Week 3 HW: Lab Automation
Assignment : Python Script for Opentrons Artwork
I had to write a Python script for a art design. I chose to create a silhouette of the Indian subcontinent, with my city being highlighted. I did that using the Opentrons Artwork website. I thought I will make a pattern of sorts with code but I realized that would time consuming and not very symbolic as such. I got a clipart of India from google and cropped it and then used that too generate my artwork. It didn’t look very good, I had to fiddle around with the contrast, brightness and other values to make it work. It still wasn’t looking how I’d expected it too. I decided to redo it.
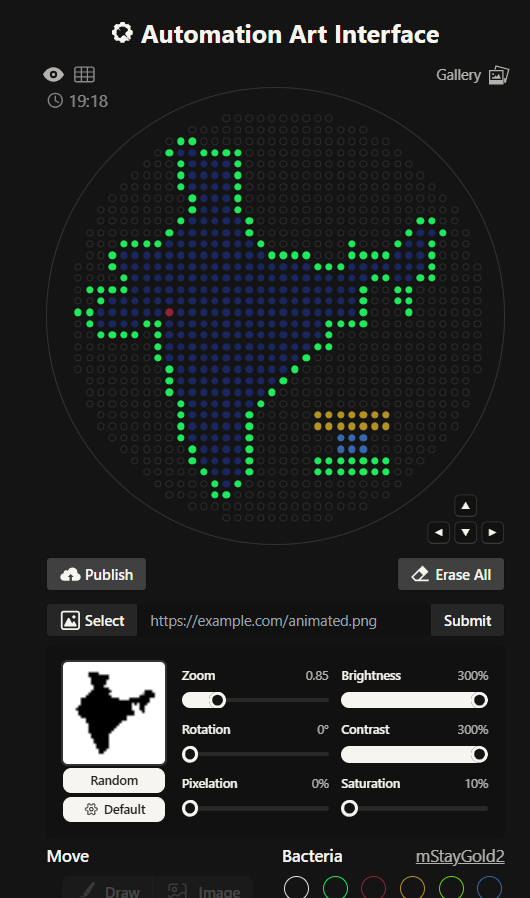

This is the India 2.0 Art (image below), I like this one much better. The green outline in the previous one was not a very good design choice. I don’t know what I was thinking. I created this one by editing my original clipart then striking a balance in the contrast, brightness values. This one look much better in my opinion.
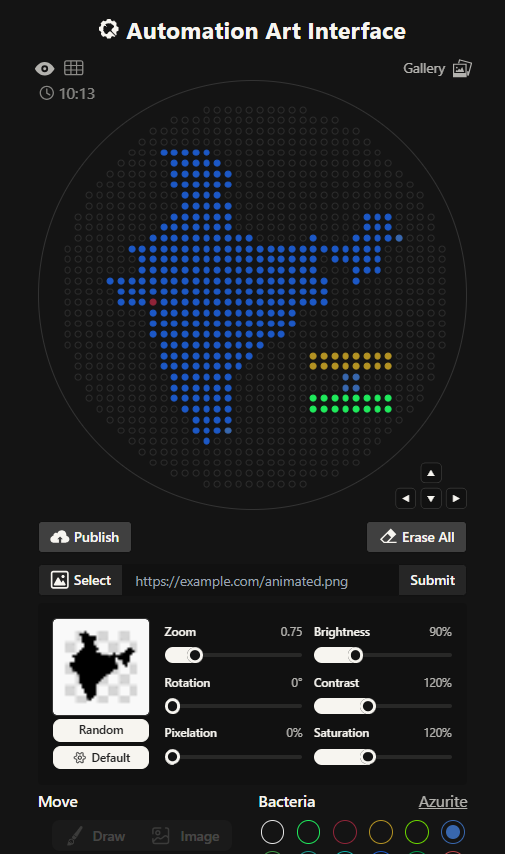



The coordinates were right below my art on the opentrons art website, I just decided to download the script. I edited the script a bit in the PyCharm (adding my name).

