Week 6 HW: Genetic Circuits Part I: Assembly Technologies
Assignment: DNA Assembly
1. What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
Phusion High-Fidelity PCR Master Mix Components:
Phusion DNA Polymerase: A high-accuracy enzyme with 3′→5′ exonuclease (proofreading) activity that minimizes mutations during amplification dNTPs (Deoxynucleotide Triphosphates): The nucleotide building blocks (A, T, C, G) used to synthesize the new DNA strand.
Mg2+ (Magnesium Ions): A critical cofactor for polymerase activity. It stabilizes the interaction between the enzyme and the DNA.
Buffer System: Maintains optimal pH and ionic strength to ensure enzyme stability and efficient primer binding.
2. What are some factors that determine primer annealing temperature during PCR?
Melting Temperature (Tm), GC content, Lenght of the primer, and Salt concentration.
3. There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
Protocol: PCR uses thermal cycling (heating and cooling) to amplify specific sequences exponentially. Restriction digests involve incubating DNA with specific enzymes at a constant temperature (usually 37∘C) to “cut” the DNA at recognition sites. Comparison: PCR is preferable when you need to amplify a specific gene from a small amount of template or add custom “overhangs” for assembly. Restriction digests are preferable for verifying plasmid maps (diagnostic digests) or using “sticky-end” cloning when appropriate sites are already present.
4. How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
To ensure success, you must design your fragments with overlapping homologous ends (usually 20−40 bp long). This is achieved by designing PCR primers where the 5′ end of the primer matches the end of the adjacent fragment. Additionally, you must ensure the fragments are free of template DNA (via DpnI digestion) and that the overlaps do not contain stable secondary structures that could interfere with the exonuclease step of the Gibson reaction.
5. How does the plasmid DNA enter the E. coli cells during transformation?
In chemically competent E. coli, the cells are treated with divalent cations (like Ca2+) to neutralize the negative charges of the DNA and the cell membrane. During Heat Shock (moving from ice to 42∘C), a thermal gradient is created that opens temporary pores or “adhesion zones” in the membrane, allowing the plasmid DNA to be pulled into the cell.
6. Describe another assembly method in detail (such as Golden Gate Assembly)
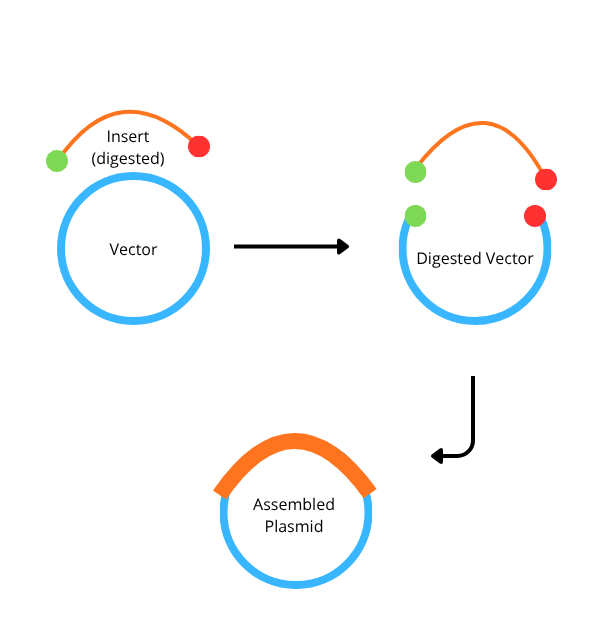
Golden Gate Assembly (GGA) is a molecular cloning method that allows for the simultaneous, one-pot assembly of multiple DNA fragments using Type IIS restriction enzymes and T4 DNA ligase. Unlike traditional restriction enzymes, Type IIS enzymes (such as BsaI) cut at a specific distance away from their non-palindromic recognition sites, creating unique 4-base pair overhangs. These custom overhangs are designed so that the recognition sites are removed from the final ligated product, making the reaction essentially irreversible and driving it toward the desired construct. This “scarless” nature allows for the seamless joining of various biological parts, such as promoters, genes, and terminators, in a predefined order. Because the enzyme and ligase work in a single cycle of temperature shifts, GGA is significantly faster and more efficient than traditional multi-step cloning. Consequently, it has become a standard tool in synthetic biology for building modular systems and large library collections.

