Week 3 HW: Lab Automation
Project Overview: Cell-Free Allergen Biosensor
I am hoping to develop a rapid, consumer-grade biosensor designed to detect trace allergens like peanut or soy in a restaurant setting. To prioritize speed and accuracy, I will use a DNA-to-RNA detection circuit.
The workflow consists of three main stages:
- Extraction and Amplification: I could use RPA (Recombinase Polymerase Amplification) to exponentially copy target DNA (like the Ara h 1 gene) at a constant 37°C.
- Transcription: T7 RNA polymerase can concurrently convert that DNA into Trigger RNA.
- RNA Toehold Detection: This Trigger RNA can bind to a synthetic Toehold Switch, and unzip an RNA hairpin to allow the translation of a reporter protein. This can create a visible color change or induce luminescence in under 20 minutes.
By using a cell-free protein synthesis system, the entire reaction is shelf-stable and functions without the need for a traditional lab environment.
Automation Stack
- Nebula: I could use this for computational modeling to predict the thermodynamic stability ($\Delta G$) of different toehold designs. It can help me pick sequences that are stable at room temperature but unzip quickly when triggered.
- Opentrons OT-2: This is a good tool for kinetic screening. I can use it to automate the distribution of reagents and synchronize the start of up to 96 reactions across a plate, ensuring my “time-to-detection” data is precise.
- Ginkgo: I can look to foundry-scale automation for validating my optimized sensors against complex food matrices (oils, acids, salts) to minimize false negatives.
Kinetic Screening Pseudo-code
This protocol could be used to screen my Nebula-designed candidates to find the one with the fastest response time.
Novel Lab Automation for Brain Organoids
Overview Organoids are 3D in vitro models that closely replicate the biology and physiology of their in vivo counterparts, making them highly valuable for developmental research and disease modeling. However, traditional manual cell culture protocols often lack consistency and expose these complex models to erratic swings in nutrient availability and the buildup of toxic metabolites, which can lead to cellular stress.
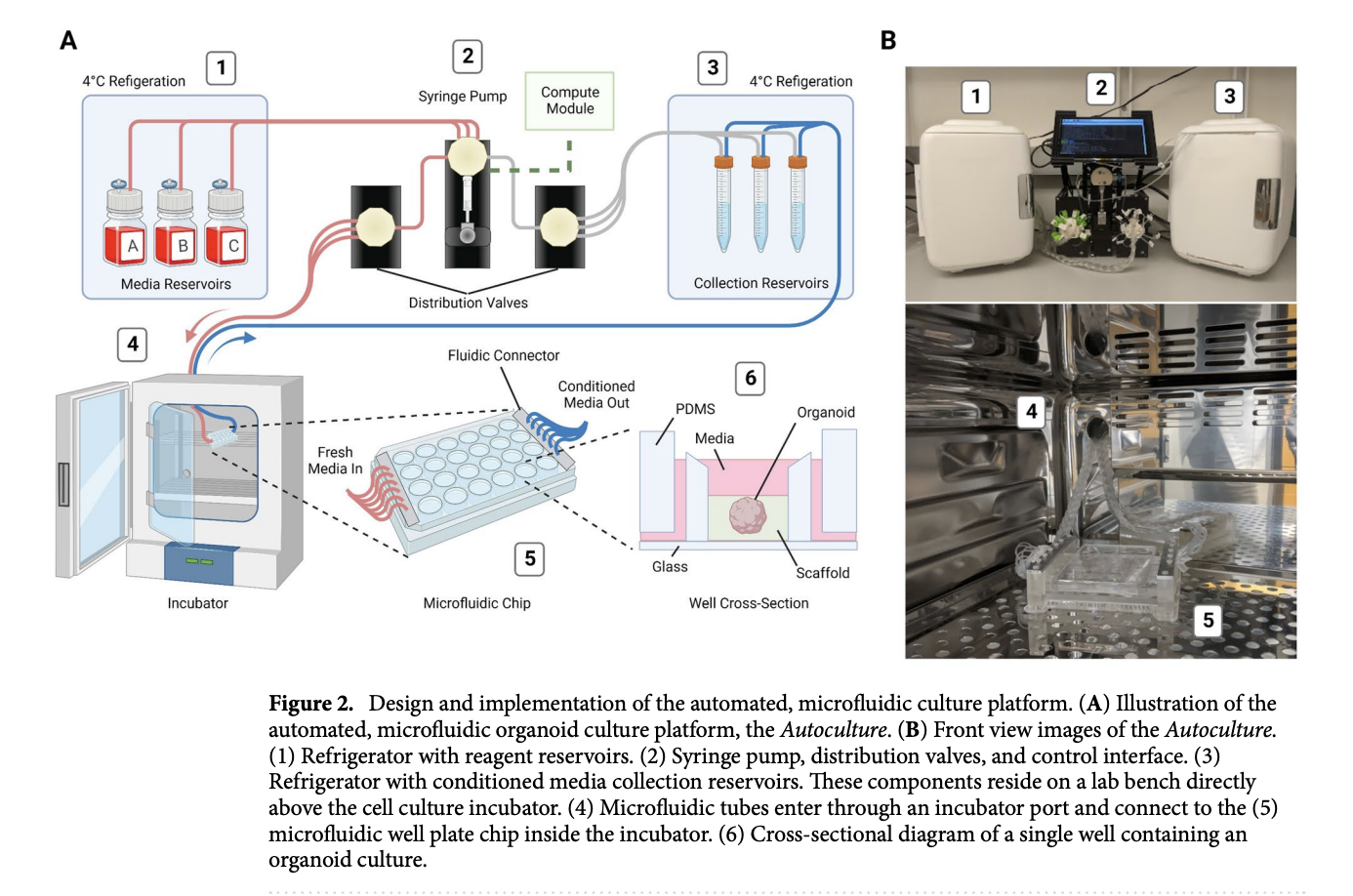
To address these limitations, researchers developed the “Autoculture” platform, a modular, automated microfluidic system designed to optimize 3D organoid growth. This Internet of Things (IoT)-enabled system uses a custom 24-well polydimethylsiloxane (PDMS) chip to house individual organoids in isolated microenvironments. The platform automatically delivers precisely scheduled media and removes waste without the need to take the cultures out of the incubator, offering high spatiotemporal resolution and customizable feeding schedules.
Findings When tested on cerebral cortex organoids over an 18-day period, the Autoculture platform supported robust growth and accurate neural differentiation comparable to conventional orbital shaker suspension methods. Crucially, RNA sequencing revealed that the automated microfluidic environment significantly reduced markers of cellular stress. Specifically, organoids grown in the automated system showed a marked downregulation in genes associated with canonical glycolysis and endoplasmic reticulum (ER) stress when compared to those maintained in traditional suspension cultures.