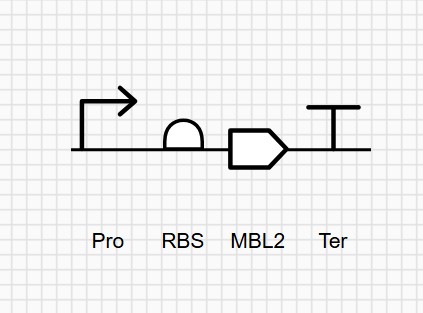
Week 2 HW: DNA Read, Write and Edit
Part 1: Benchling & In-silico Gel Art
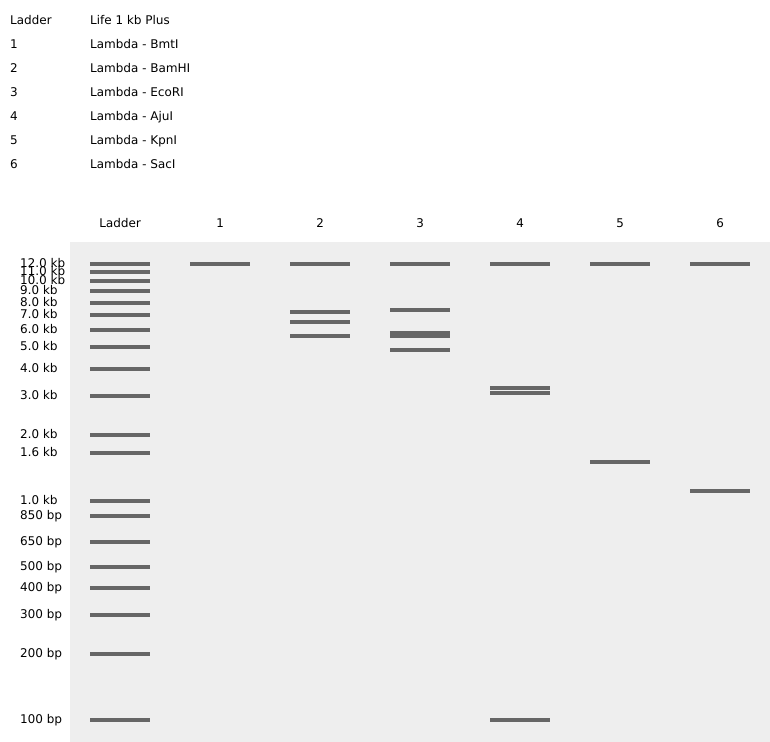


Part 3: DNA Design Challenge
Protein: mannose-binding protein C precursor
Reverse Translate:
Aminoacids
mslfpslpll llsmvaasys etvtcedaqk tcpaviacss pgingfpgkd grdgtkgekg epgqglrglq gppgklgppg npgpsgspgp kgqkgdpgks pdgdsslaas erkalqtema rikkwltfsl gkqvgnkffl tngeimtfek vkalcvkfqa svatprnaae ngaiqnlike eaflgitdek tegqfvdltg nrltytnwne gepnnagsde dcvlllkngq wndvpcstsh lavcefpi
Nucleotid sequence
atgagcctgtttccgagcctgccgctgctgctgctgagcatggtggcggcgagctatagc gaaaccgtgacctgcgaagatgcgcagaaaacctgcccggcggtgattgcgtgcagcagc ccgggcattaacggctttccgggcaaagatggccgcgatggcaccaaaggcgaaaaaggc gaaccgggccagggcctgcgcggcctgcagggcccgccgggcaaactgggcccgccgggc aacccgggcccgagcggcagcccgggcccgaaaggccagaaaggcgatccgggcaaaagc ccggatggcgatagcagcctggcggcgagcgaacgcaaagcgctgcagaccgaaatggcg cgcattaaaaaatggctgacctttagcctgggcaaacaggtgggcaacaaattttttctg accaacggcgaaattatgacctttgaaaaagtgaaagcgctgtgcgtgaaatttcaggcg agcgtggcgaccccgcgcaacgcggcggaaaacggcgcgattcagaacctgattaaagaa gaagcgtttctgggcattaccgatgaaaaaaccgaaggccagtttgtggatctgaccggc aaccgcctgacctataccaactggaacgaaggcgaaccgaacaacgcgggcagcgatgaa gattgcgtgctgctgctgaaaaacggccagtggaacgatgtgccgtgcagcaccagccat ctggcggtgtgcgaatttccgatt
Codon optimization:
ATG AGC CTT TTT CCG AGC CTT CCT CTG CTT TTA CTG TCG ATG GTG GCC GCC AGC TAC AGT GAA ACT GTG ACC TGT GAG GAC GCC CAA AAA ACG TGT CCT GCA GTT ATC GCG TGC AGC TCC CCG GGT ATC AAT GGC TTC CCC GGC AAG GAC GGG CGT GAT GGG ACT AAA GGC GAG AAA GGT GAA CCG GGA CAG GGC TTA CGT GGT TTA CAG GGC CCG CCG GGT AAA TTG GGG CCG CCA GGC AAT CCG GGT CCG AGT GGC TCC CCA GGG CCG AAA GGT CAG AAA GGC GAT CCA GGC AAA AGT CCG GAT GGT GAT TCA AGT CTG GCG GCC AGC GAA CGT AAG GCC CTT CAG ACC GAA ATG GCT CGT ATC AAA AAA TGG TTA ACG TTC AGC CTG GGG AAA CAA GTG GGG AAT AAG TTT TTT CTG ACT AAT GGC GAG ATC ATG ACG TTT GAG AAA GTG AAA GCG CTG TGT GTG AAG TTC CAG GCC AGC GTG GCG ACG CCA CGT AAC GCG GCG GAA AAT GGC GCG ATT CAA AAC CTT ATC AAA GAA GAG GCC TTC CTG GGT ATT ACG GAC GAA AAA ACG GAG GGC CAG TTT GTC GAT CTG ACT GGT AAC CGC TTA ACA TAT ACC AAT TGG AAT GAG GGC GAA CCT AAC AAC GCA GGC AGC GAT GAG GAC TGC GTG CTG TTA TTG AAA AAC GGC CAG TGG AAC GAC GTA CCT TGT TCC ACT AGC CAT TTA GCG GTA TGC GAA TTT CCG ATT
Why is it necessary to optimize codon usage?
Because several codons encode the same amino acid, but the frequency at which these codons are used varies among organisms. Each species has preferences for particular codons, a phenomenon known as codon usage bias. When expressing a gene from one organism in a different host without prior codon optimization, translation efficiency can be reduced, leading to lower protein production or even affecting proper protein folding.
Which organism have I chosen to optimize the codon sequence for, and why?
