Week 2 HW: DNA Read Write and Edit
Part 1: Benchling & In-silico Gel Art:
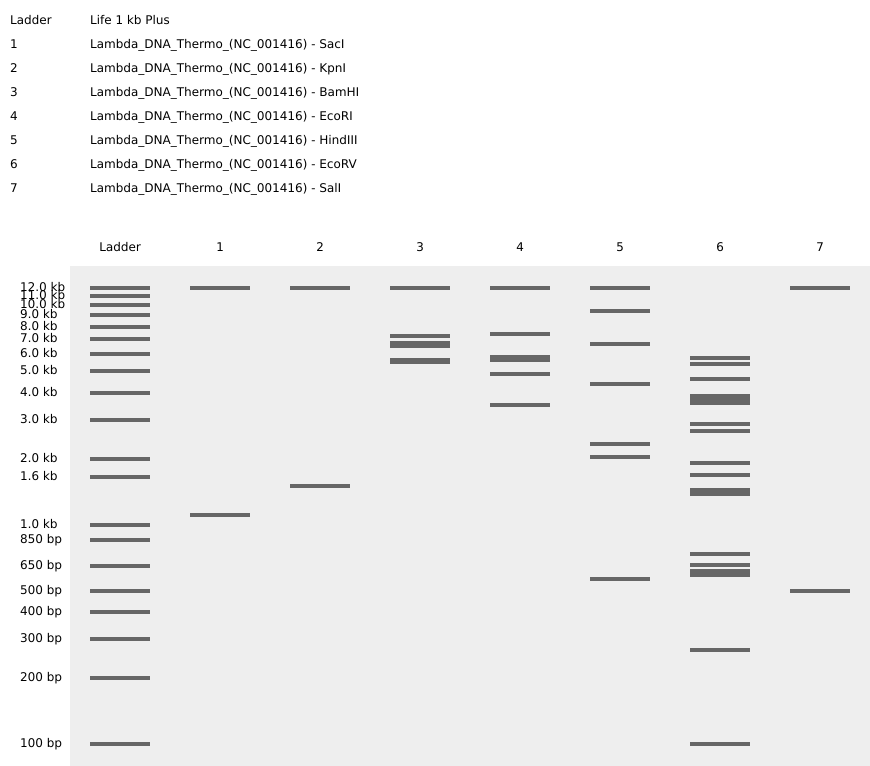
- Simulate Restriction Enzyme Digestion with the following Enzymes:
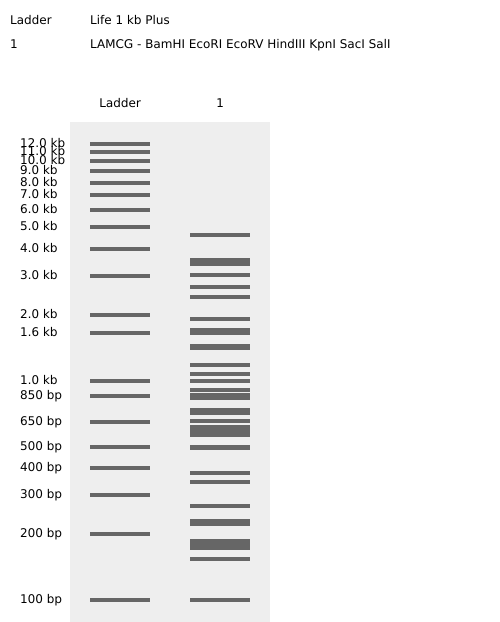

I tried to make a pattern that looked like a staircase going down by using logic but couldn’t quite seem to get it right…
Part 3: DNA Design Challenge:
Protein Choice: AFP III (Antifreeze Protein Type III) I chose AFP III because it has an incredible practical application: allowing organisms to survive in sub-zero temperatures. In biotechnology, it is used to create smoother ice cream textures (by preventing large ice crystals) and in the preservation of organs for transplant. It is a fascinating example of how nature solves extreme physical challenges through molecular engineering.