Assignment: Asimov Kernel
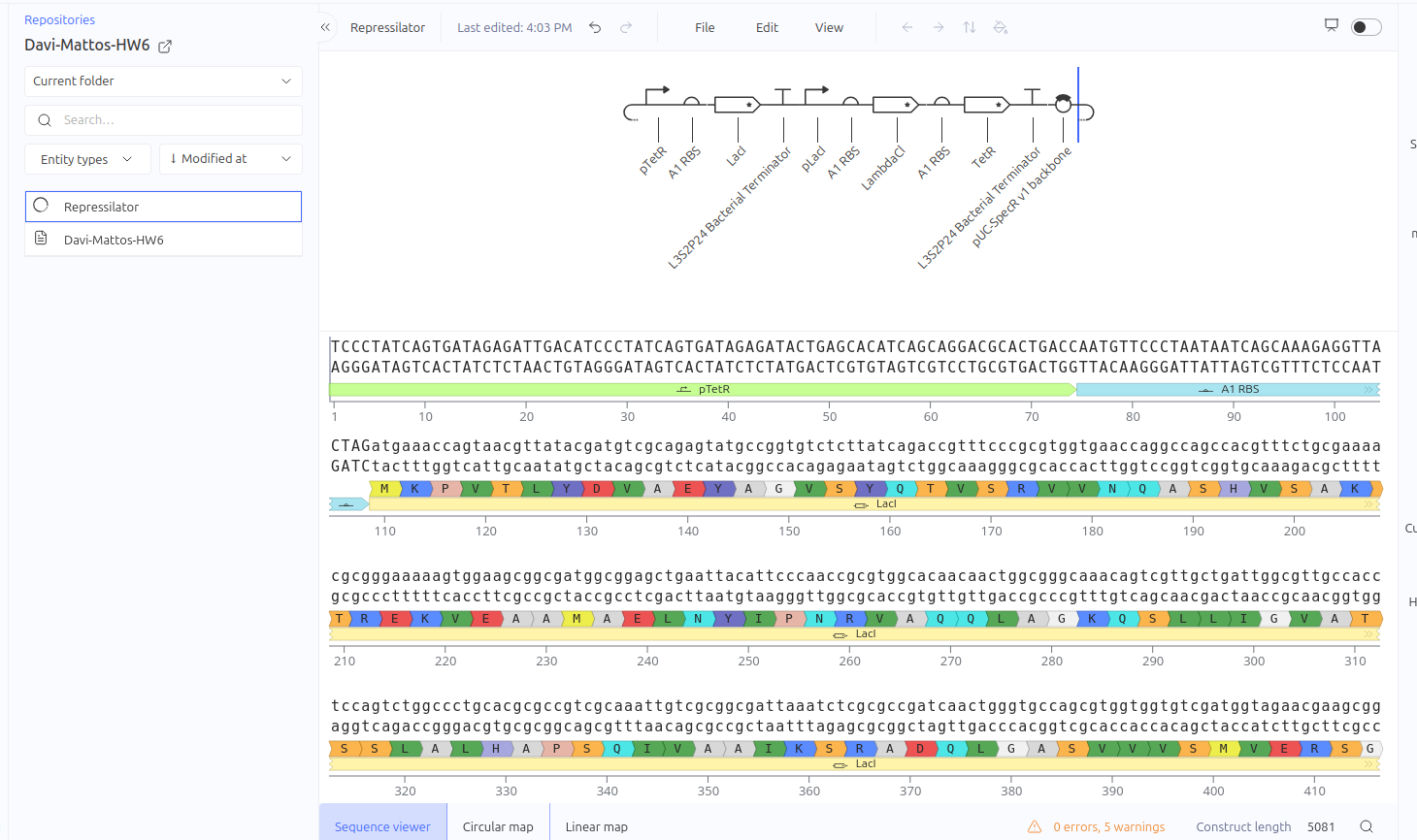
Repressilator
Construct 1: Self-Repressor - Simple Negative Feedback Loop
This circuit was designed as a simple negative feedback loop. The pLacI promoter initiates the transcription of the LacI gene. As the LacI protein is produced and its concentration increases, it binds to its own promoter (pLacI), blocking RNA polymerase and inhibiting its own production. This mechanism is fundamental for cellular homeostasis, allowing the protein to reach a steady state more quickly and in a controlled manner, preventing the waste of cellular resources.
