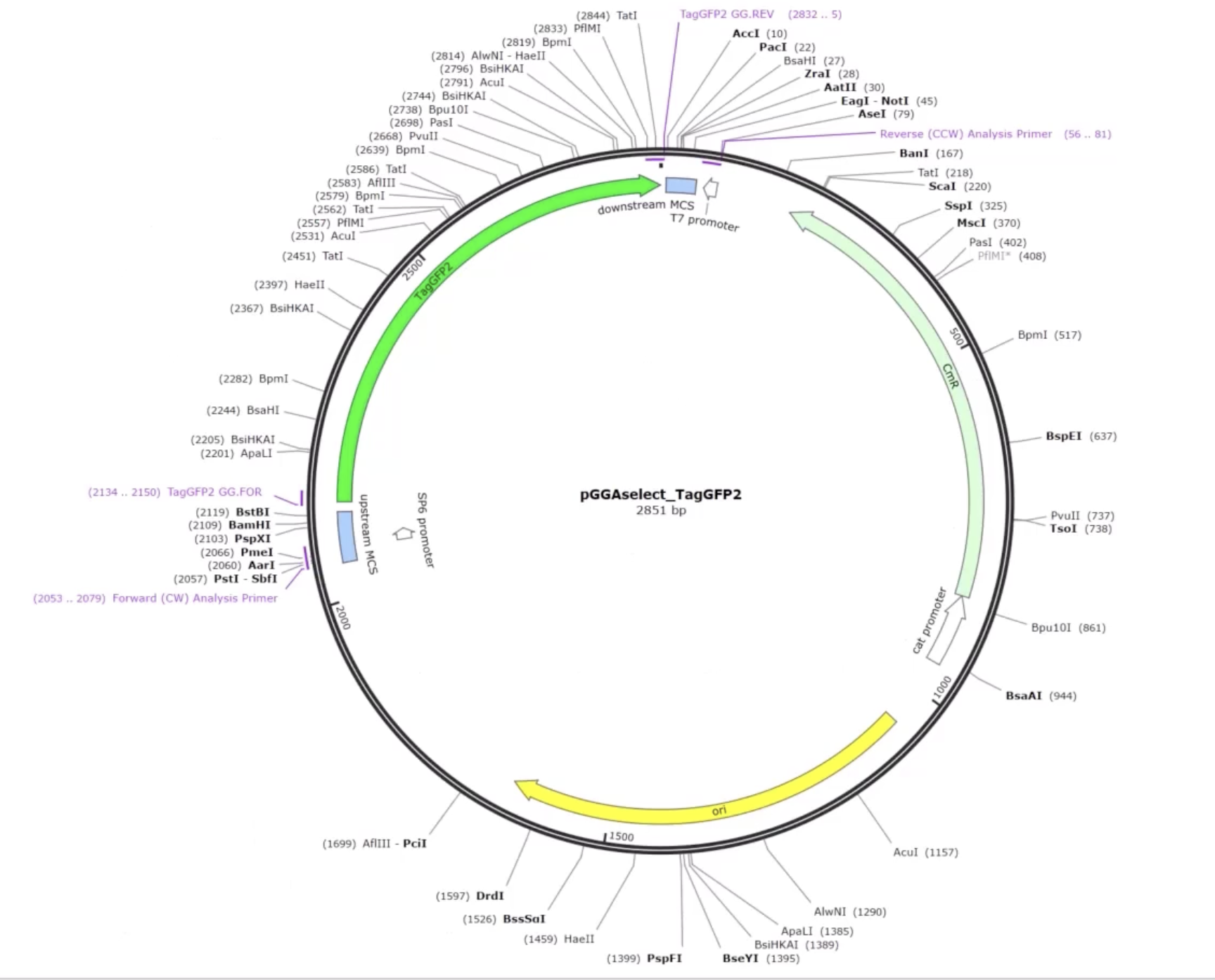
Week 6 HW: Genetic Circuits
HW Questions
- What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
- Phusion DNA Polymease, which synthesized new DNA by adding new nucleotides to the template DNA during PCR
- dNTPs, which are the nucleotide building blocks (dATP, dGTP, dCTP, and dTTP)
- reaction buffer, which acts as a chemical stabilizer that maintains the ideal pH and salt balance so the enzyme stays active and can accurately build new DNA strands.
- What are some factors that determine primer annealing temperature during PCR?
- Melting temperature of the primer, which is the temperature at which half of the DNA complex dissociates
- Primer length, since longer primers usually require higher annealing temperatures
- GC content, since higher GC content typically increases the primer melting temperature
- Salt concentration, since higher salt concentrations can stabilize the DNA and thus may require higher annealing temperatures
- There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
| Feature | PCR (Polymerase Chain Reaction) | Restriction Enzyme Digest |
|---|---|---|
| Mechanism | Enzymatic Synthesis: Building new strands from primers. | Enzymatic Cleavage: Cutting phosphodiester bonds at specific sites. |
| Protocol | Thermal Cycling: Repeated steps of denaturation (95°C), annealing (55-65°C), and extension (72°C). | Isothermal Incubation: DNA and enzymes are mixed in a buffer and held at a constant temp (usually 37°C). |
| Reagents | DNA template, Primers, dNTPs, Taq Polymerase, MgCl2, Buffer. | DNA template, Restriction Enzymes, specific BSA/Salt Buffer, Water. |
| Pros | High sensitivity; amplifies DNA; creates specific fragments without needing existing cut sites. | Simple setup; highly reproducible; great for verifying known sequences or circular DNA. |
| Cons | Prone to contamination; requires known flanking sequences; potential for polymerase errors. | Does not amplify DNA; limited by the location of natural recognition sites. |
| When to Use | When you have minimal DNA, need a custom fragment, or want to add “tails” for cloning. | When linearizing plasmids, performing diagnostic checks, or subcloning existing inserts. |
- How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
- Both the PCR and digested fragments must share identical overlapping terminal sequences (15–40 bp) with their neighboring fragments to allow for seamless homology-directed assembly.