I chose to optimize the sequence for Escherichia coli because it is one of the most widely used systems for recombinant protein production. It is easy to cultivate, grows quickly, and is cost‑effective. In addition, there are well‑established genetic tools that enable efficient protein expression when codons are adapted to its natural codon bias, and it is the organism I am most familiar with.
You have a sequence! Now what?:
What technologies could be used to produce this protein from your DNA?
Recombinant gene expression in bacteria, yeasts, or cell‑free systems
Cell-dependent transcription and translation
The gene is inserted into a plasmid and then introduced into a host organism. The process begins with cloning, where the codon‑optimized gene is inserted into a plasmid. During transformation or transfection, the plasmid is delivered into the host cell. Inside the cell, the host RNA polymerase recognizes the promoter and transcribes the DNA into mRNA. This mRNA is then read by ribosomes, which synthesize the protein by assembling amino acids according to the codon sequence. Finally, the newly synthesized protein folds and, if necessary, undergoes further modifications.
Part 4: Prepare a Twist DNA Synthesis Order
Benchling: https://benchling.com/s/seq-9syL7gvyZin8DtEAROGk?m=slm-yWJdRnMP3mRA7qSkwxTO
Part 5: DNA Read/Write/Edit
READ
What DNA would you want to sequence and why?
MBL for be used as an initial viral capture molecule due to its affinity for high-mannose glycans present on the viral envelope protein.
What technology or technologies would you use to perform sequencing on your DNA and why?
To sequence DNA, I would use Illumina sequencing for its high accuracy and cost-effectiveness, since my idea is targeted gene analysis and Illumina technology provides low error rates and high throughput, making it ideal for detecting deletions.
Is your method first-, second or third-generation or other? How so?
Second generation, because it performs massive sequencing in parallel, use clonal amplification (bridge amplification) before reading and is necessary perform the amplification before sequencing
What is your input? How do you prepare your input? List the essential steps.
Input: MBL2
Preparation: Using linear synthetic DNA MBL2 sequence
- End repair.
- A-tailing.
- Adapter ligation for Illumina-compatible adapters
- PCR enrichment of adapter-ligated fragments.
- Cleanup and quantification.
What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample? Essential steps:
Flow Cell Binding MBL2 gene fragments with adapters are loaded onto the flow cell. The fragments hybridize to complementary oligonucleotides immobilized on the surface.
Bridge Amplification Each fragment is locally amplified, forming clonal clusters, generating many identical copies of each fragment, increasing the detectable signal during sequencing.
Sequencing by Synthesis Fluorescently labeled nucleotides with reversible terminators. In each cycle: • A labeled nucleotide (A, T, C, or G) is added. • Only one nucleotide is incorporated per cycle. • A camera detects the emitted fluorescence. • The system records the corresponding color. • The fluorophore and chemical blocking group are then removed. • The cycle is repeated.