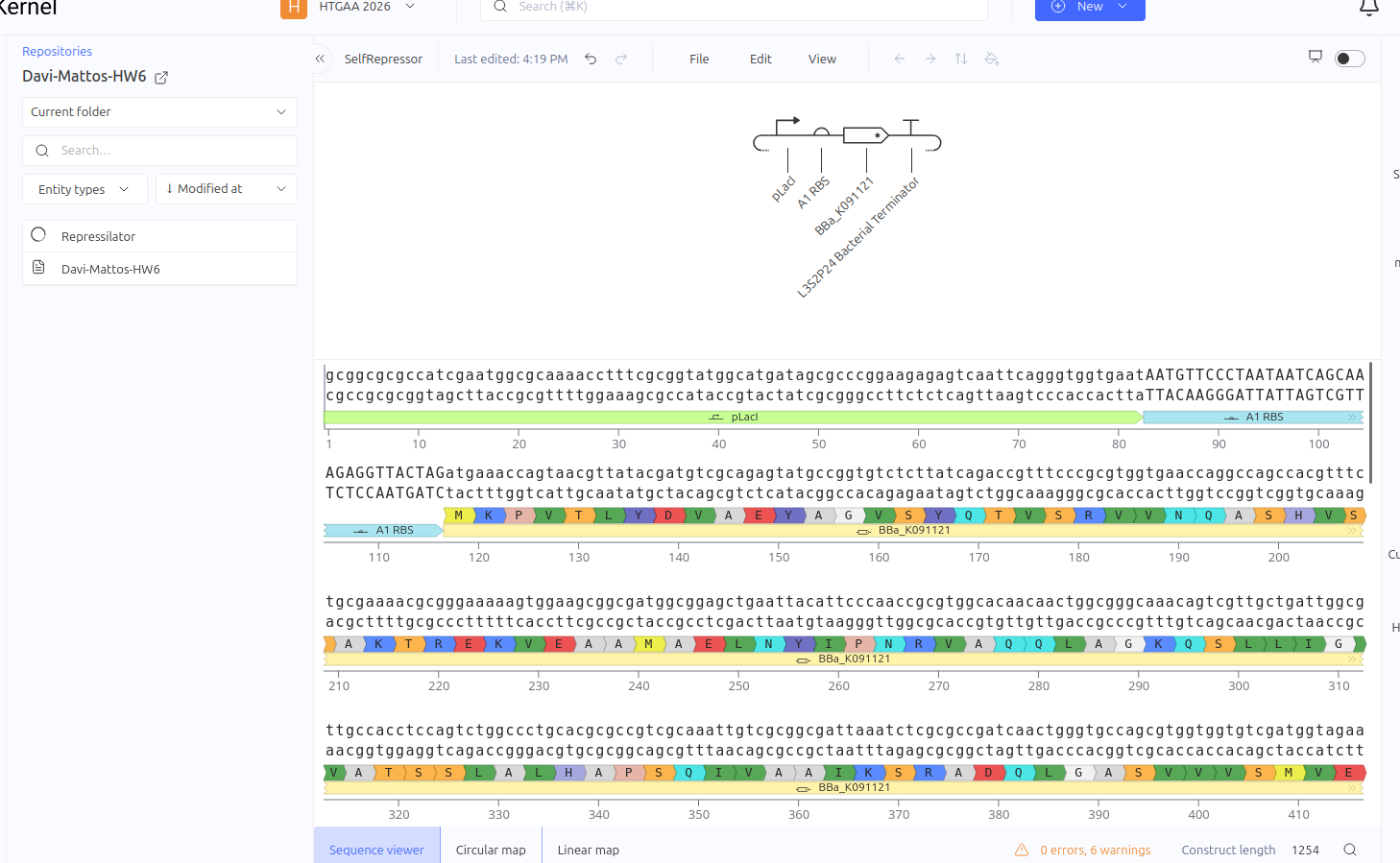
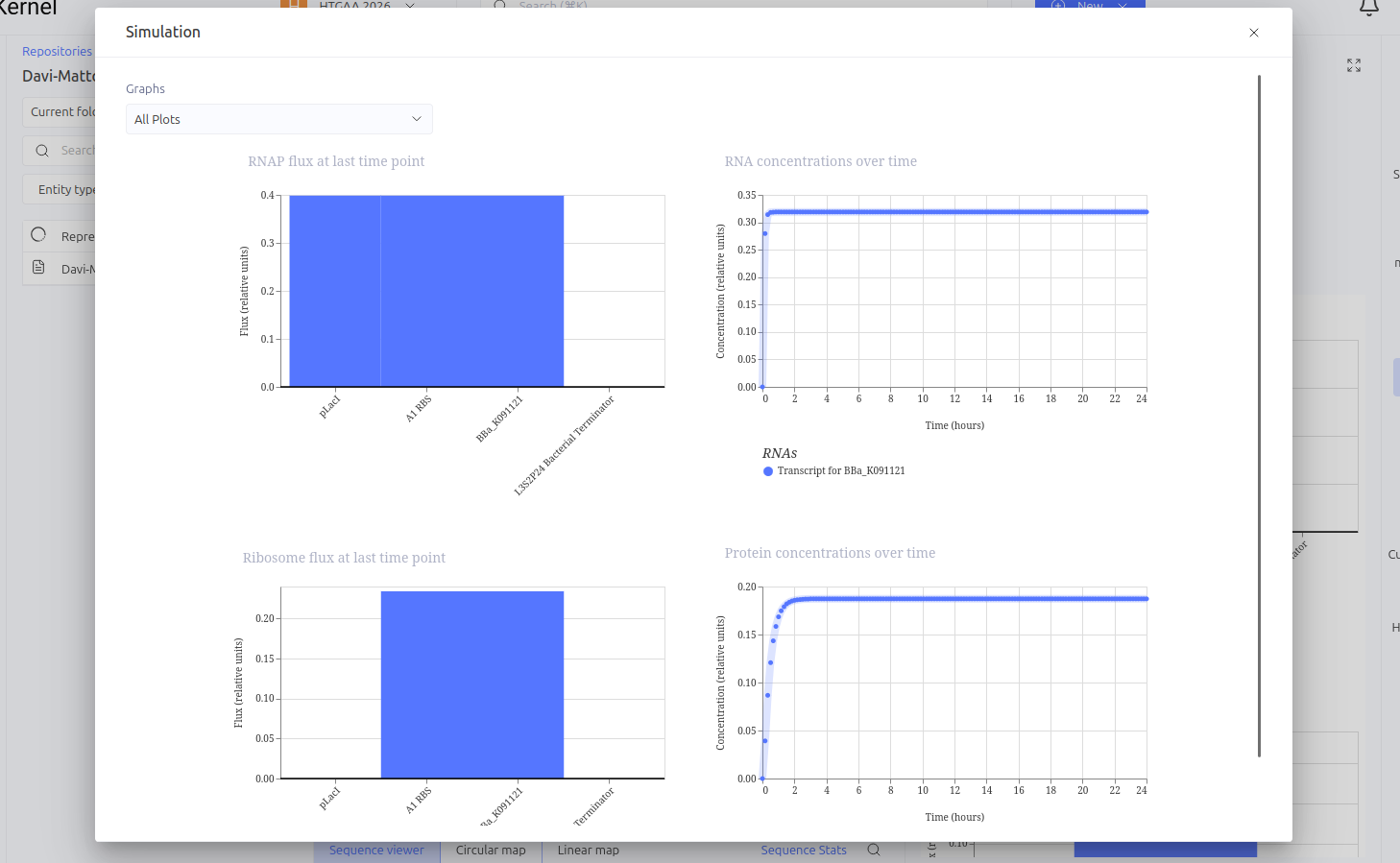
 Expected vs. Observed Results: In the simulator, it was observed that the protein concentration does not grow indefinitely; instead, it reaches a plateau (steady state) within a short time. Initially, a construction error occurred by using two promoters in series, resulting in a lack of translation. After correcting the sequence to Promoter -> RBS -> CDS (LacI), the self-regulating behavior was confirmed by the graphs."
