<Dariya> — HTGAA Spring 2026
About me
Hello! My name is Dariya, I am a Master’s student at Yonsei University, Designer Cells Lab. Currently working on developing a project about the clustering of T cell receptors :)
Hello! My name is Dariya, I am a Master’s student at Yonsei University, Designer Cells Lab. Currently working on developing a project about the clustering of T cell receptors :)
Week 1 HW: Principles and Practices
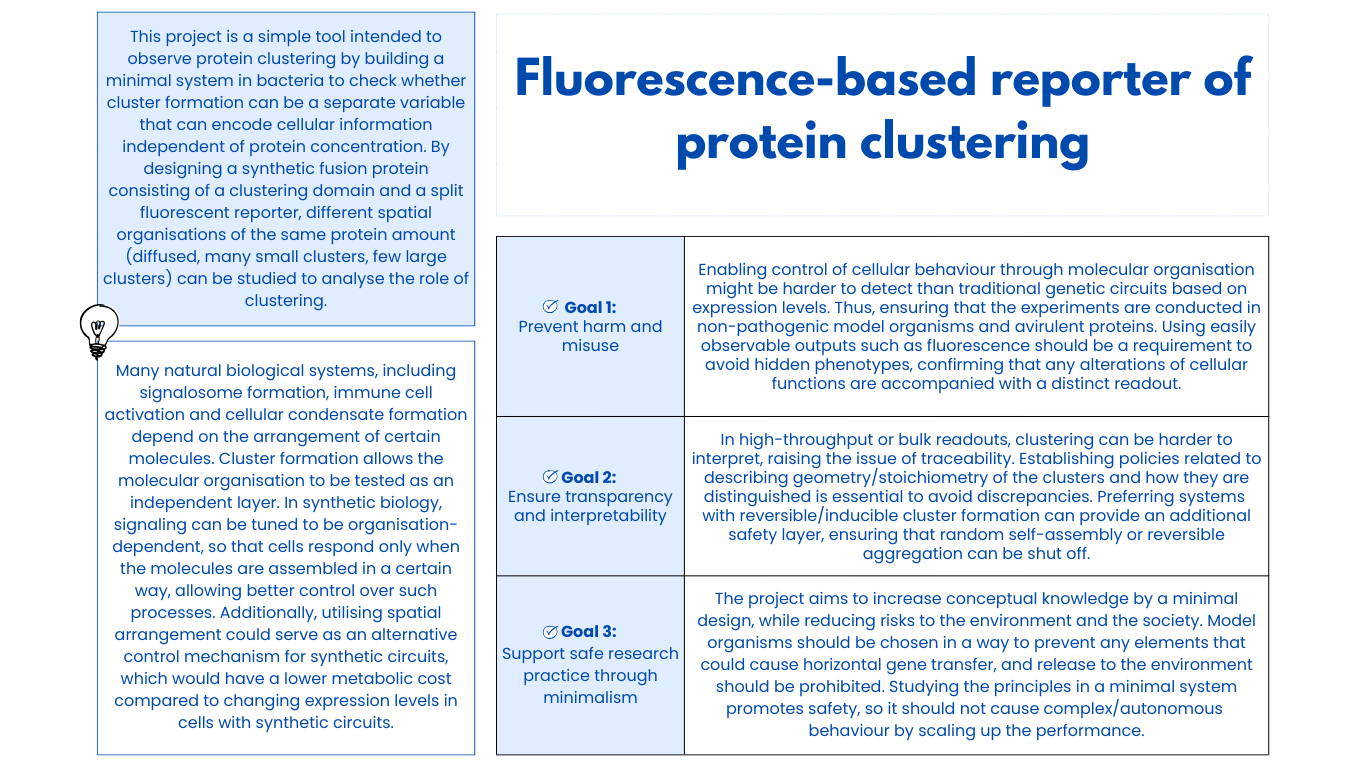
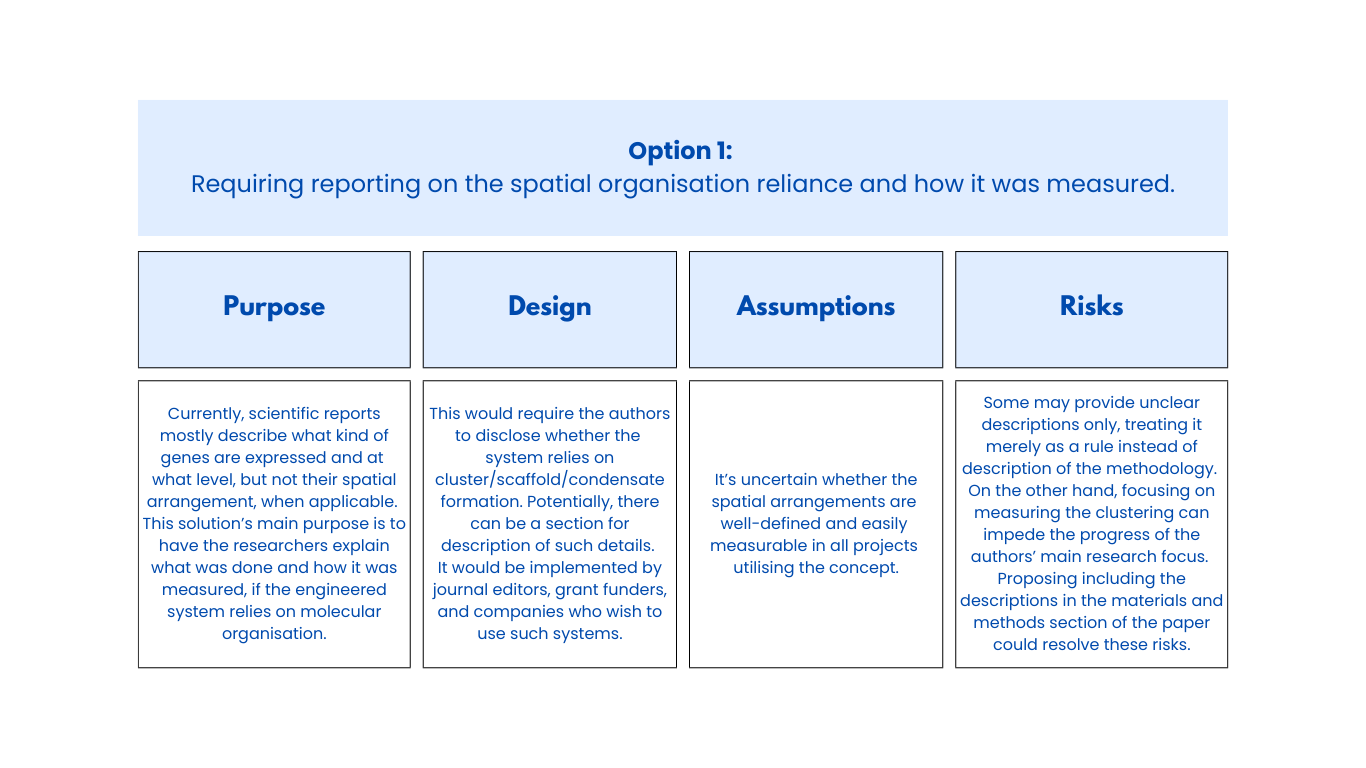
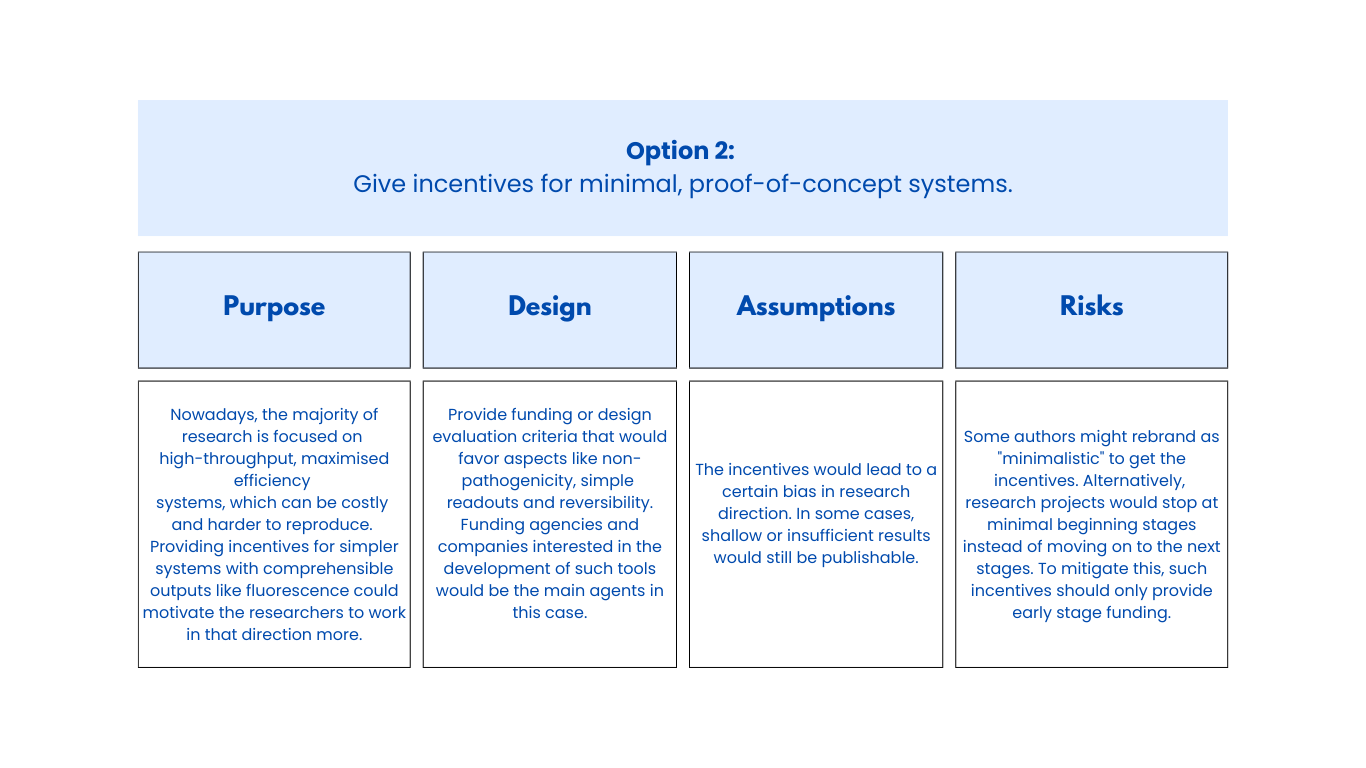
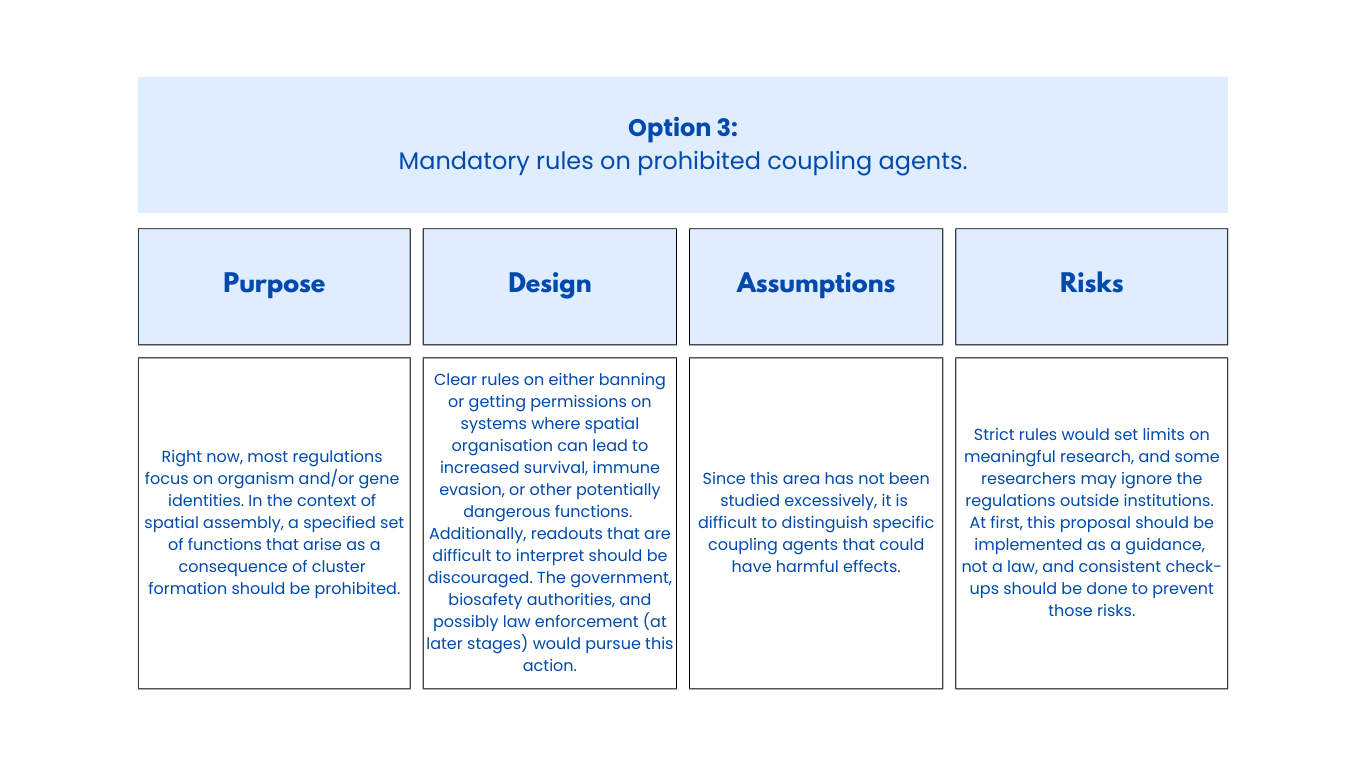
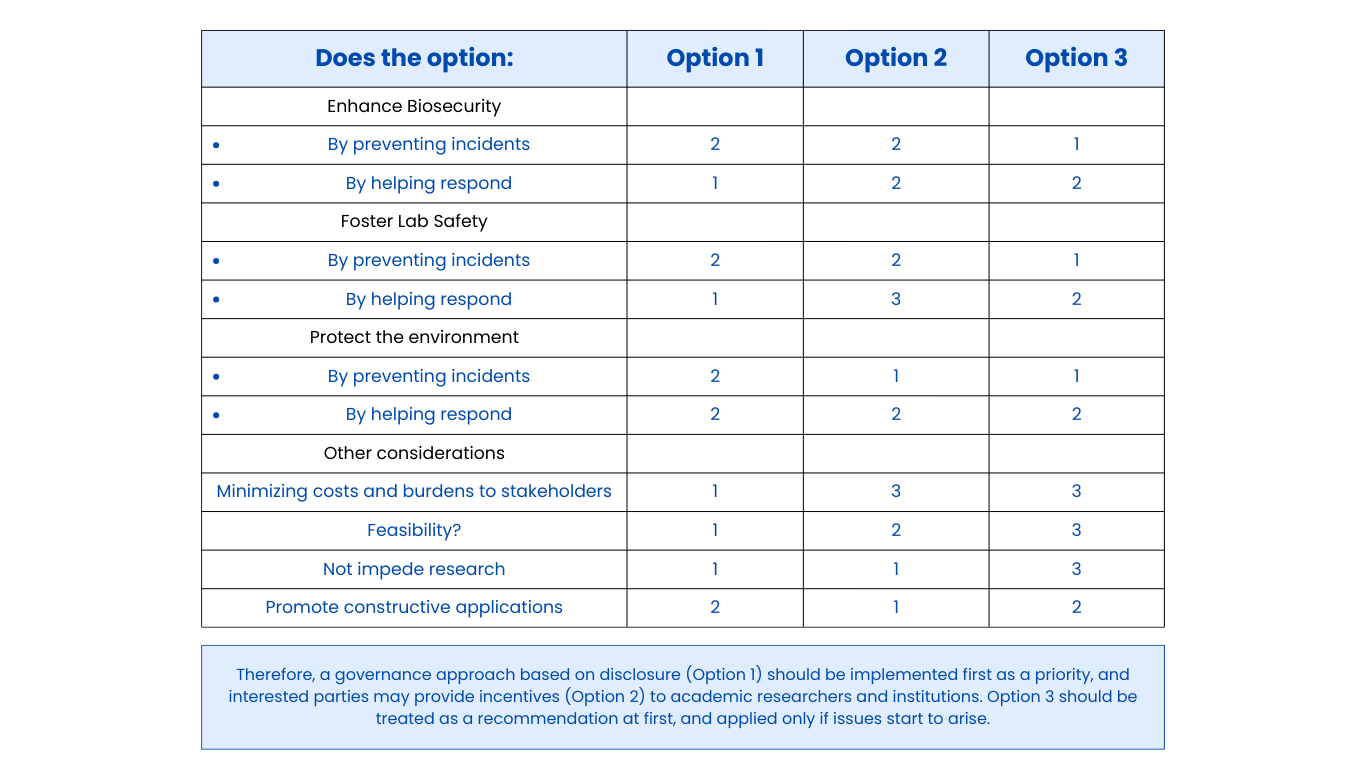
Class Assignment First, describe a biological engineering application or tool you want to develop and why. Next, describe one or more governance/policy goals related to ensuring that this application or tool contributes to an “ethical” future, like ensuring