Week 1 HW: Principles and Practices
Part 1: Yeast BioShields: Engineering Yeast for Wound Protection
The global problem of multi-drug-resistant bacteria (MDR) has been a widespread issue for many years now. The World Health Organization has declared this phenomenon as a major global health threat, further emphasizing the need for a solution to this problem (Renata Urban-Chmiel et al, 2022). In the pursuit of this solution, I structured my idea around a way to reduce the threat of antibiotic-resistant bacteria.
My first proposal was to use GRAS bacteria as a chassis, and modify them to be receptive to the pheromones of infectious bacteria and recognize them as competitors. These modified bacteria could then be cultured and used as a natural shield for the human wounds against infections. This idea was based around the knowledge that bacteria possess a diverse range of mechanisms for inhibiting competitors, including bacteriocins, tailocins, type VI secretion systems and contact-dependent inhibition (CDI) (Booth, S.C., Smith, W.P.J. & Foster et al, 2023).


Figure 1. The dynamics of non-engineered bacteriocin-producing bacteria are complex, guaranteeing the ecological co-existence of antibiotic-resistant and antibiotic-sensitive strains. I believe that, by using yeasts as a naturally immune producer, this ecological balance may be turned in favor of eliminating resistant strains. From Booth, S.C., Smith, W.P.J. & Foster et al, 2023.
But I shifted my focus from using bacteria, be as cheap and fast to grow as they may be, to yeasts, as the former lack the ability to create more complex peptides and are endangered by the antimicrobial peptides (AMPs) they produce themselves. I plan to focus on using genetically engineered yeast within a patients gut or near an infection-vulnerable wound (such as that of a surgery, specially long ones) to eliminate antibiotic resistant bacteria.
While their growth is slightly more time-taking and expensive, yeasts hold many advantages in comparison with GRAS bacteria:
- Yeasts, as eukaryotes, have the ability to create complex peptides and antibodies, and contain way less restrictions due to their ability to contain endosymbiotic.
- Unlike bacteria which can, at times, eliminate themselves using their peptides, yeast do not share that weakness. Due to their eukaryotic membranes, yeast are immune to the peptides that can target and destroy bacteria.
- Yeast like Saccharomyces cerevisiae have the potential for huge implementations within the medical field. For example, they can be used as Biosensors, Pharmaceutical cell factories, and to create vaccines (see Figure 2).
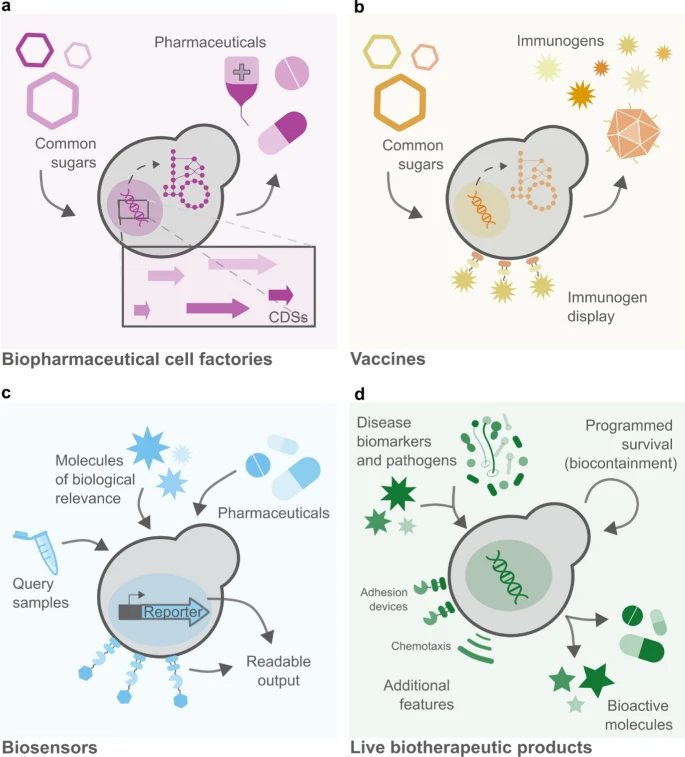

Figure 2. Different applications of the baker’s yeast in medical biotechnology. My idea relates to panel d, demonstrating yeast as a potent biotherapeutic agent. From Maneira et al., 2025.
Part 2: Governance Goals and Sub-goals
Genetically engineered yeast can be a promising boon towards the struggle against MDRs, they can also be health risk themselves if not used carefully. For example, there has been reports of a case of bloodstream co-infection of Saccharomyces cerevisiae and Candida glabrata while using micafungin (Furuya K, Ito K, Sugiyama K, Tokuda S, Kanemoto H, Kamei K, Shimada T et al, 2023), which can cause complications regarding the use of yeast near open wounds. So, several policies are needed in order to ensure that yeast remain harmless towards patients.
- Ensuring biosafety (aligns with non-maleficence): The safety of using the yeast must be guaranteed in terms of its biological functions. i. Rendering the yeast incapable of causing harm to patients. ii. Ensuring its containment and inability to spread outside its specified area of use.
