-better-than-cell-based-systems. Accessed 7 Apr. 2026.
- Steinkühler, Jan, et al. “Improving Cell-Free Expression of Model Membrane Proteins by Tuning Ribosome Cotranslational Membrane Association and Nascent Chain Aggregation.” ACS Synthetic Biology, vol. 13, no. 1, 27 Dec. 2023, pp. 129–140, https://doi.org/10.1021/acssynbio.3c00357. Accessed 11 Dec. 2025.
- ---. “Improving Cell-Free Expression of Model Membrane Proteins by Tuning Ribosome Cotranslational Membrane Association and Nascent Chain Aggregation.” ACS Synthetic Biology, vol. 13, no. 1, 27 Dec. 2023, pp. 129–140, https://doi.org/10.1021/acssynbio.3c00357. Accessed 11 Dec. 2025.
- UMBC. “CAST | Center for Advanced Sensor Technology.” Umbc.edu, 2020, cast.umbc.edu/research-new/manufacturing-of-insulin-using-cell-free-systems-cfs/.
- ---. “CAST | Center for Advanced Sensor Technology.” Umbc.edu, 2020, cast.umbc.edu/research-new/manufacturing-of-insulin-using-cell-free-systems-cfs/.Wan, Xinyi. “Synthetic Biology Enabled Cellular and Cell-Free Biosensors for Environmental Contaminants.” Ed.ac.uk, The University of Edinburgh, 6 July 2019, era.ed.ac.uk/items/eada344c-6d52-4c51-8714-73b612b1848b. Accessed 7 Apr. 2026.
- ---. “Synthetic Biology Enabled Cellular and Cell-Free Biosensors for Environmental Contaminants.” Ed.ac.uk, The University of Edinburgh, 6 July 2019, era.ed.ac.uk/items/eada344c-6d52-4c51-8714-73b612b1848b. Accessed 7 Apr. 2026.
- “What Are the Advantages of Cell-Free Protein Expression System over the Traditional in Vivo Systems? | AAT Bioquest.” Aatbio.com, 2023, www.aatbio.com/resources/faq-frequently-asked-questions/what-are-the-advantages-of-cell-free-protein-expression-system-over-the-traditional-in-vivo-systems. Accessed 7 Apr.
- 2026.Whittaker, James W. “Cell-Free Protein Synthesis: The State of the Art.” Biotechnology Letters, vol. 35, no. 2, 21 Oct. 2012, pp. 143–152, https://doi.org/10.1007/s10529-012-1075-4.
- ---. “Cell-Free Protein Synthesis: The State of the Art.” Biotechnology Letters, vol. 35, no. 2, 21 Oct. 2012, pp. 143–152, https://doi.org/10.1007/s10529-012-1075-4.“Why Use Cell-Free Protein Expression?” Cube Biotech, 2014, cube-biotech.com/our-science/cell-free-lysates/cell-free-expression/.
- “Why Use Cell-Free Protein Expression?” Cube Biotech, 2014, cube-biotech.com/our-science/cell-free-lysates/cell-free-expression/.
- Wyss Institute. “Wearable Synthetic Biology: Clothing That Can Detect Pathogens and Toxins.” YouTube, 29 June 2021, www.youtube.com/watch?v=_pHiIlNDAGk. Accessed 7 Apr. 2026.
- ---. “Wearable Synthetic Biology: Clothing That Can Detect Pathogens and Toxins.” YouTube, 29 June 2021, www.youtube.com/watch?v=_pHiIlNDAGk. Accessed 7 Apr. 2026.
- Zemella, Anne, et al. “Cell-Free Protein Synthesis: Pros and Cons of Prokaryotic and Eukaryotic Systems.” ChemBioChem, vol. 16, no. 17, 19 Oct. 2015, pp. 2420–2431, www.ncbi.nlm.nih.gov/pmc/articles/PMC4676933/, https://doi.org/10.1002/cbic.201500340.---.