non-malfeasance (preventing harm). Break big goals down into two or more specific sub-goals. Next, describe at least three different potential governance “actions” by considering the four aspects below (Purpose, Design, Assumptions, Risks of Failure & “Success”). Next, score (from 1-3 with, 1 as the best, or n/a) each of your governance actions against your rubric of policy goals. Last, drawing upon this scoring, describe which governance option, or combination of options, you would prioritize, and why. Outline any trade-offs you considered as well as assumptions and uncertainties. Week 2 Lecture Prep Homework Questions from Professor Jacobson: Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy? A: The error rate of polymerase is 1:106. The human genome is approximately 3 billion base pairs long. DNA Polymerases have a proofreading mechanism, and remaining errors can also be fixed through mismatch repair after DNA synthesis.








A: The error rate of polymerase is 1:106. The human genome is approximately 3 billion base pairs long. DNA Polymerases have a proofreading mechanism, and remaining errors can also be fixed through mismatch repair after DNA synthesis.
A: The average size of a human protein is 1036 bp ≈ 345 amino acids, and if we consider that the codon size is 3, the possibility would be 3345, which is approximately 10164. Some codes don’t work due to phenomena like codon bias, degeneracy, and even the GC content.
A: The most commonly used method for oligo synthesis currently is phosphoramidite DNA synthesis.
A: With each step, the yield decreases, meaning that the resulting product will have only a small ratio of the intended oligo. Even though the error rate is not very high, incorrect sequences will accumulate.