- Accessibility (aligns with equity) Keeping the product affordable and accessible. i. Lowering the cost of making the final yeast-based product through bioprocess optimization. ii. Improving production capacity and upscaling the process.
Part 3: Governance Actions
For my final project and according to the aforementioned sub-goals, I suggest the following actions: A) Implementing a genetic kill switch: Actors: Researchers, Academia.
By creating a designed kill switch within the yeast, we can design the yeast to be neutralized in the case of it entering the patient’s blood.
- Purpose: This ensures that the yeast cannot harm the patient by potentially infecting their blood.
- Design: We can use Rim101 pathway in yeast (Mira et al., 2009) and couple its responsive promoter to a suicide gene.
- Assumptions: The kill switch could not activate due to some evolutionary factors, or the yeast becoming resistant to it.
- Risks: Infections caused by the kill switch not working as intended.
B) Optimization of growth methods for the yeast: Actors: Researchers, Medical practitioners.
This action reduces the cost of production by increasing the yield of the yeast, using aerobic conditions and fed-batch cultivation to avoid the Crabtree effect (Gregory J. O. Martin and Sitha Chan et al, 2024).
- Purpose: Lowering the cost of yeast growth and increasing mass yield.
- Design: To avoid the Crabtree effect and resulting ethanol production, fed-batch cultivation is used in which low concentrations of glucose are maintained by controlled feeding of substrate during the cultivation to provide the necessary flow of carbon.
- Assumptions: The biologically engineered sample may need different conditions in order to increase yield or grow at all.
- Risk: Hindering the process of growth through wrong methods.
- Securing governmental or private funds: Actors: Governments, Industries, investors.
In order to fund the bioprocess, we can encourage investors and academic institutes. This action can ensure the resources needed to fully develop the product.
- Purpose: Securing the needed investment in order to run project.
- Design: Writing proposals, pitching to investors and applying for governmental and academic fundings.
- Assumptions: The investors and companies could see the project as too risky or non-profitable.
- Risks: The bioprocess being met with a shortage of needed equipment and resources.
Part 4: Scoring Rubric
| Does the option: | Action 1: Implementing a genetic kill switch | Action 2: Optimization of growth methods for the yeast | Action 3: Securing governmental or private funds |
|---|---|---|---|
| Ensuring biosafety | |||
| • By rendering the yeast incapable of causing harm to patients? | 2 | 0 | 0 |
| • By ensuring its containment and inability to spread outside its specified area of use? | 3 | 0 | 0 |
| Keeping the product affordable and accessible | |||
| • By lowering the cost of making the final yeast-based product through bioprocess optimization? | 0 | 3 | 1 |
| • By improving production capacity and upscaling the process? | 0 | 2 | 3 |
| Other considerations | |||
| • Minimizing costs and burdens to stakeholders | 1 | 3 | 2 |
| • Feasibility? | 2 | 3 | 3 |
Part 5: Prioritization and Trade-offs
During my research I have made some assumptions:
- That the use of yeast as a way of combating antibiotic resistant bacteria and preventing their infection is a more effective method than others of its kind.
- That the genetically modified yeast is cost-effective compared to other ways of treatment for this area. According to the fact that insuring biosafety is crucial, and that the project could not develop as fast without proper funding, I prioritized action 1 over 2 and 3, and action 3 over 2. While mass production methods and reducing the cost of growing the yeast could have a considerable impact in upscaling the process, I believe it to be an acceptable trade off for the short- and mid-term.
Part 6: Pre-lecture Questions for Week 2
6.1: Prof. Jacobson’s Questions
- Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy?
Answer: Even though different eukaryotic and prokaryotic DNA polymerases have different error rates, which is also subject to the environment (Balint et al., 2024) in which they function, the apporixmate error rate of all polymerases is about 1 mutation per 10,000 bases replicated. Considering the fact that the human genome is roughly 3.2 billion base pairs long, after each round of replication, one should expect to see 320,000 mutations, which is far from what is the case in the real world. This discrepancy is explained by the active role of polymerase proof-reading and error correction systems active during and after DNA is replicated by DNA polymerase.
- How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice what are some of the reasons that all of these different codes don’t work to code for the protein of interest?
Answer: There are 64 different codons that make up the genetic code of all living organisms, of which 61 combinations are used to code amino acids and 3 are used to stop translation. The main reason behind the existence of more than 1 codon for each of the amino acids goes back to the previous question. Even with error correction in place, there may be some mutations that pop up after each round of replication. Therefore, having more than 1 codon for certain amino acids helps preserve the function and amino acid composition of the proteins mutated genes code for.
6.2: Dr. LeProust’s Questions
- What’s the most commonly used method for oligo synthesis currently?
Answer: The typical method for synthesizing DNA (chemically) is phosphoramidite synthesis (Hoose et al., 2023). As far as I could understand it, this method involves loading modified bases on a solid silicon platform, removing a protecting group from them, and binding them to another base with the protecting group still attached. See the figure below for more information.