- How does the plasmid DNA enter the E. coli cells during transformation?
- Membrane pores open due to a thermal pressure imbalance during the heat shock, allowing the plasmid DNA (which has been neutralized by calcium ions) to be pulled into the cell.
- Describe another assembly method in detail (such as Golden Gate Assembly)
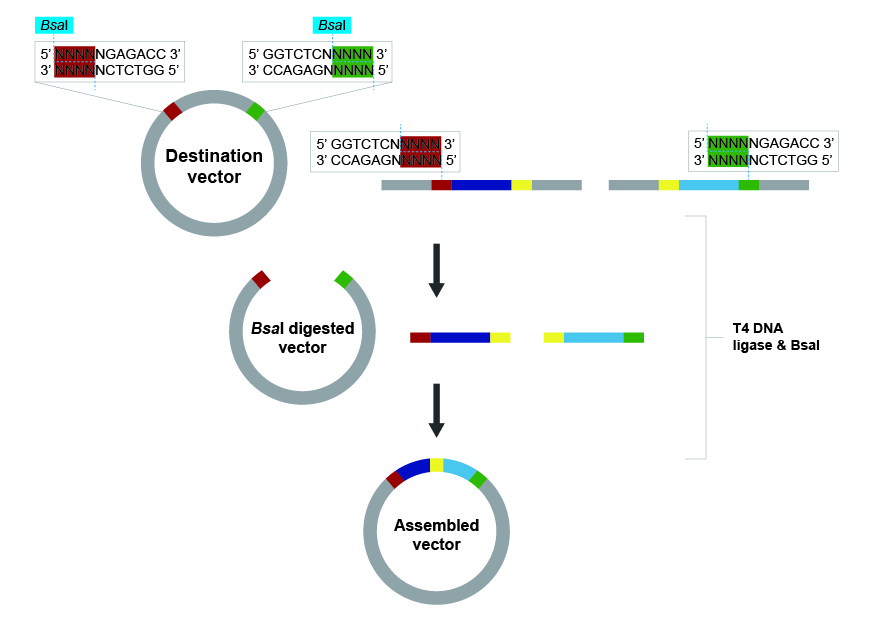
- Golden Gate Assembly is a highly efficient “one-pot” cloning method that allows you to join multiple DNA fragments together simultaneously using Type IIS restriction enzymes and T4 DNA ligase. Unlike standard enzymes, Type IIS enzymes like BsaI bind to a specific recognition sequence but cut the DNA several