Reading further, I found out that I didn’t even need to download my script. I could just publish my design on the opentrons art website and then share the link. I did exactly that and submitted my script to the google form. But in case, Murphy’s Law decides to apply, here’s my code:
https://drive.google.com/file/d/1lLArYDpEx2YWh4lM2WEjRLa5W41kLSq0/view?usp=sharing
Post-Lab Questions
Question 1. Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
I thought that I will need to put in effort to find a paper but the Opentrons website has its own embedded search box thingie to find papers. I just selected the area of my preference and then got a paper I liked.

The paper I chose was titled “Optimizing automated layer‑by‑layer deposition of engineered ECM‑like microenvironments for mammalian cell culture” I chose the paper because last year I’d attended a workshop on 3D Bioprinting and I’d learnt about organoid cultures and replication of the ECM-like conditions for better cell cultures, so I was aware about the what the paper was about. Here’s a link to the paper. https://doi.org/10.1557/s43579-025-00912-9


Question 1 Detour: Reading a Paper and Consolidation.
Back in my graduate coursework, I didn’t have to read a lot of papers but my final dissertation did require me to read a lot of papers. I’d chosen a broad topic “Machine Learning in Life Sciences” There were a lot of papers to be read and a lot of distillation to be done. I used to read papers like header-to-footer but I realized that it was not an optimal approach. The very act of reading 30+ papers for my bachelor’s dissertation led me to evolve, iterate on my paper reading method. So now what I do is this.
- ABSTRACT! I read the abstract to get a basic idea of what the paper is about. If I don’t get it, perhaps because of a lot of jargons. I search up on the jargons, revisit the fundamentals and then read the abstract again. I write it all down (what I understood from the abstract)
- QUESTIONS! I try and see the abstract as a summary of the paper and the expectations I can make, then I skim the paper looking for the references to the abstract. For example, in this case the abstract talked about using heparin and collagen, I skimmed and found out the information related to it, WHY they were using it, WHAT were they creating. I try to connect the components via questions (you can see what I mean in my distillation sheet of sorts)
- SKETCHES and FLOWS! I make a lot of sketches where there’s methods/compositions involved. I also try to properly write workflows in a sequence as it aids in a narratively coherent understanding. I DO NOT try to polish it, nobody else has to understand it, only me. I try to not be a perfectionist as then looking of a proper sketch/metaphor often leads to time wasting. (In case, I am making some kind of content, I use this sheet as a start and polish it further. NEVER try to be clean while understanding especially if your sheet isn’t going to be used by someone else!)
- DATA! I have an inherent problem with graphs (I am actively trying to mitigate it) so I spend a lot of time, trying to understand graphs and then convert them into legit statements (for my ease, graphs don’t instantly make sense to me for some reason) I verify if the data makes sense with respect to the abstract, how they’ve validated the results, the metrics they’ve chosen (this part can take some time, but with exposure to more papers, it starts getting efficient as validation methods/approaches often follow a core principle that can repeat (it’s like a six basic plots type thing))
I don’t try to make it clean. This is one might seem clean but as I’ve read a lot of papers before this and iterated on my method, I have sort of gained a knowledge of what to write, where and what can be expected so due to prior experience/iterations, the running clutter is naturally reduced.
My distillation sheet for the paper:


Question 2. Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more.
