Week 11 — Bioproduction & Cloud Labs


Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork
1. Contribute at least one pixel to this global artwork experiment before the editing ends on Sunday 4/19 at 11:59 PM EST.
- A personalized URL was sent to the email address associated with your Discourse account, and you can discuss the artwork on the Discourse.
- If you did not have a chance to contribute, it’s okay, just make sure you become a TA this fall! 😉
Although I was not able to contribute a pixel, I noticed the suggestion about becoming a TA this fall. I think it could be an interesting opportunity, as it would allow for more involvement in collaborative projects like this one and a way to support the course community.
2. Make a note on your HTGAA webpages, including:
- What you contributed to the community bioart project (e.g., “I made part of the DNA on the bottom right plate”)
- What you liked about the project, and what about this collaborative art experiment could be made better for next year.
I explored the final artwork and the collaborative process behind it.
What I liked most about the project is the idea of building a collective artwork where each participant contributes a minimal unit (a pixel), and the result collects a complex and meaningful image. I also found it engaging that the project connects science, art, and community participation.
One aspect that could be improved for next year is increasing reminders and accessibility to the participation link, since it is easy to miss the contribution window. Additionally, having a longer editing period or sending follow-up notifications could help ensure more people are able to contribute.
Overall, even without directly contributing, the project highlights the value of collaboration and creativity in scientific communities.
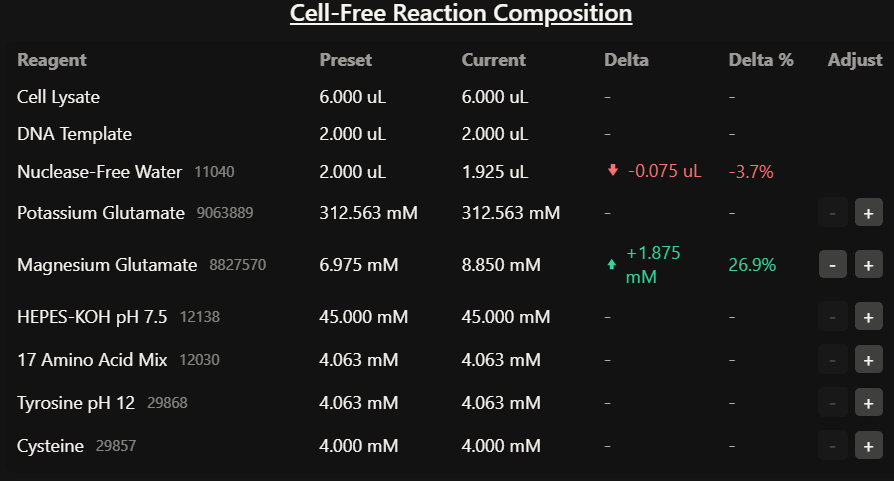
Part B: Cell-Free Protein Synthesis | Cell-Free Reagents
1. Referencing the cell-free protein synthesis reaction composition (the middle box outlined in yellow on the image above, also listed below), provide a 1-2 sentence description of what each component’s role is in the cell-free reaction.
E. coli Lysate
- BL21 (DE3) Star Lysate (includes T7 RNA Polymerase)
Salts/Buffer
- Potassium Glutamate
- HEPES-KOH pH 7.5
- Magnesium Glutamate
- Potassium phosphate monobasic
- Potassium phosphate dibasic
Energy / Nucleotide System
- Ribose
- Glucose
- AMP
- CMP
- GMP
- UMP
- Guanine
Translation Mix (Amino Acids)
- 17 Amino Acid Mix
- Tyrosine
- Cysteine
Additives
- Nicotinamide
Backfill
- Nuclease Free Water
BL21 (DE3) Star Lysate (includes T7 RNA Polymerase): This is a specialized lysate from an expression strain that also contains T7 RNA polymerase, allowing very efficient transcription from T7 promoters.
Potassium Glutamate: This supplies potassium ions and helps stabilize the reaction conditions needed for ribosome activity and enzyme function.
HEPES-KOH pH 7.5: This is the main buffering agent that keeps the reaction at a near-neutral pH, which is optimal for protein synthesis.
Magnesium Glutamate: This provides magnesium ions, which are essential for ribosome stability, nucleotide binding, and overall enzymatic activity.
Potassium Phosphate Monobasic: This contributes to buffering capacity and helps maintain stable chemical conditions in the reaction.
Potassium Phosphate Dibasic: This works with the monobasic form to adjust and stabilize the pH of the buffer system.
Ribose: This serves as a carbon source and helps support nucleotide regeneration pathways in the reaction.
Glucose: This provides a long-lasting energy source that helps sustain ATP regeneration and overall reaction activity.
AMP: This is a nucleotide precursor that helps maintain the adenylate pool and supports energy recycling.