Expected vs. Observed Results: In the simulator, it was observed that the protein concentration does not grow indefinitely; instead, it reaches a plateau (steady state) within a short time. Initially, a construction error occurred by using two promoters in series, resulting in a lack of translation. After correcting the sequence to Promoter -> RBS -> CDS (LacI), the self-regulating behavior was confirmed by the graphs."

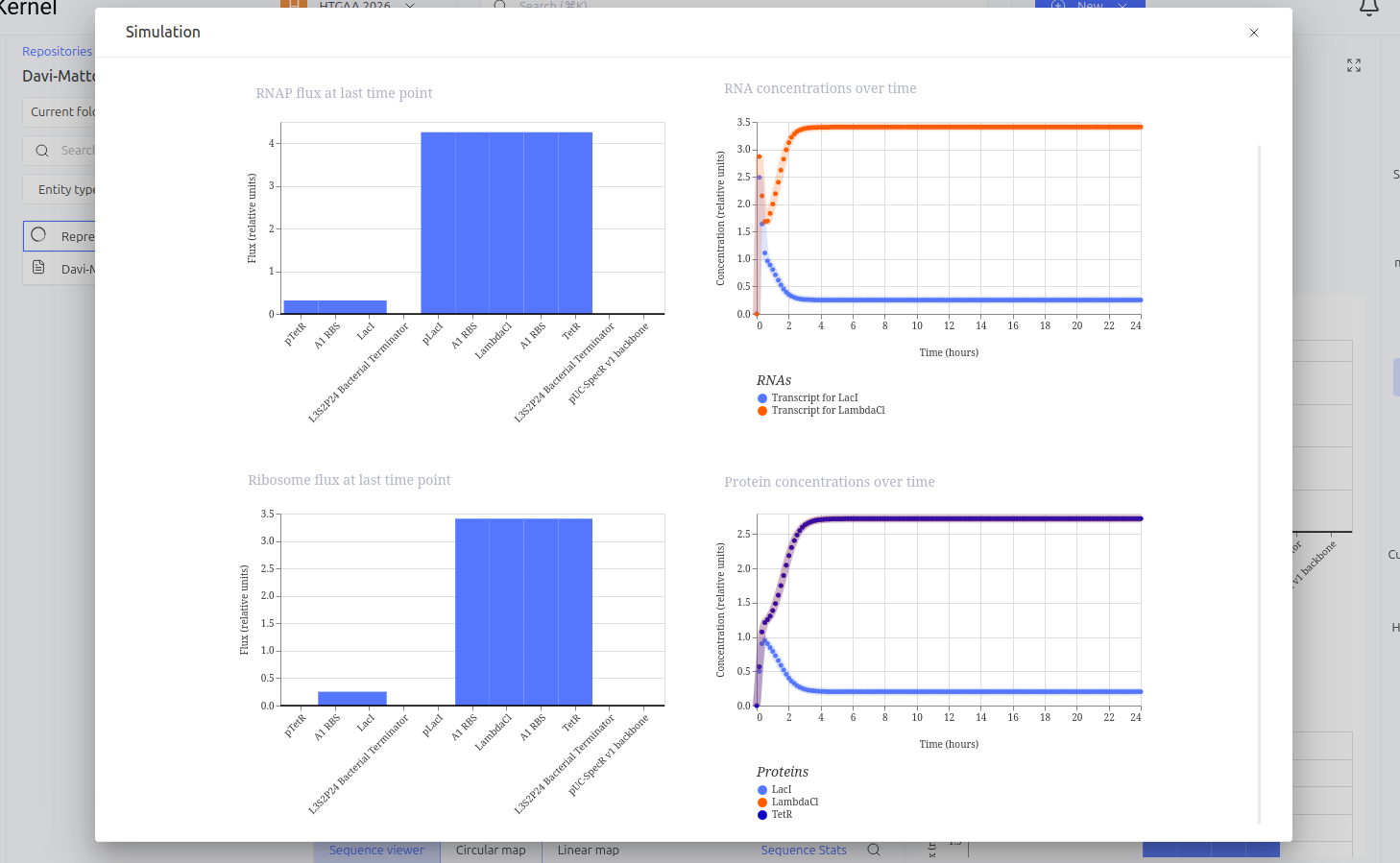
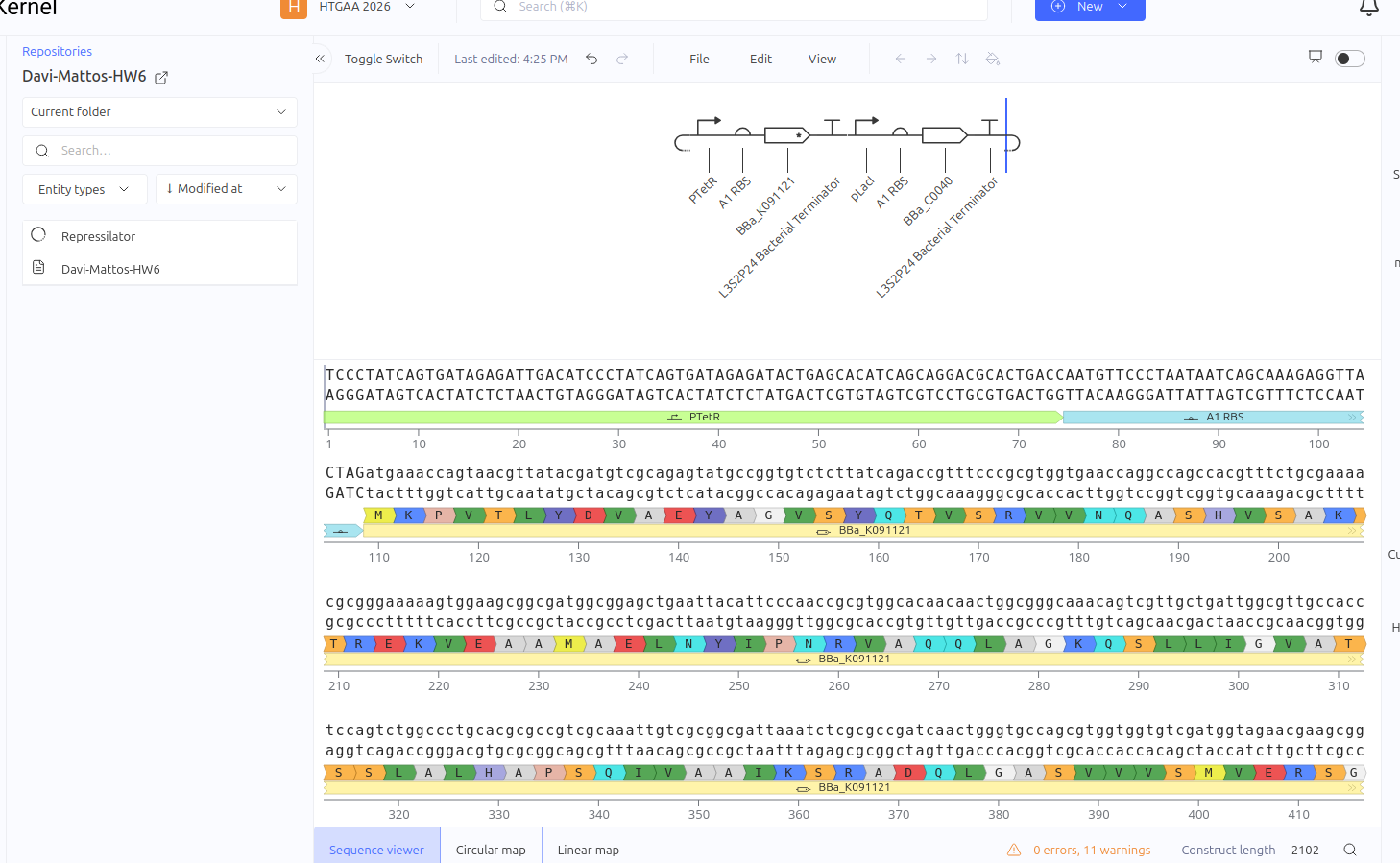
Construct 2: Genetic Toggle Switch - Bistable Switch
The Toggle Switch is a circuit composed of two repressors that mutually inhibit each other’s expression (LacI and TetR). The design consists of two cassettes: the first where the pTetR promoter controls the expression of LacI, and the second where the pLacI promoter controls the expression of TetR. This arrangement creates a bistability system, functioning as a biological memory that can flip between two stable states: (1) High LacI / Low TetR or (2) High TetR / Low LacI.