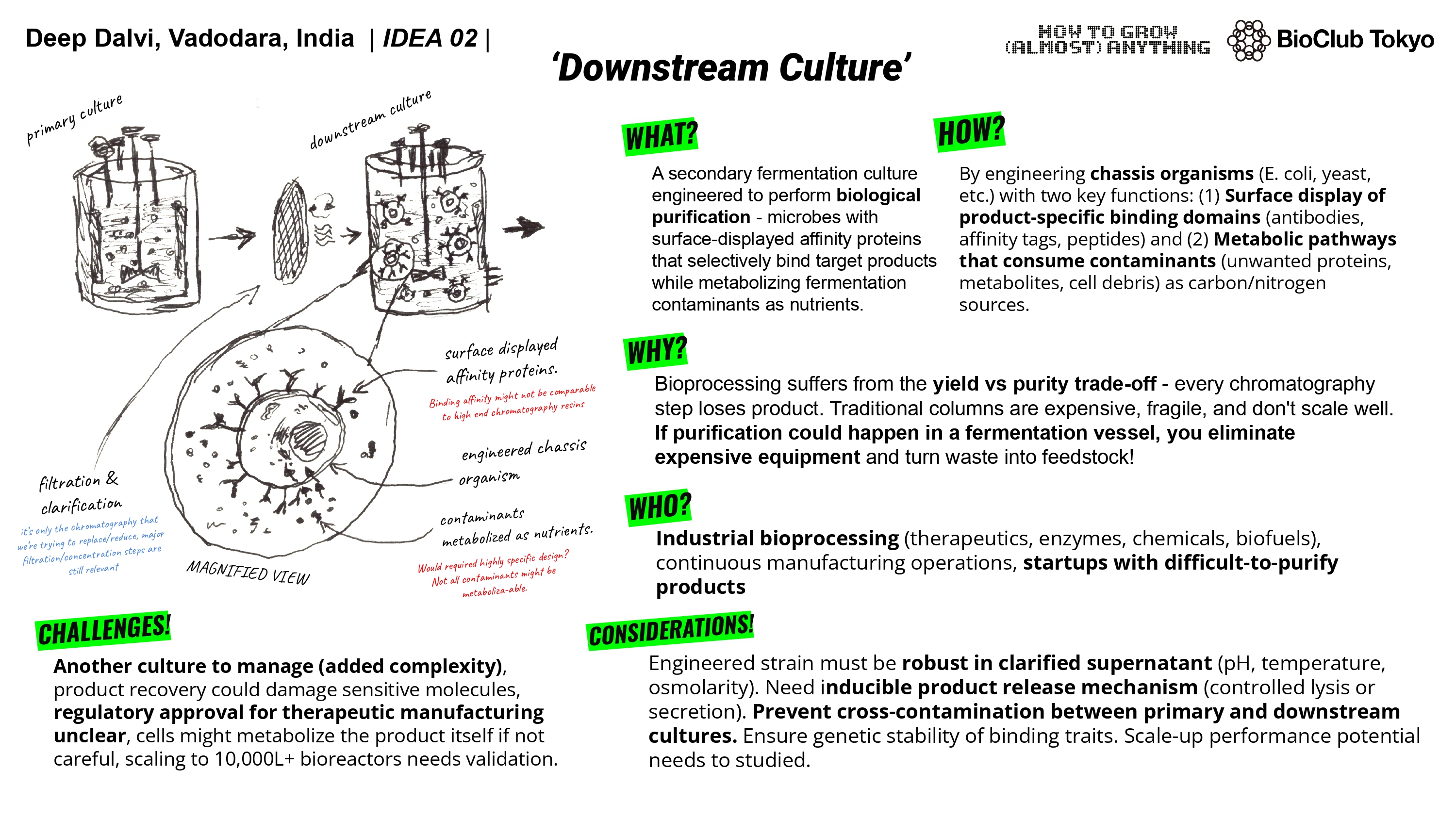
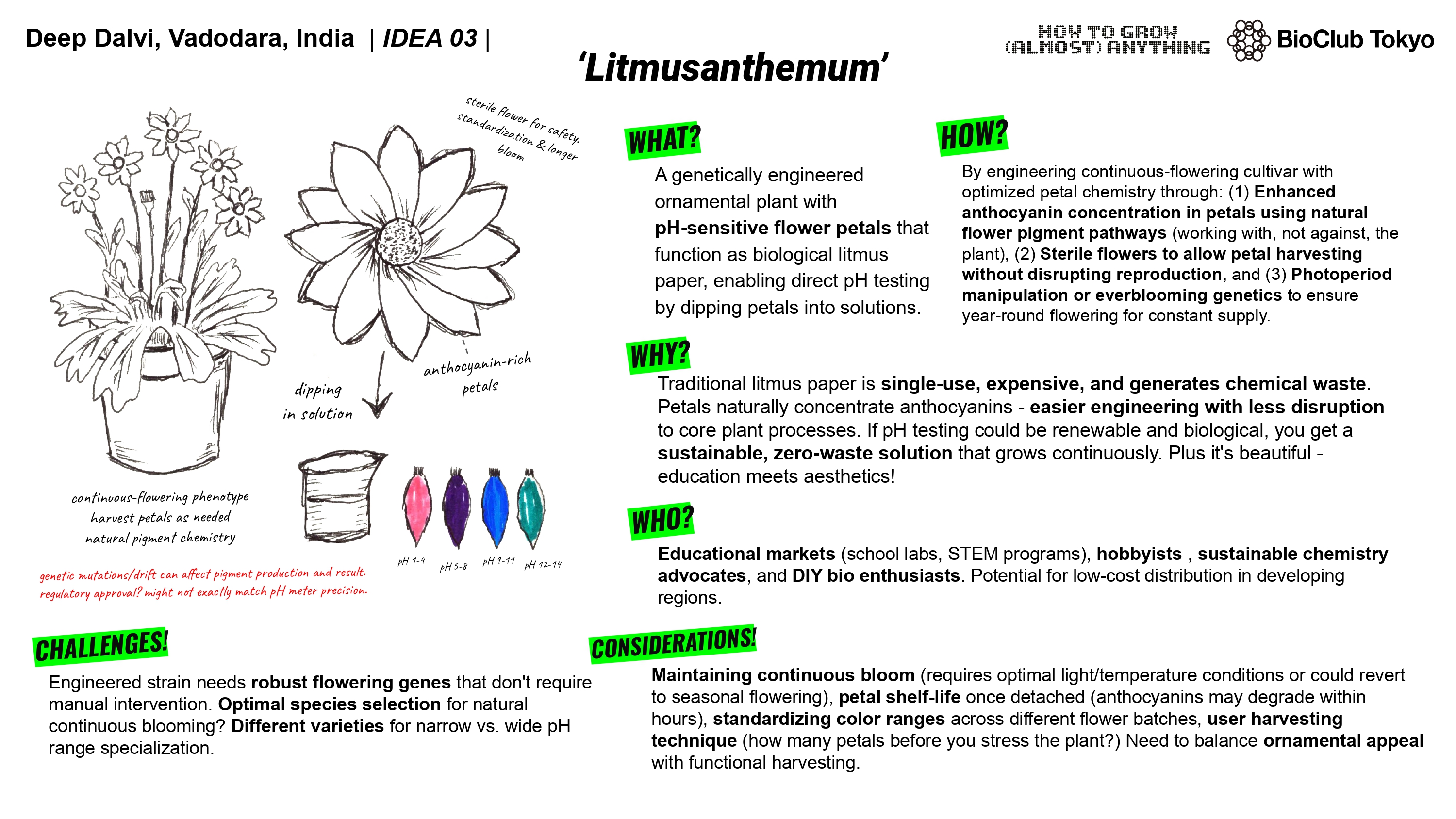
For my final project, I have multiple ideas. I could use lab automation to help me perfect the accessory/secondary culture for the second step. Allow me to test various strains of bacteria with the clarified broth from a benchtop bioreactor. The benchtop reactor would contain the target culture producing our product and then I could use multiple engineered cultures with different affinity proteins and test their efficiency in binding to the product, allowing to perfect the downstream culture. this would only leave the engineering / design part to the scientists and take away the tedious repetition.
The lab automations could also help me in testing out an engineered yeast culture for my astrobioreactor. Also if the opentrons could also have a custom automated temperature controlled centrifuge then I could test and iterate on the creation of modified yeast for better shear durability and heat resistance to support fermentation in space.