nucleotides away, creating custom 4-base overhangs. By strategically designing these overhangs to be complementary, you can ensure that multiple fragments assemble in a specific, directional order. During the reaction, you cycle the temperature to repeatedly cut and ligate the DNA until the fragments are perfectly joined. A key advantage is that the enzyme’s recognition sites are positioned to be “cut off” and removed during the process, meaning the final product cannot be re-cut. This makes the reaction irreversible and drives the assembly toward the final, seamless circular plasmid. Because of this precision, Golden Gate is the gold standard for modular cloning and building complex multi-gene constructs.

Simulating Golden Gate using AddGene’s tool
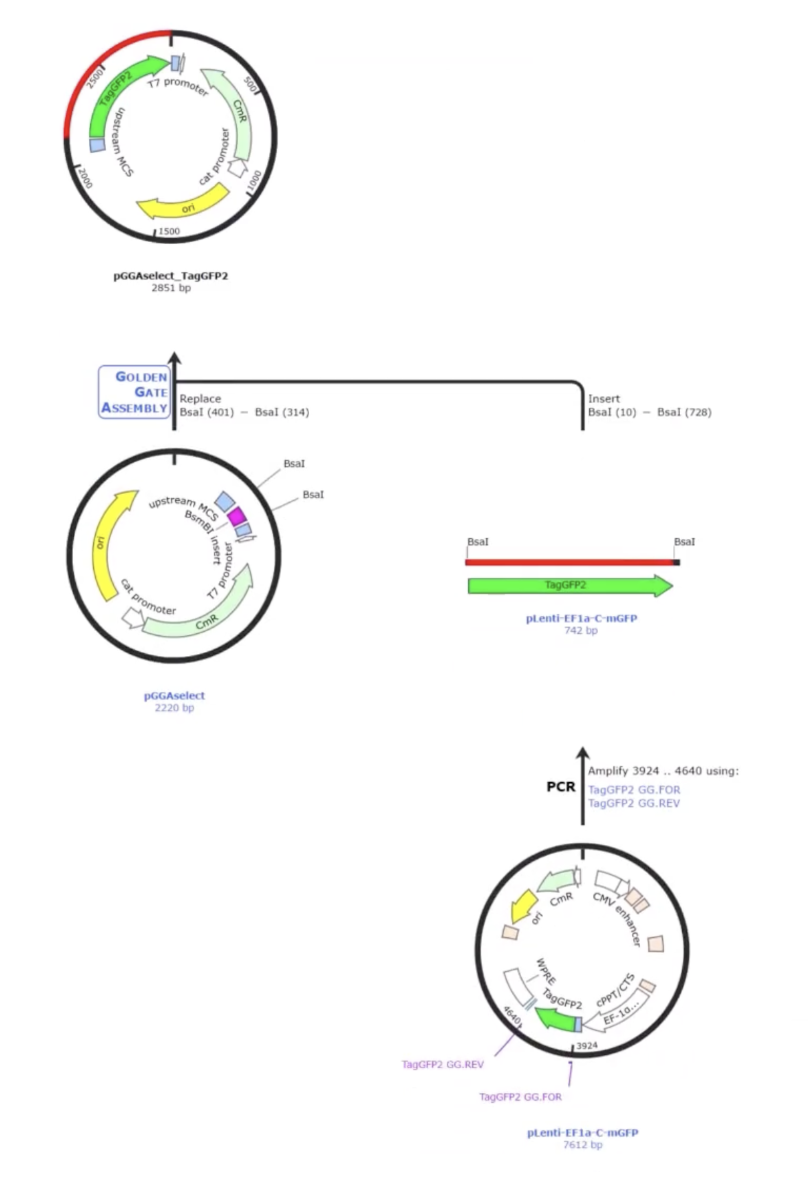

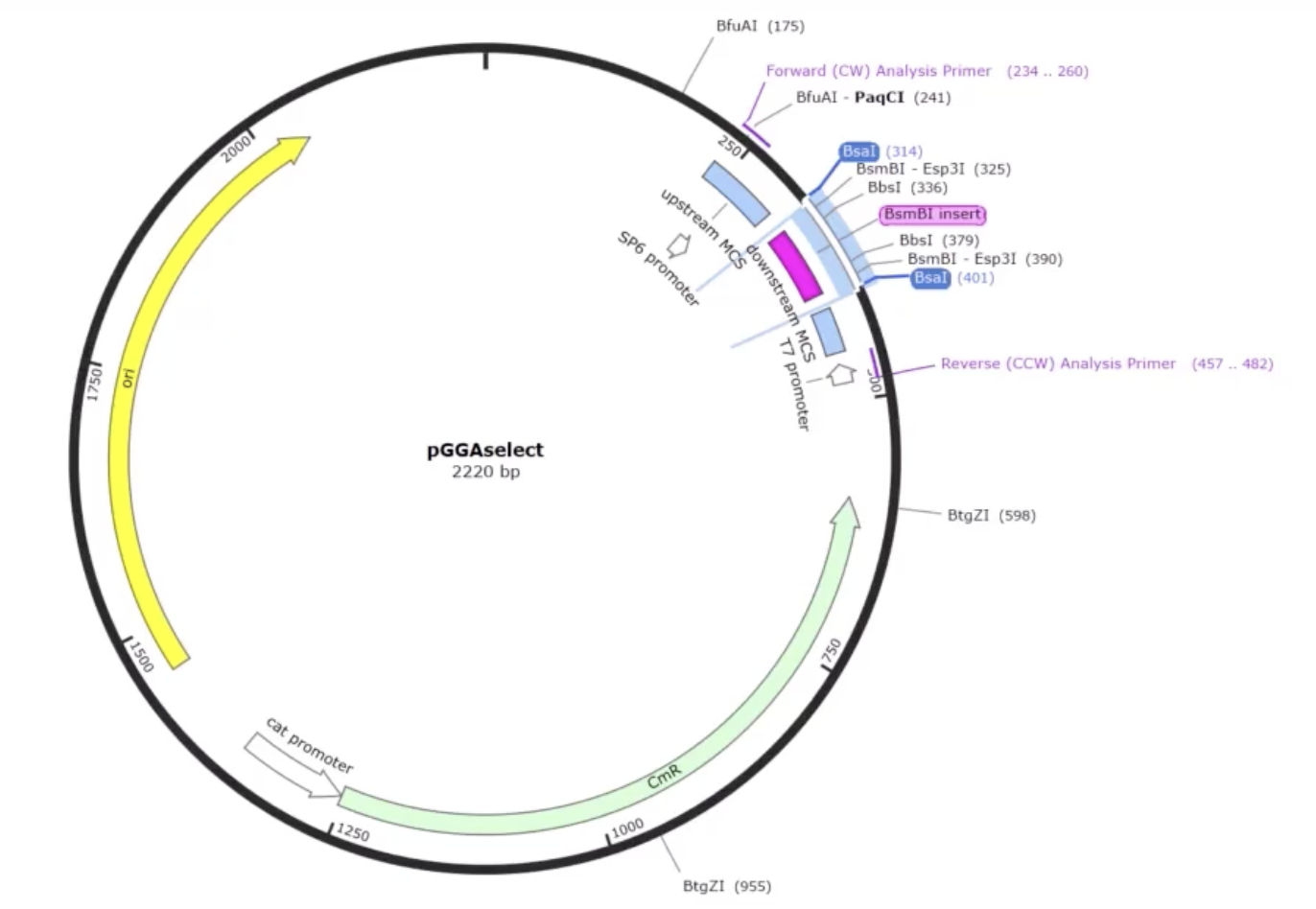
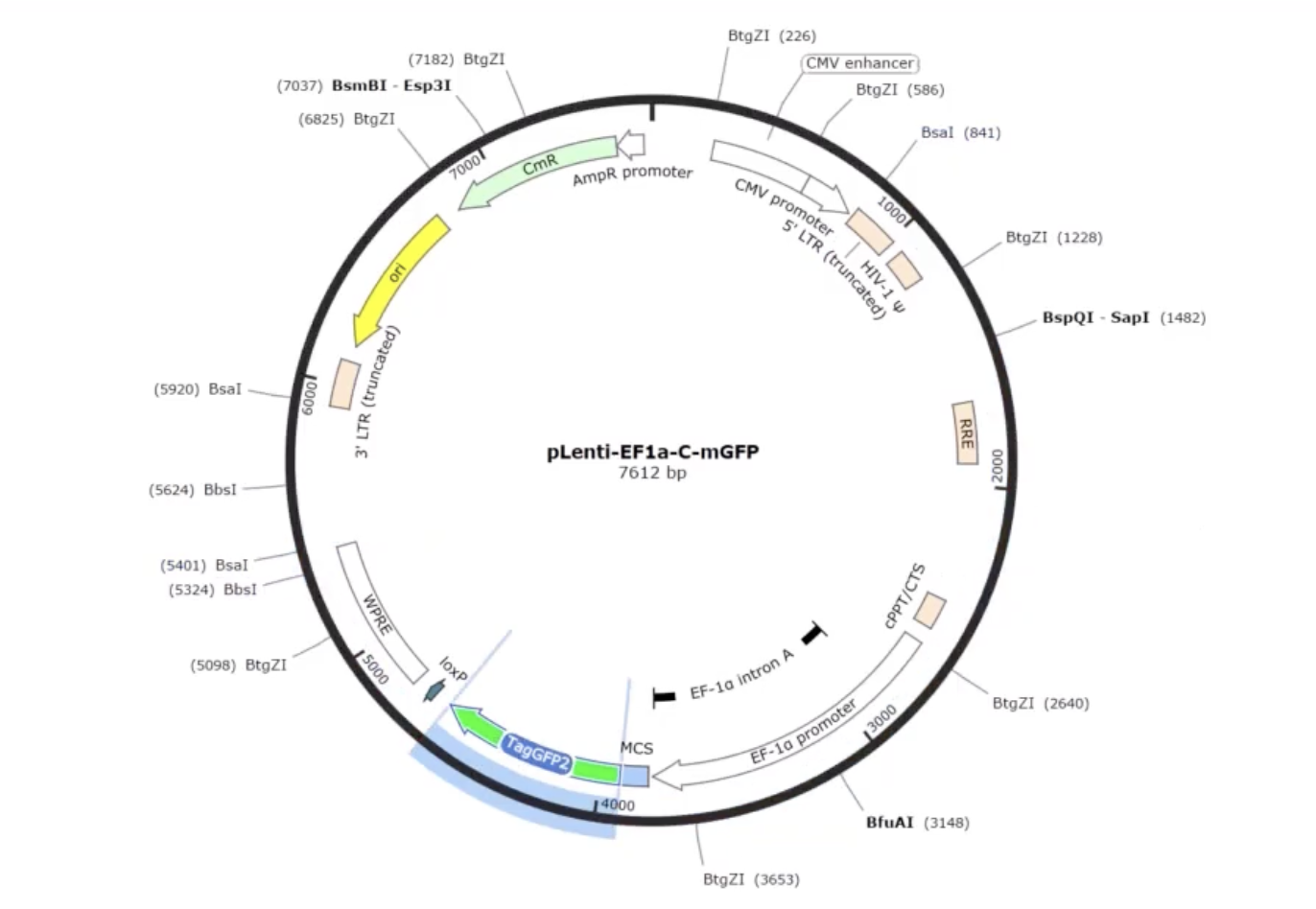
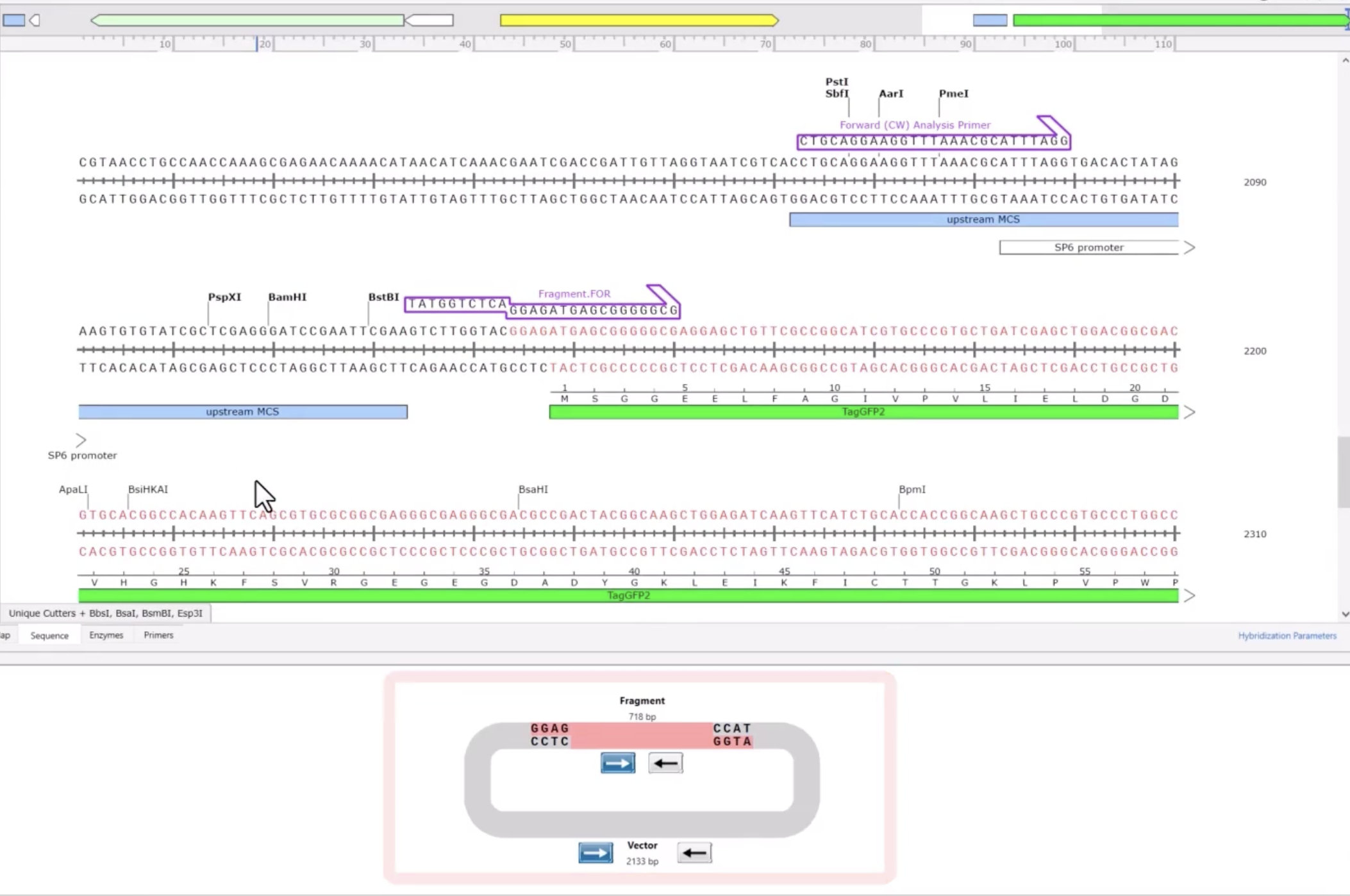
- AddGene’s tool allows you to work with either a single insert or multiple fragments (up to 100), and it handles both vectors that already have Type IIS restriction sites and those that don’t. When sites are missing, it automatically designs the PCR primers needed to add them. You can also set your preferred PCR primer Tm. The tool selects the appropriate enzyme recognition sites, predicts how overhangs will interact, and flags any