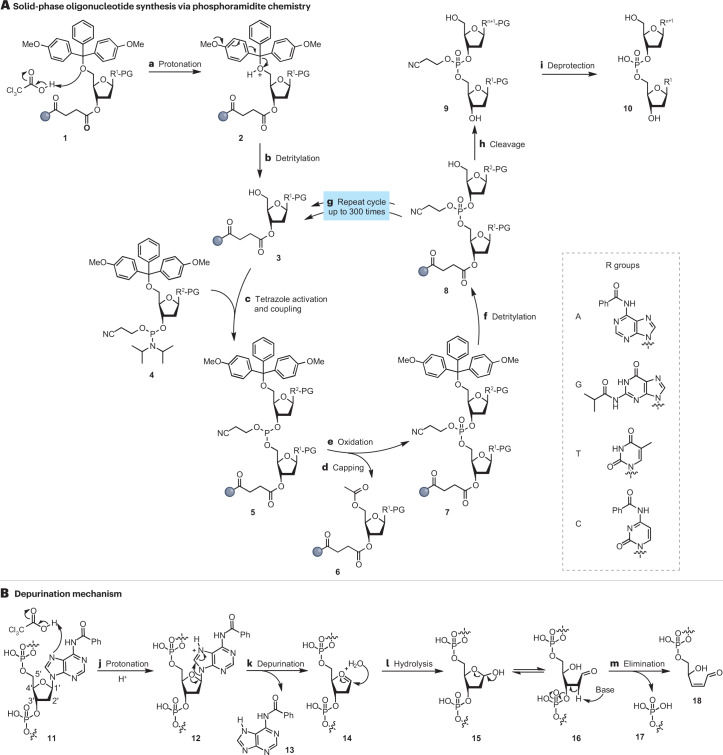

Figure 3. The phosphoramidite synthesis method. Steps h and i are important for the answer to the next question.
- Why is it difficult to make oligos longer than 200nt via direct synthesis?
Answer: As it can be observed in the Figure 3, sometimes the added bases do not stay in the oligo synthesized and separate from it as an unwanted byproduct. The article introduces this as acid-catalyzed depurination, which is the reason behind the inefficiency of direct synthesis for breaking the 200nt cap.
- Why can’t you make a 2000bp gene via direct oligo synthesis?
Answer: As it was implicitly mentioned in the previous question, direct oligo synthesis using the phosphoramidite method has ‘imperfect’ cycles, meaning that at each cycle there is a chance for one or more nucleotides to break apart from the growing oligo and hinder its growth. This problem, which becomes more serious and ultimately breaks and stops oligo synthesis as the oligo grows longer, disables us from creating a 2000bp gene by direct synthesis.
Hoose et al., 2023 instead names two other methods that can be used to assemble synthesized DNA molecules like LEGO bricks: Gibson Assembly and Polymerase Cycling Assembly (see Figure 4).
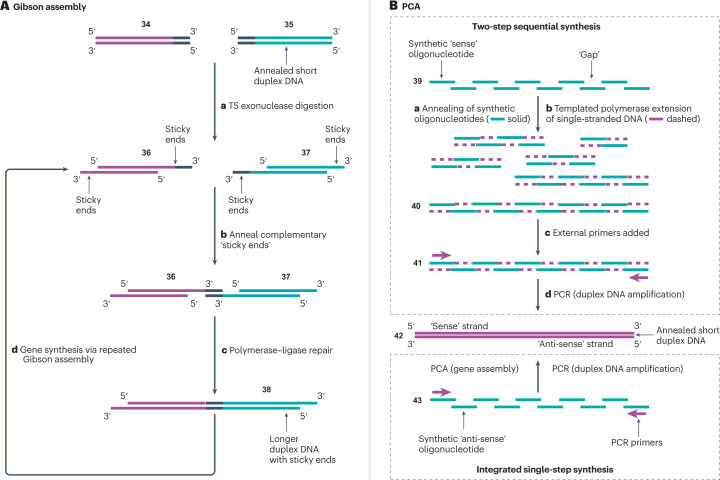

Figure 4. Two DNA assembly methods.
6.3: Prof. George Church’s Questions
- What are the 10 essential amino acids in all animals and how does this affect your view of the “Lysine Contingency”?
Answer: Cysteine, Histidine, Isoleucine, Leucine, Lysine, Methionine, Phenylalanine, Threonine, Tryptophan, Tyrosine, and Valine are considered essential amino acids for all animals (Hou et al, 2018).
Interestingly, the concept behind ‘Lysine contingency’ is similar to auxotrophy, a situation where an organism lacks the ability to make one or more compounds necessary for its growth and sustenance (usually with reference to its wild-type relatives). However, since Lysine is naturally an essential amino acid and not produced by the animal itself, the ‘Lysine contingency’ is more fiction than (feasible) reality.
Acknowledgement: Use of AI
I acknowledge the use of AI, particularly Gemini, for early fact checking of my raw ideas. “One thing that I was curious about: Can yeast cells produce and secrete more potent and a more diverse array of engineered/heterologous antimicrobial peptides (AMPs) than probiotic bacterial cells creating bacteriocin and other types of antimicrobial factors or AMPs?”