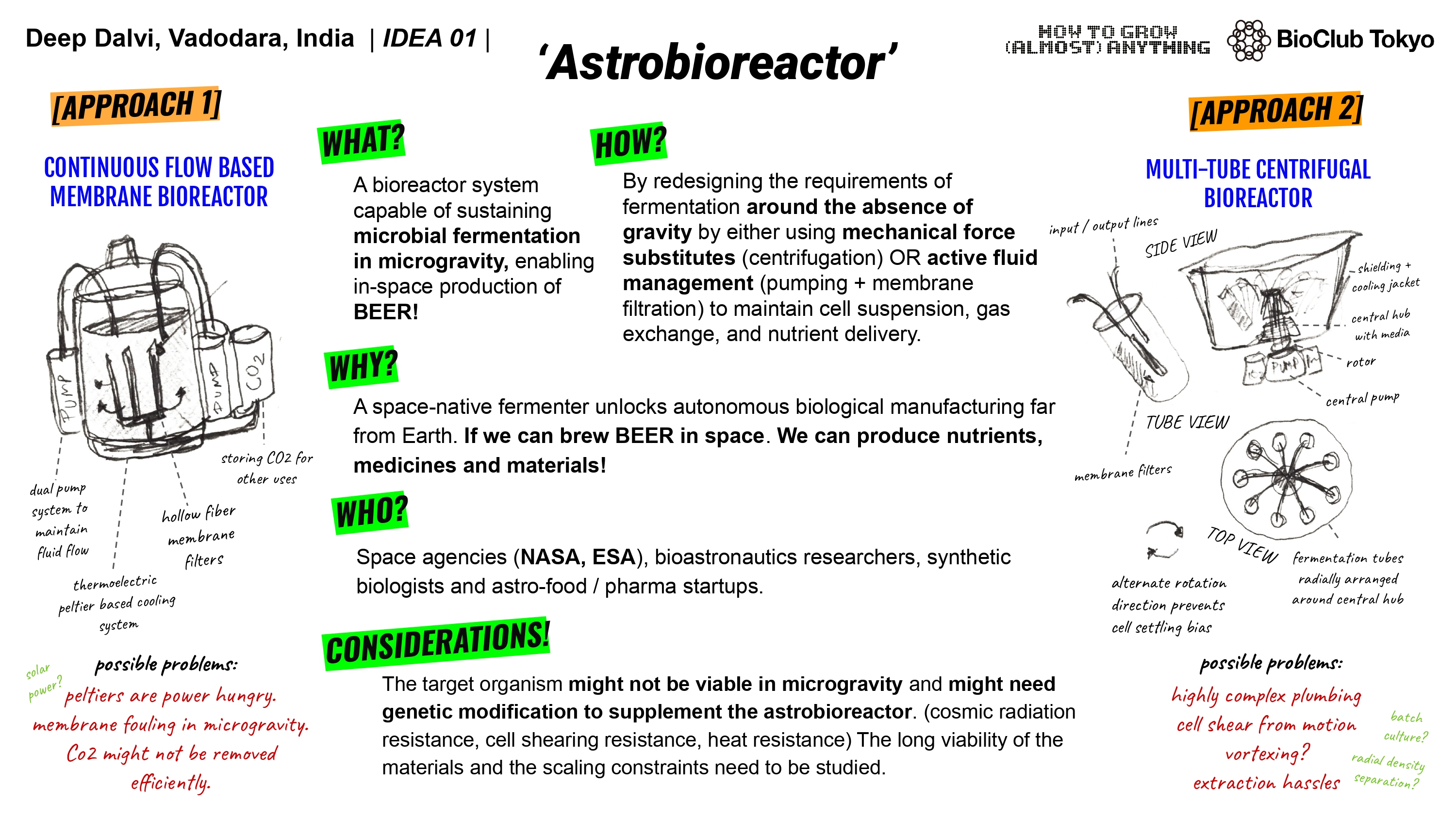
Final Project Ideas