CMP: This is a cytidine nucleotide precursor needed to maintain the pool of RNA building blocks.
GMP: This is a guanosine nucleotide precursor required for RNA synthesis and nucleotide balance.
UMP: This is a uridine nucleotide precursor that supports RNA production.
Guanine: This is a base precursor that can be used to help replenish guanosine nucleotide pools.
17 Amino Acid Mix: This supplies most of the amino acids required for protein synthesis.
Tyrosine: This amino acid is often added separately to fine-tune its concentration and improve solubility or balance.
Cysteine: This is added separately because it is important for protein structure and may require special handling in the mixture.
Nicotinamide: This supports redox-related reactions and can help maintain metabolic activity in the lysate.
Nuclease-Free Water This is used to complete the reaction volume while avoiding degradation of DNA or RNA by nucleases.
2. Describe the main differences between the 1-hour optimized PEP-NTP master mix and the 20-hour NMP-Ribose-Glucose master mix shown in the Google Slide above. (2-3 sentences)
The 1-hour optimized PEP-NTP master mix is designed for fast, high-energy protein synthesis, using PEP as a rapid ATP-regenerating substrate and preloaded NTPs to support quick transcription. In contrast, the 20-hour NMP-Ribose-Glucose master mix is built for longer reactions, relying on glucose and ribose plus NMPs as a more sustained, lower-cost energy/nucleotide regeneration system that supports extended protein production over time.
Part C: Planning the Global Experiment | Cell-Free Master Mix Design
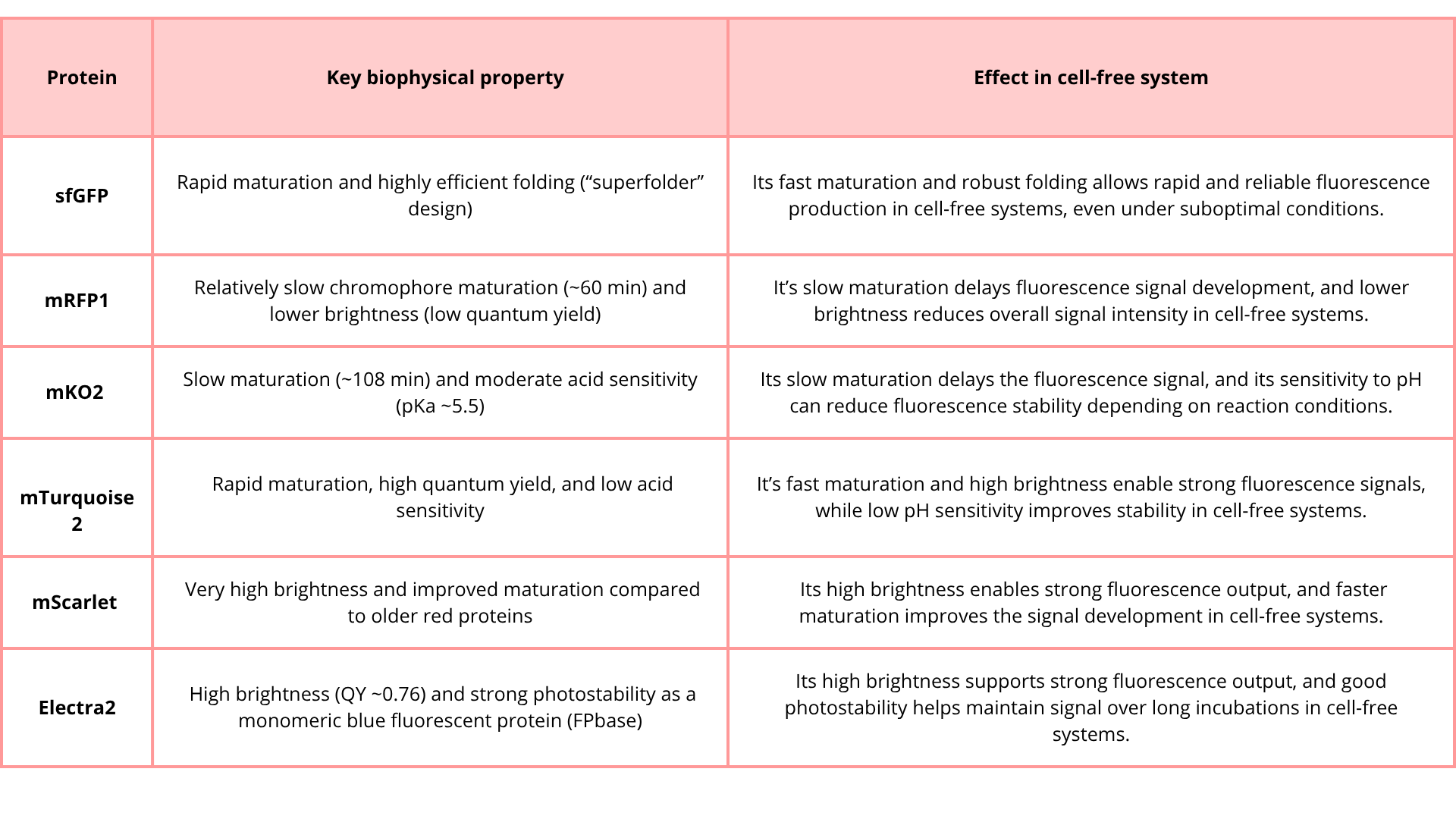
1. Given the 6 fluorescent proteins we used for our collaborative painting, identify and explain at least one biophysical or functional property of each protein that affects expression or readout in cell-free systems. (Hint: options include maturation time, acid sensitivity, folding, oxygen dependence, etc) (1-2 sentences each)
- sfGFP
- mRFP1
- mKO2
- mTurquoise2
- mScarlet_I
- Electra2
The amino acid sequences are shown in the HTGAA Cell-Free Benchling folder.