potential mis-ligation issues through an assembly fidelity score. Once you’re happy with the design, it simulates the full digest-and-ligate reaction and generates the predicted product sequence, which you can inspect at the fusion points to confirm everything is in frame and in the right order. It also exports your primer sequences ready to order.
Asimov Kernel
Repressilator
I reacreated the represillator by searching dragging parts one by one from the Parts Search.
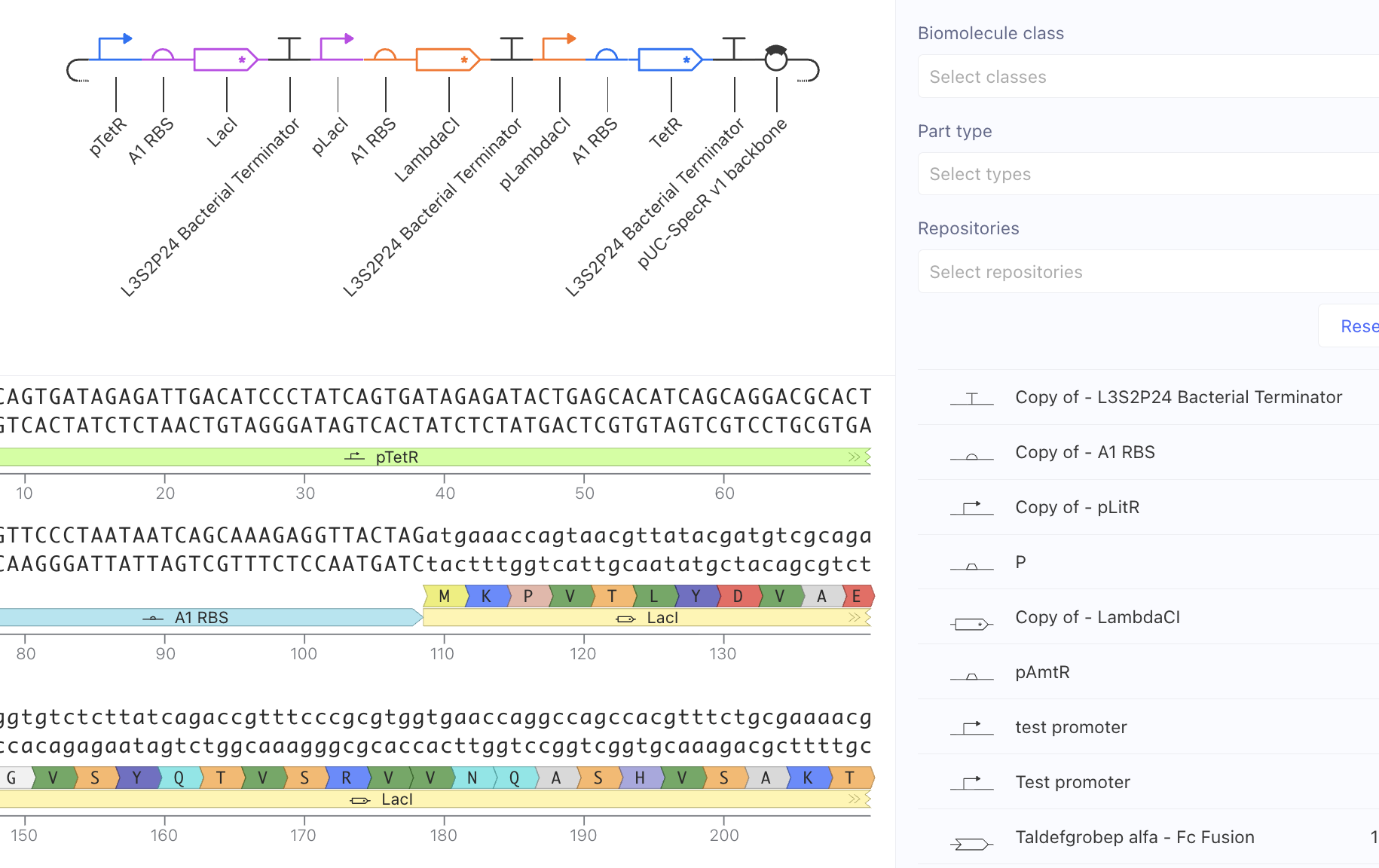

The results of the simulation were similar to the simulation of the original represillator. The simulation shows oscillations in expression levels for all three proteins.