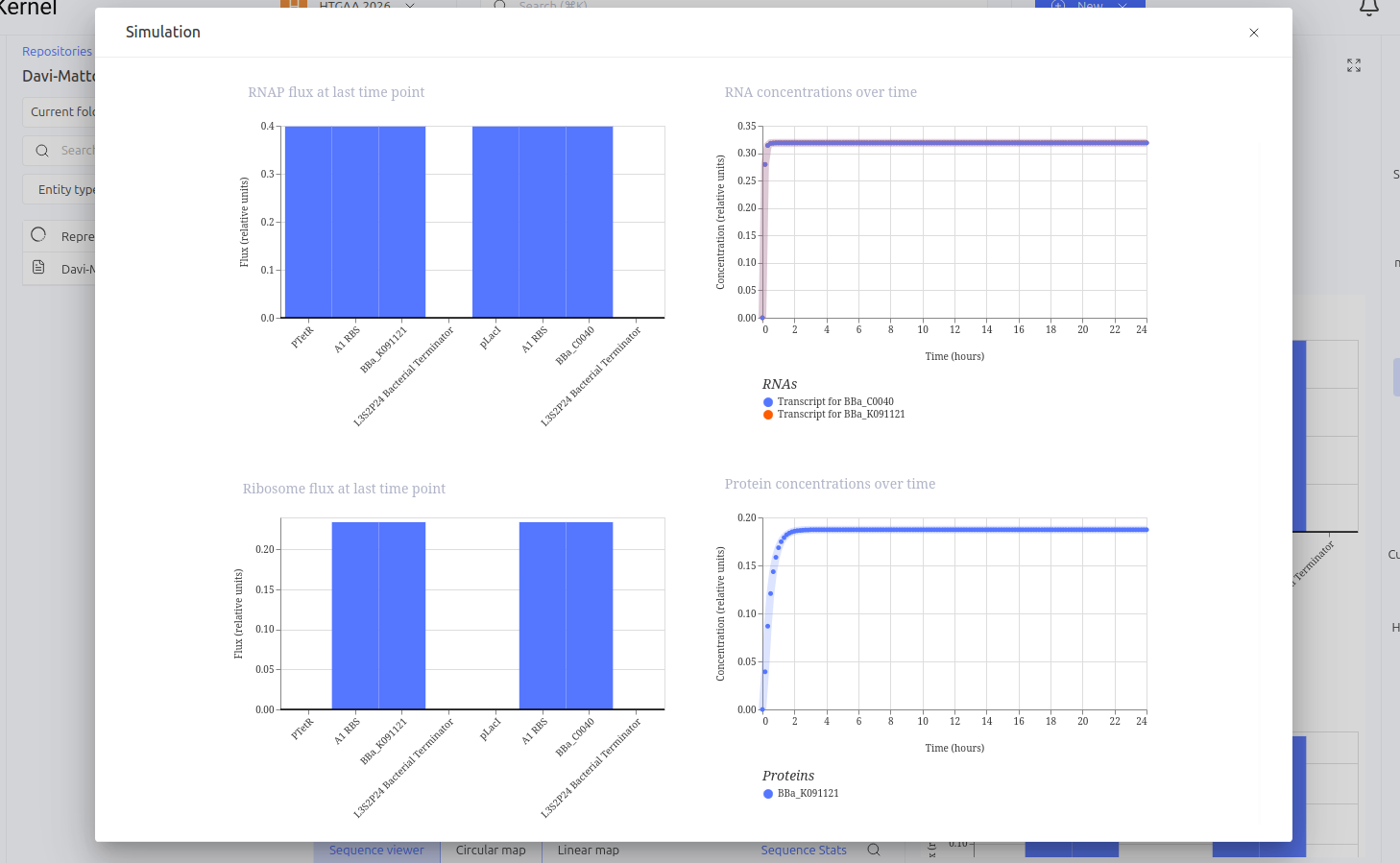
 Expected vs. Observed Results: Simulation results showed the system ‘choosing’ one of the stable states. The LacI protein (BBa_K091121) dominated the system, keeping the expression of TetR (BBa_C0040) repressed and close to zero throughout the simulated time. This demonstrates the circuit’s ability to maintain a stable state even in the absence of external inducers.