Base calling performed through:
Optical detection of the fluorescent signal in each cluster. Each color corresponds to a specific base (A, T, C, or G), the software converts the light signal into a nucleotide sequence, and a quality score is assigned to each base.
What is the output of your chosen sequencing technology?
Millions of short reads. Generated in FASTQ format files, with the nucleotide sequence of each read and a quality score assigned to every base.
WRITE
What DNA would you want to synthesize (e.g., write) and why?
would synthesize a codon-optimized version of the MBL2 coding sequence (CDS) to produce recombinant MBL protein for use as a viral capture molecule in a diagnostic assay.
ATG AGC CTT TTT CCG AGC CTT CCT CTG CTT TTA CTG TCG ATG GTG GCC GCC AGC TAC AGT GAA ACT GTG ACC TGT GAG GAC GCC CAA AAA ACG TGT CCT GCA GTT ATC GCG TGC AGC TCC CCG GGT ATC AAT GGC TTC CCC GGC AAG GAC GGG CGT GAT GGG ACT AAA GGC GAG AAA GGT GAA CCG GGA CAG GGC TTA CGT GGT TTA CAG GGC CCG CCG GGT AAA TTG GGG CCG CCA GGC AAT CCG GGT CCG AGT GGC TCC CCA GGG CCG AAA GGT CAG AAA GGC GAT CCA GGC AAA AGT CCG GAT GGT GAT TCA AGT CTG GCG GCC AGC GAA CGT AAG GCC CTT CAG ACC GAA ATG GCT CGT ATC AAA AAA TGG TTA ACG TTC AGC CTG GGG AAA CAA GTG GGG AAT AAG TTT TTT CTG ACT AAT GGC GAG ATC ATG ACG TTT GAG AAA GTG AAA GCG CTG TGT GTG AAG TTC CAG GCC AGC GTG GCG ACG CCA CGT AAC GCG GCG GAA AAT GGC GCG ATT CAA AAC CTT ATC AAA GAA GAG GCC TTC CTG GGT ATT ACG GAC GAA AAA ACG GAG GGC CAG TTT GTC GAT CTG ACT GGT AAC CGC TTA ACA TAT ACC AAT TGG AAT GAG GGC GAA CCT AAC AAC GCA GGC AGC GAT GAG GAC TGC GTG CTG TTA TTG AAA AAC GGC CAG TGG AAC GAC GTA CCT TGT TCC ACT AGC CAT TTA GCG GTA TGC GAA TTT CCG ATT
What technology or technologies would you use to perform this DNA synthesis and why? Array-based or column-based phosphoramidite DNA synthesis. Because it is a method with high sequence fidelity and control over sequence design
What are the essential steps of your chosen sequencing methods?
a) Solid-phase attachment b) Deprotection c) Coupling d) Capping e) Oxidation f) Cleavage and deprotection g) Sequence verification
What are the limitations of your sequencing method in terms of speed, accuracy, scalability?
In terms of accuracy, each coupling step is not 100% efficient, so errors can accumulate as the sequence length increases. Regarding length, synthesis is limited to about 150–200 base pairs per oligonucleotide, meaning longer genes must be assembled from multiple overlapping fragments. In terms of speed, synthesis is relatively fast for short oligos, but full gene synthesis requires extra assembly and validation steps,increasing turnaround time. Concerning scalability, array-the individual yield per oligo may be lower compared to column-based synthesis. Finally, cost increases with more complex constructs due to the need for assembly and error correction.
EDIT
What DNA would you want to edit and why?
MBL2 gene, which encodes Mannose-binding lectin, to enhance its expression in individuals with naturally low serum MBL levels. Increased MBL expression could potentially improve early immune recognition of viral pathogens
What kinds of edits might you want to make to DNA? Why?
Precise regulatory modification rather than altering the protein-coding sequence. Introducing promoter variants associated with higher transcriptional activity could increase protein levels without changing protein structure, minimizing unintended functional consequences while enhancing host defense.
What technology or technologies would you use to perform these DNA edits and why?
CRISPR-Cas9 with Homology-Directed Repair, because the precise control over the type of genetic modification and flexibility depending on whether the goal is a single-base change or a larger regulatory insertion.
How does your technology of choice edit DNA? What are the essential steps?
Using a guide RNA (gRNA) to direct the Cas9 nuclease to a specific genomic sequence. Target recognition DNA cleavage DNA repair
What preparation do you need to do and what is the input for the editing?
Design a specific guide RNA targeting for MBL2, verifying minimal off-target sites using bioinformatic tools. Inputs: Cas9 protein, Guide RNA, delivery system and target cell
What are the limitations of your editing methods (if any) in terms of efficiency or precision?
Efficiency: HDR efficiency is often low in non-dividing cells, and editing rates depend on the cell type and genomic context.
Precision: Potential off-target edits if the guide RNA is not highly specific and NHEJ repair may introduce unintended insertions or deletions.