Protein Sequence (UniProt P05140):
sp|P05140|ANP3_MACAM Antifreeze protein type 3 NQASVVANQLIPINTALTLVMMKAEVVTPMGIPAEEIPNLVGMQVNRAVPLGTTLMPDMV KNY
Reverse Translation
aatcaagcctctgtagtagccaaccagctgatccccatcaacaccgccctcaccctggtcatgatgaaggccgaggtcgtcacccccatgggcatccccgccgaggagatccccaacctggtcggcatgcaggtcaaccgcgccgtccccctcggcaccaccctgatgcccgacatggtcaagaactac
Codon Optimization
In your own words, describe why you need to optimize codon usage. Which organism have you chosen to optimize the codon sequence for and why?
This answer takes back to last homework, due to codon bias, using a sequence that’s not ideal for your chosen organism, you may have some problems with codons used tipically by the species that produces said protein naturally. I picked E.Coli because it is the standart for easy cultivation.
I ran VectorBuilder’s tool with some uncertainty but it seemed good because it was the only one that said ‘free’ on the google search.
This is the optimized sequence:
AACCAGGCGAGCGTGGTGGCGAATCAGCTGATTCCGATTAATACCGCCCTGACCCTGGTGATGATGAAAGCCGAAGTGGTGACCCCGATGGGCATTCCGGCGGAAGAAATTCCGAACCTGGTGGGCATGCAGGTGAATCGCGCGGTGCCGCTGGGCACCACCCTGATGCCGGATATGGTGAAAAATTAT
You have a sequence, now what? Cell-Dependent Expression (In Vivo): The most common method is to transform E. coli cells with the plasmid. When an inducer (like IPTG) is added to the growth media, the bacteria’s internal machinery (RNA Polymerase and Ribosomes) begins to transcribe the DNA into mRNA and translate that mRNA into the physical AFP III protein.
Cell-Free Protein Synthesis (In Vitro): Alternatively, one could use a “cell-free” extract. This involves mixing the DNA directly with a “molecular soup” containing ribosomes, enzymes, and amino acids in a test tube.
Part 4: Prepare a Twist DNA Synthesis Order:
I made an account and twist doesn’t seem to allow me to use the website, saying I need to contact a distributor…
I checked with a fellow brazilian colleague and he seems to be having the same issue, maybe it’s a regional block?

Part 5: DNA R/W/E:
In graduation I learned a ton about mangroves and their importance, so, to sequence the mangrove microbiome, I’d target the DNA in the sediment and root zones of these coastal forests. Mangroves are massive “blue carbon” sinks, and by reading the genetic material of the microbes living there, we can find hidden enzymes that fix nutrients or even break down plastics in salty water. This helps us understand how these forests fight climate change and gives us new “bio-tools” for cleaning up the environment. I would use Illumina Sequencing for this because it’s the best for handling the complex mix of species found in soil. This is a second-generation technology that uses “sequencing by synthesis” to read millions of DNA fragments at once. First, I’d extract the DNA, break it into tiny pieces, and attach “adapters” so they can stick to a glass flow cell. Inside the machine, the DNA is copied into clusters, and as fluorescently labeled bases are added, they flash a specific color for each letter (A,T,C, or G). A camera captures these flashes, and the software translates them into digital FASTQ files that we can piece together like a giant puzzle.
For the “DNA Write” part, I’d synthesize a caffeine biosynthetic pathway to put into yeast. This would let us “brew” caffeine in a lab without needing huge coffee or tea plantations, saving a lot of land and water. I’d use silicon-based synthesis (like Twist Bioscience), which uses a silicon chip to build thousands of DNA strands at once using a chemical process called phosphoramidite chemistry. We’d print short pieces of the caffeine genes and then stitch them together into a full circuit. The main catch is that it’s hard to write very long or repetitive sequences, but it’s incredibly fast and scalable for these kinds of metabolic projects.
Finally, I’d use CRISPR-Cas9 to edit a banana or tomato so it grows with the caffeine pathway built right in. The idea is to create a “Caffeine Fruit” for a natural, healthy morning energy boost. CRISPR works like a molecular GPS and scissors; a guide RNA leads the Cas9 enzyme to a specific spot in the plant’s DNA to make a cut. By providing a “donor template” with our caffeine genes, the plant’s own repair system pastes the new instructions into its genome. We’d use a “shuttle” like Agrobacterium to get the CRISPR tools into the plant cells. It’s a powerful method, though it can sometimes be slow to grow a full plant from an edited cell, and we have to be careful that the enzyme doesn’t cut the DNA in the wrong place.