Expected vs. Observed Results: Simulation results showed the system ‘choosing’ one of the stable states. The LacI protein (BBa_K091121) dominated the system, keeping the expression of TetR (BBa_C0040) repressed and close to zero throughout the simulated time. This demonstrates the circuit’s ability to maintain a stable state even in the absence of external inducers.

Construct 3: Repression Cascade (Inverter / NOT Gate)
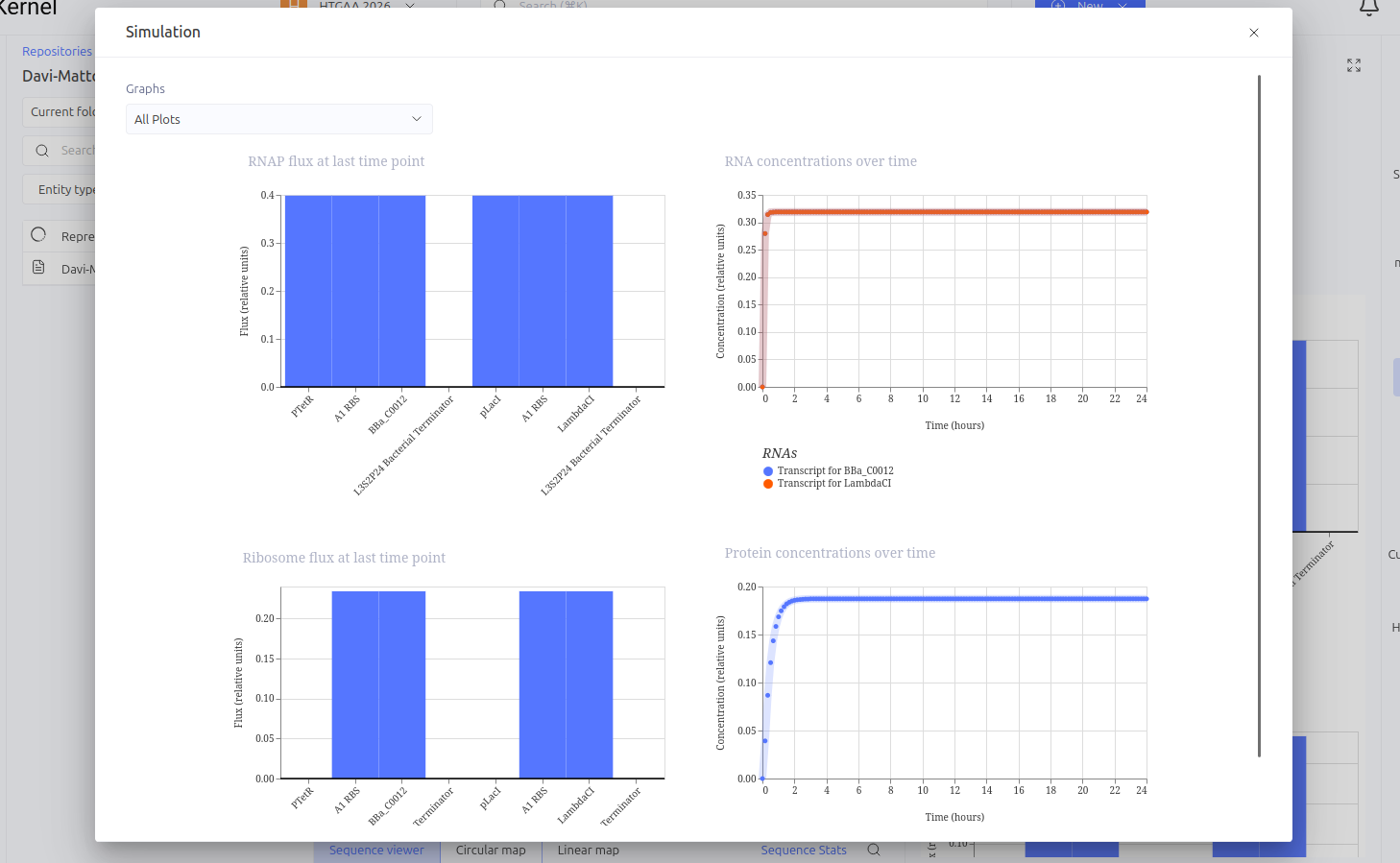
This circuit functions as a linear repression cascade, simulating a NOT logic gate. The first stage uses the pTetR promoter to express the LacI gene constitutively (in the absence of TetR). The second stage contains the LambdaCI gene under the control of the pLacI promoter. The system’s logic dictates that the presence/activation of the first stage necessarily results in the shutdown of the second stage.