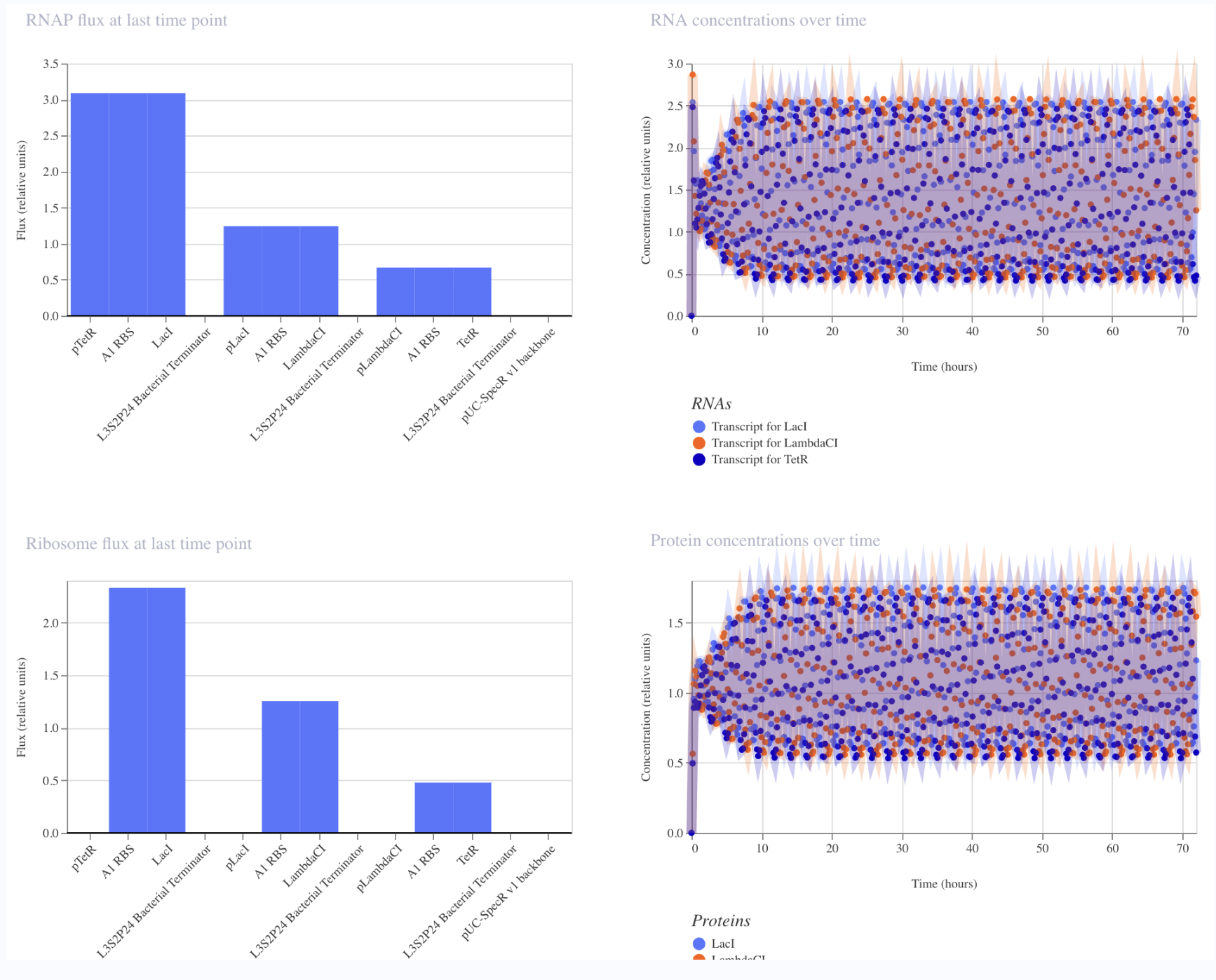

Original Constructs
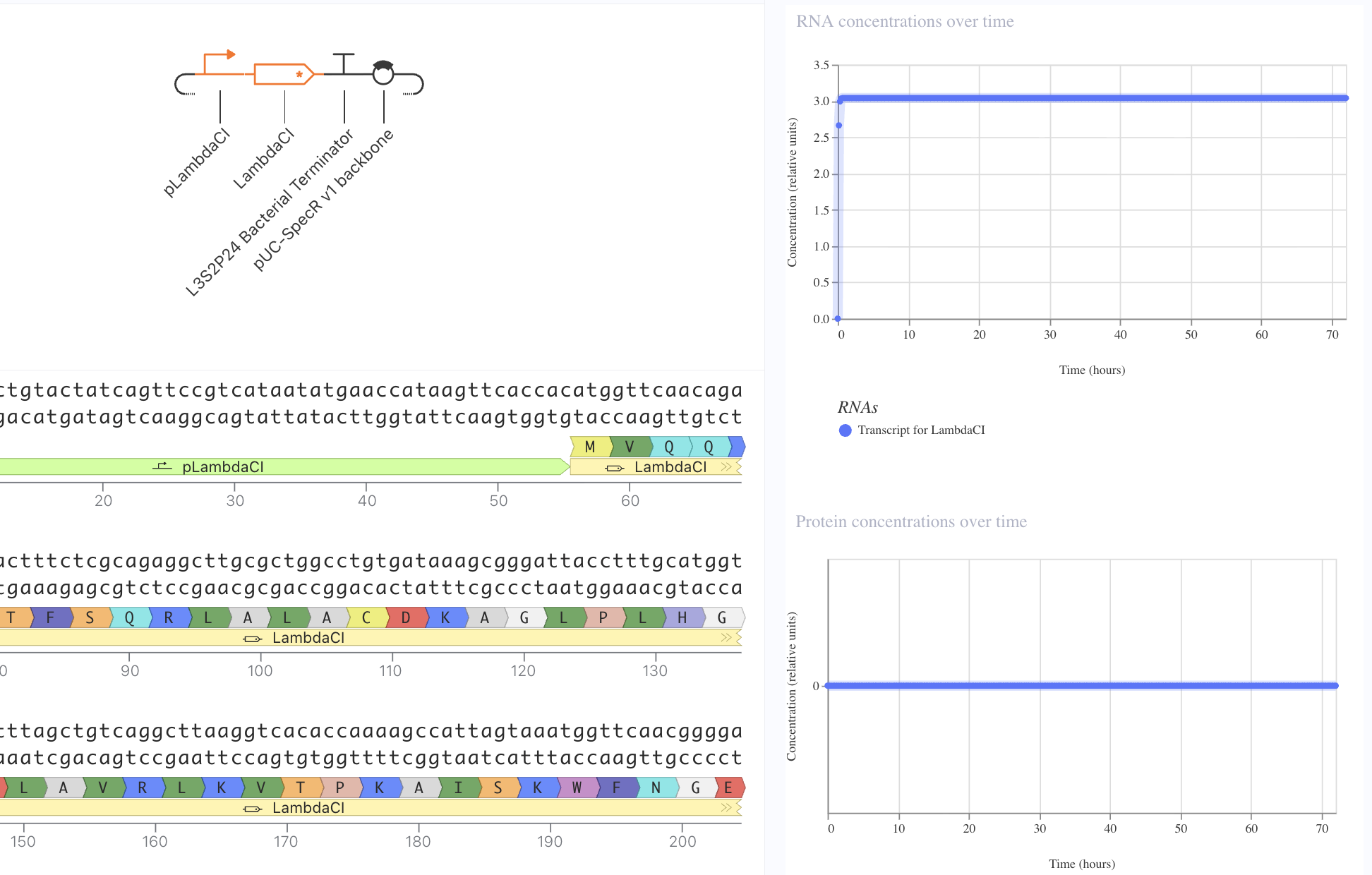
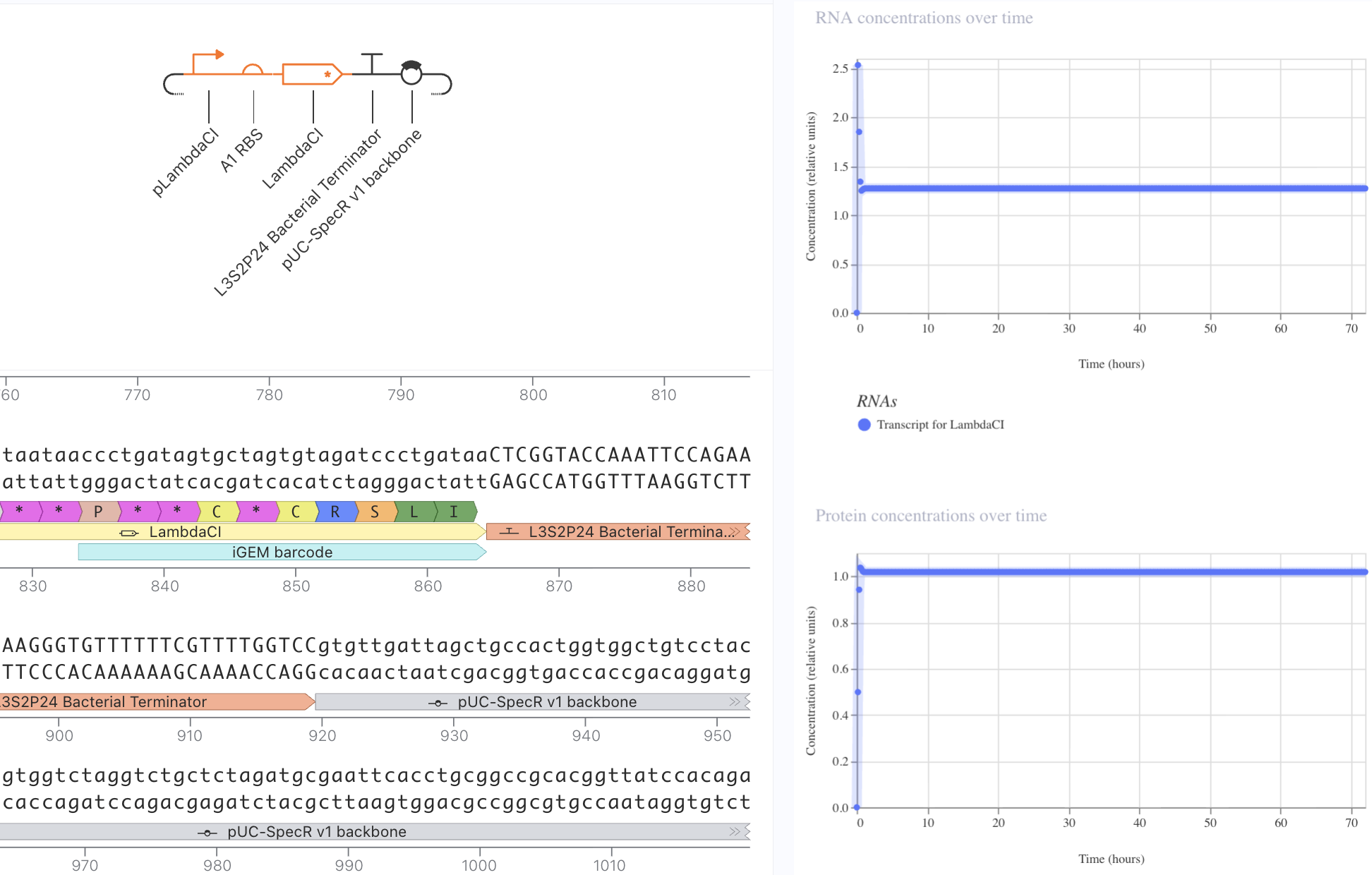
I first started with a simple construct consisting of a LambdaCI promoter, an A1 RBS, the LambdaCI CDS, a bacterial terminator, and a backbone. Simulating showed constant concentration of RNA, and highly expressed protein levels.