2. Create a hypothesis for how adjusting one or more reagents in the cell-free mastermix could improve a specific biophysical or functional property you identified above, in order to maximize fluorescence over a 36-hour incubation. Clearly state the protein, the reagent(s), and the expected effect.
Protein: mRFP1
Hypothesis: Adding molecular chaperones (e.g., GroEL/ES or DnaK system) and increasing the energy regeneration system in the cell-free mastermix will improve folding efficiency and overall protein yield, leading to increased and more stable fluorescence signal over a 36-hour incubation period.
3. The second phase of this lab will be to define the precise reagent concentrations for your cell-free experiment. You will be assigned artwork wells with specific fluorescent proteins and receive an email with instructions this week (by April 24). You can begin composing master mix compositions here
For mRFP1, I increased magnesium glutamate in the CFPS mastermix because Mg2+ is essential for efficient transcription-translation activity, ribosome stability, and formation of productive translation complexes. In cell-free systems, optimizing magnesium is important because it can directly affect protein yield, folding efficiency, and the total amount of fluorescent protein that accumulates over time, which is especially relevant for a 36-hour incubation.
My original hypothesis also included adding molecular chaperones to improve the folding of mRFP1. Chaperones would be expected to help the newly synthesized protein reach a functional conformation more efficiently, which would increase the amount of properly matured fluorescent protein and therefore improve fluorescence output.
References
- Bartsch, Tabea, et al. “Cell-Free Protein Synthesis with Technical Additives – Expanding the Parameter Space of in Vitro Gene Expression.” Beilstein Journal of Organic Chemistry, vol. 20, 4 Sept. 2024, pp. 2242–2253, https://doi.org/10.3762/bjoc.20.192. Accessed 29 Nov. 2024.
- Calhoun, Kara A., and James R. Swartz. “Energizing Cell-Free Protein Synthesis with Glucose Metabolism.” Biotechnology and Bioengineering, vol. 90, no. 5, 2005, pp. 606–613, onlinelibrary.wiley.com/doi/abs/10.1002/bit.20449, https://doi.org/10.1002/bit.20449.
- “CLS Cell Lines Service GmbH.” Cytion, 2026, www.cytion.com/es/Acerca-de-Cytion/Centro-de-conocimiento/Articulos-y-novedades/Sistemas-sin-celulas-para-la-produccion-de-proteinas-Ventajas-sobre-las-celulas-vivas/. Accessed 23 Apr. 2026.
- Gregorio, Nicole E., et al. “A User’s Guide to Cell-Free Protein Synthesis.” Methods and Protocols, vol. 2, no. 1, 12 Mar. 2019, https://doi.org/10.3390/mps2010024.
- Guzman-Chavez, Fernando, et al. “Constructing Cell-Free Expression Systems for Low-Cost Access.” ACS Synthetic Biology, vol. 11, no. 3, 8 Mar. 2022, pp. 1114–1128, https://doi.org/10.1021/acssynbio.1c00342.
- Lang, Xianshengjie, et al. A Simplified and Highly Efficient Cell-Free Protein Synthesis System for Prokaryotes. 11 Dec. 2025, elifesciences.org/reviewed-preprints/109495, https://doi.org/10.7554/elife.109495.1.
- Lara, Álvaro R. “Producción de Proteínas Recombinantes En Escherichia Coli.” Revista Mexicana de Ingeniería Química, vol. 10, no. 2, 1 Aug. 2011, pp. 209–223, www.scielo.org.mx/scielo.php?script=sci_arttext&pid=S1665-27382011000200006.
- Rubin, Harry. “Intracellular Free Mg2+ and MgATP2- in Coordinate Control of Protein Synthesis and Cell Proliferation.” Nih.gov, University of Adelaide Press, 2024, www.ncbi.nlm.nih.gov/books/NBK507263/.