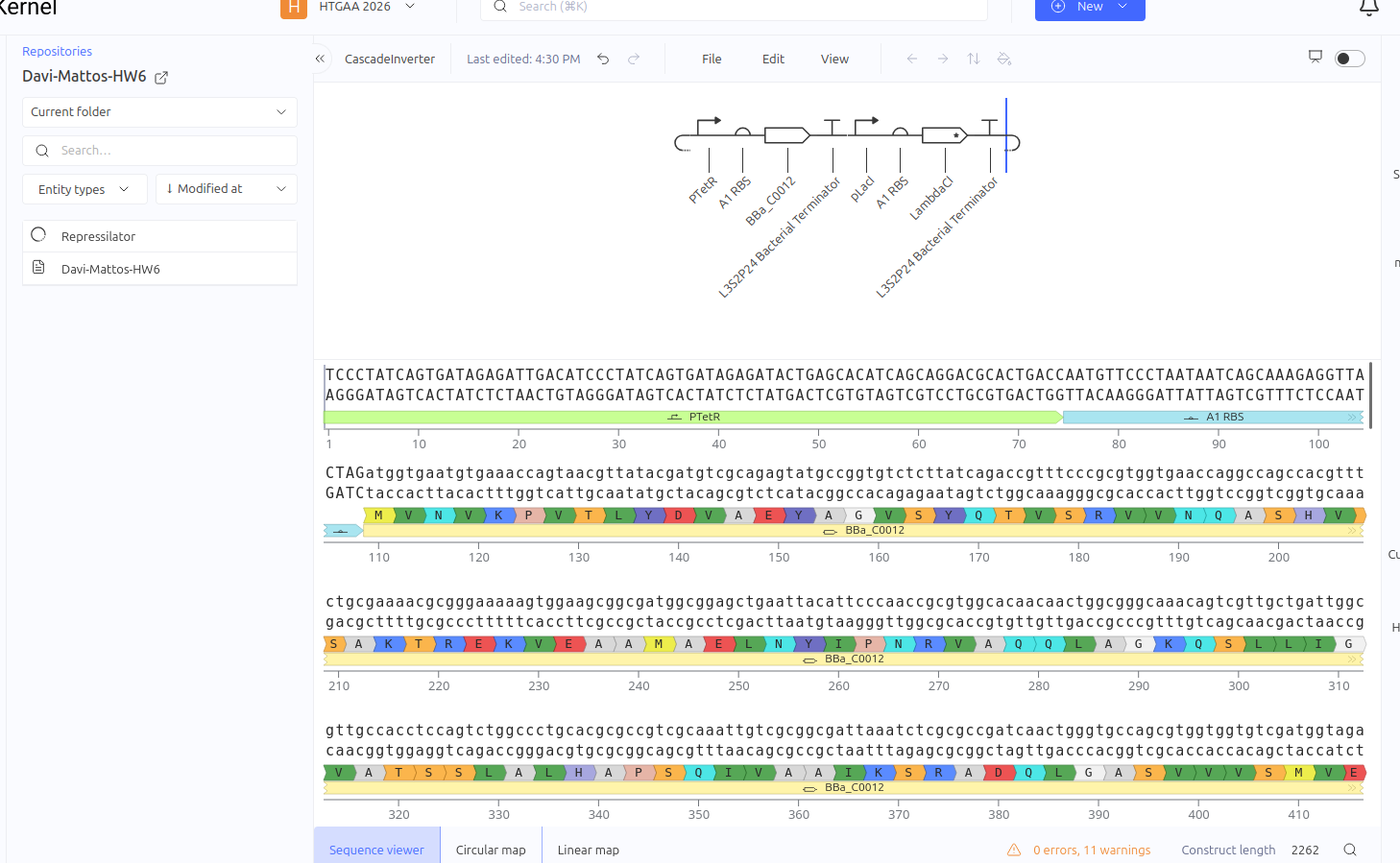
 Expected vs. Observed Results: As expected, the simulator showed high levels of RNA and protein for the first gene (LacI), while the second gene (LambdaCI) showed only a brief initial transient peak of RNA before being completely repressed. The ribosome flux for the second gene dropped drastically, confirming that the biological signal inversion was successful.
Expected vs. Observed Results: As expected, the simulator showed high levels of RNA and protein for the first gene (LacI), while the second gene (LambdaCI) showed only a brief initial transient peak of RNA before being completely repressed. The ribosome flux for the second gene dropped drastically, confirming that the biological signal inversion was successful.