Next, I removed the promoter. This meant that no presence of RNA or protein expression were predicted.
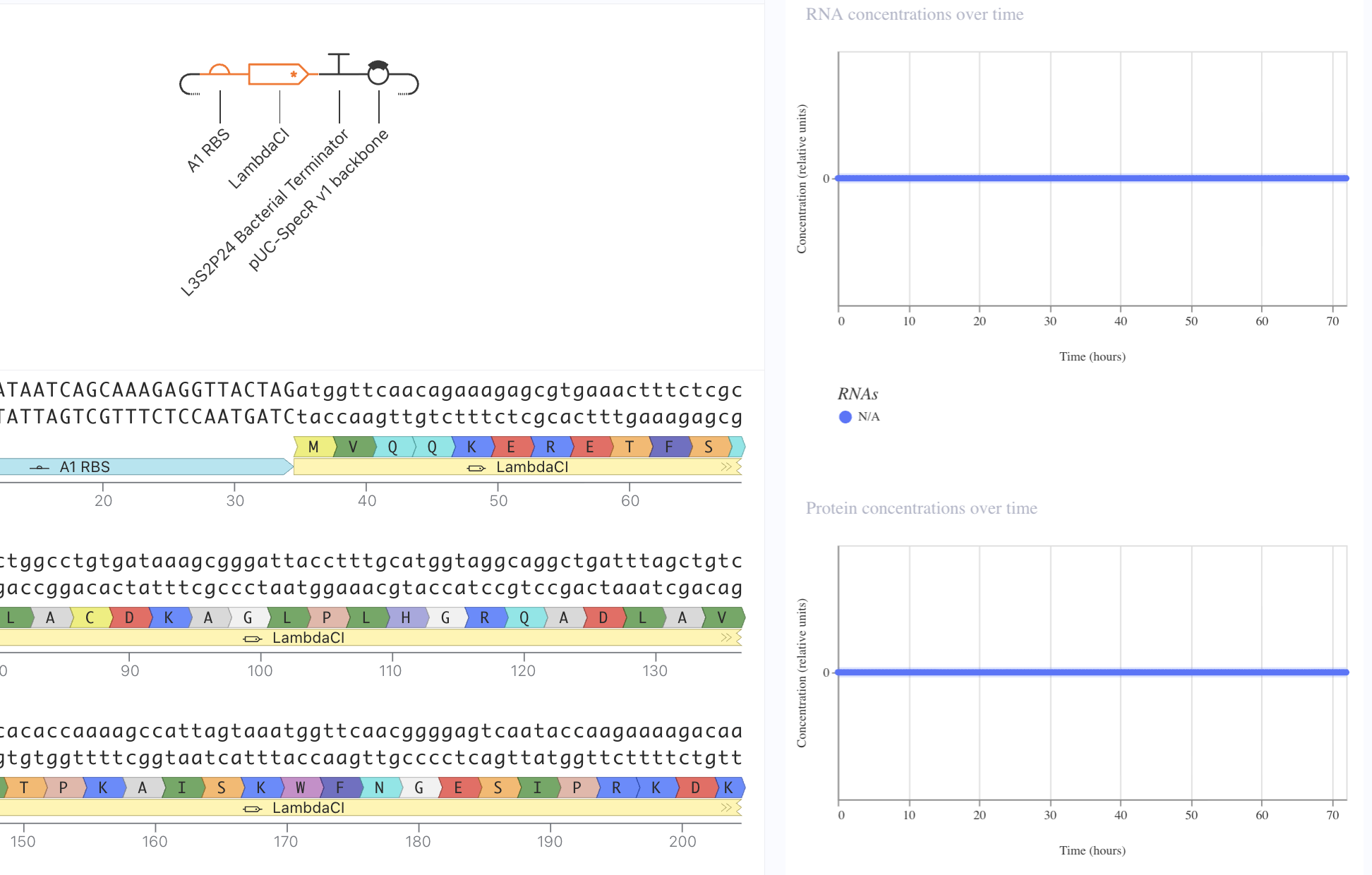

Lastly, I removed the RBS. Here, despite predicted presence of RNA, there was no protein expression. This reveals the necessity of the RBS for protein epression.