A: Similar to the reasons described above, yield drops, and synthesis of long genes becomes expensive. The risks of errors increase with each step. Additionally, longer sequences are more prone to forming secondary structures.
A: The 10 essential amino acids are isoleucine, leucine, lysine, threonine, tryptophan, methionine, histidine, valine, and phenylalanine. “Lysine Contingency” is a concept from the Jurassic Park, in which the scientist introduced a genetic modification to prevent the dinosaurs from synthesising lysine, thus making them reliant on lysine supplements. However, dinosaurs would survive by consuming food that contains lysine.
Background and Rationale T-cell activation is initiated when the T-cell receptor (TCR) and associated CD3 complex form nanoscale clusters at the cell membrane. Although anti-CD3 antibodies are widely used to activate T cells and redirect them in cancer therapy, conventional antibodies poorly control cluster size, geometry, spacing, and signaling strength because they are large, bivalent, and structurally inflexible. Nanobodies (VHHs) or scFvs offer a strong alternative. This project uses anti-CD3 nanobodies as programmable building blocks to precisely control TCR clustering and define how receptor geometry determines T-cell activation.
Hypothesis The magnitude and quality of T-cell activation are governed not only by CD3 binding affinity, but by the valency, spacing, and geometry of clustered TCR-CD3 complexes. Engineered multivalent nanobody scaffolds can tune these parameters and generate predictable signaling outputs.
Research Objective Engineer anti-CD3 nanobody-based proteins with defined valency and spacing to identify the structural rules of productive TCR clustering and apply them to safer T-cell activation platforms.
Expected Outcomes This project is expected to establish: