Week 5 HW: Genetic circuits part 1
Assignment: DNA Assembly
- What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
Phusion High-Fidelity DNA polymerase: enzyme responsible for synthesizing new DNA strands while possessing activity, which reduces errors during replication.
dNTPs (deoxynucleotide triphosphates): Nucleotide substrates incorporated by DNA polymerase into the elongating DNA strand during synthesis
reaction buffer: contain compounds as Tris-HCl that maintains the correct pH and salts like KCl which help stabilize primer binding and enzyme activity
MgCl₂: Essential cofactor required for DNA polymerase catalytic activity.
- What are some factors that determine primer annealing temperature during PCR?
Tm Range: reflects the temperature at which half of the primer–template duplex dissociates, it depends largely on the primer nucleotide composition, particularly the GC content.
Primer length: As longer primers higher melting temperatures due to more base-pair interactions with the template.
Secondary structures: May require adjustment of the annealing temperature.
Reaction conditions: can alter primer–template stability and thus influence the optimal annealing temperature.
- There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
PCR and restriction digests serve different purposes: PCR amplifies a specific DNA fragment using primers and an enzyme like Taq polymerase, making it ideal when you need a lot of a precise sequence or start with very little DNA. In contrast, restriction enzymes such as EcoRI cut DNA at specific sites, which is useful for cloning or checking constructs. So basically, PCR = amplify, restriction digest = cut, and they’re often used together.
- How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
Primers should be designed to add some bp of overlapping homologous sequences between the insert and the vector. After PCR or digestion, fragments are verified by gel electrophoresis to confirm the correct size and purity, and the overlap regions should be checked in silico, for example by calculating the predicted digest in Benchling to verify the expected band sizes.
- How does the plasmid DNA enter the E. coli cells during transformation?
Heat shock: Generate pores in bacterial cell wall with an abrupt temperature change
Electroporation: Generate pores in bacterial cell wall with high electrical voltage
In both methods the cells are shocked causing the cell membrane to “open up”
- Describe another assembly method in detail
Golden Gate Assembly
Allows the assembly of multiple DNA fragments in a single reaction, using a Type IIS restriction enzyme, which cuts DNA outside of its recognition sequence to create specific overhangs. DNA fragments and the vector are designed so that these overhangs are complementary, ensuring that the fragments assemble in the correct order. During the reaction, the restriction enzyme cuts the DNA while T4 DNA Ligase simultaneously ligates the compatible ends. Because the restriction sites are removed after ligation, the final assembled plasmid cannot be cut again, which improves assembly efficiency. This method is widely used for assembling multiple fragments quickly and accurately in a one-pot reaction.
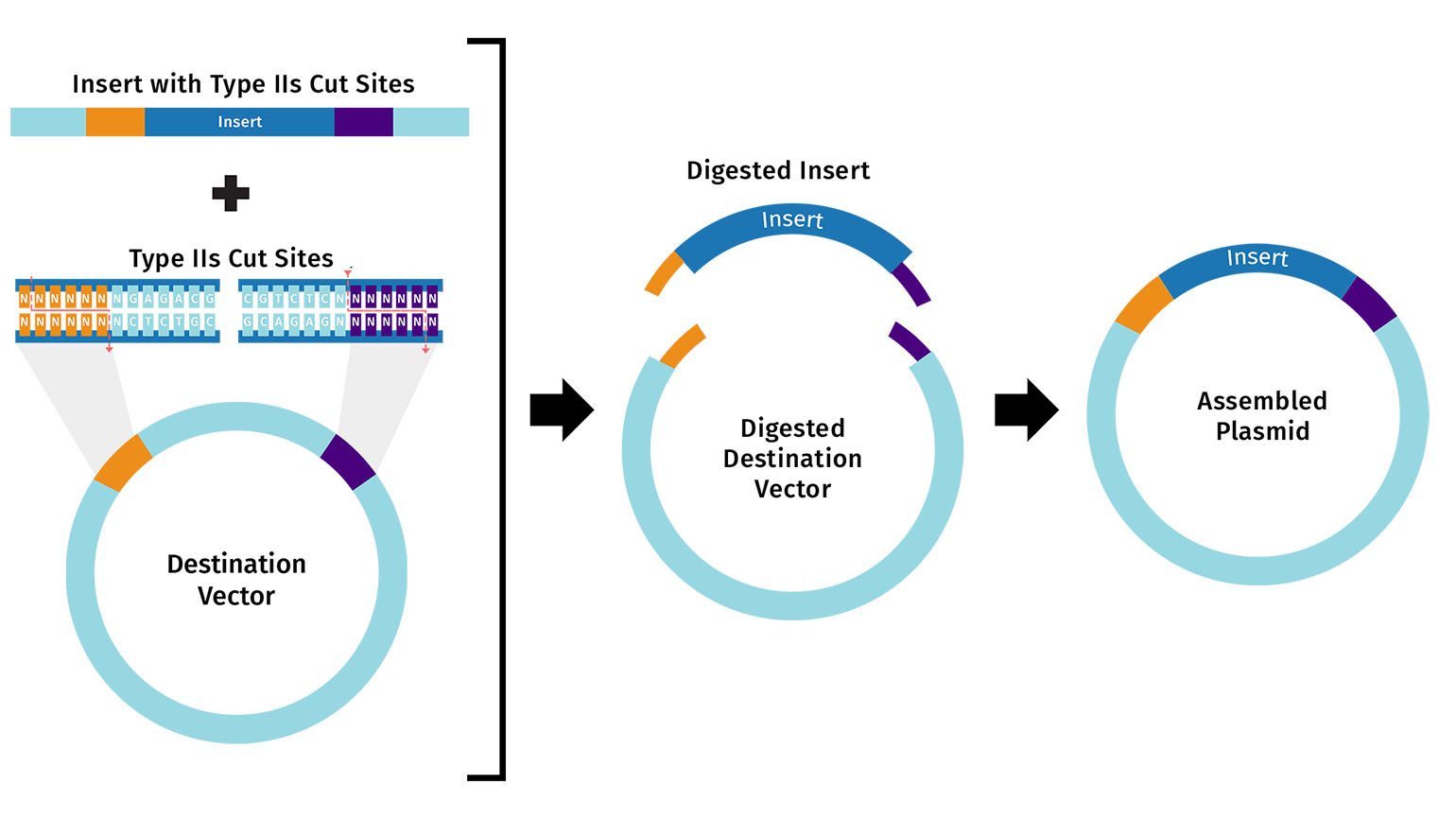
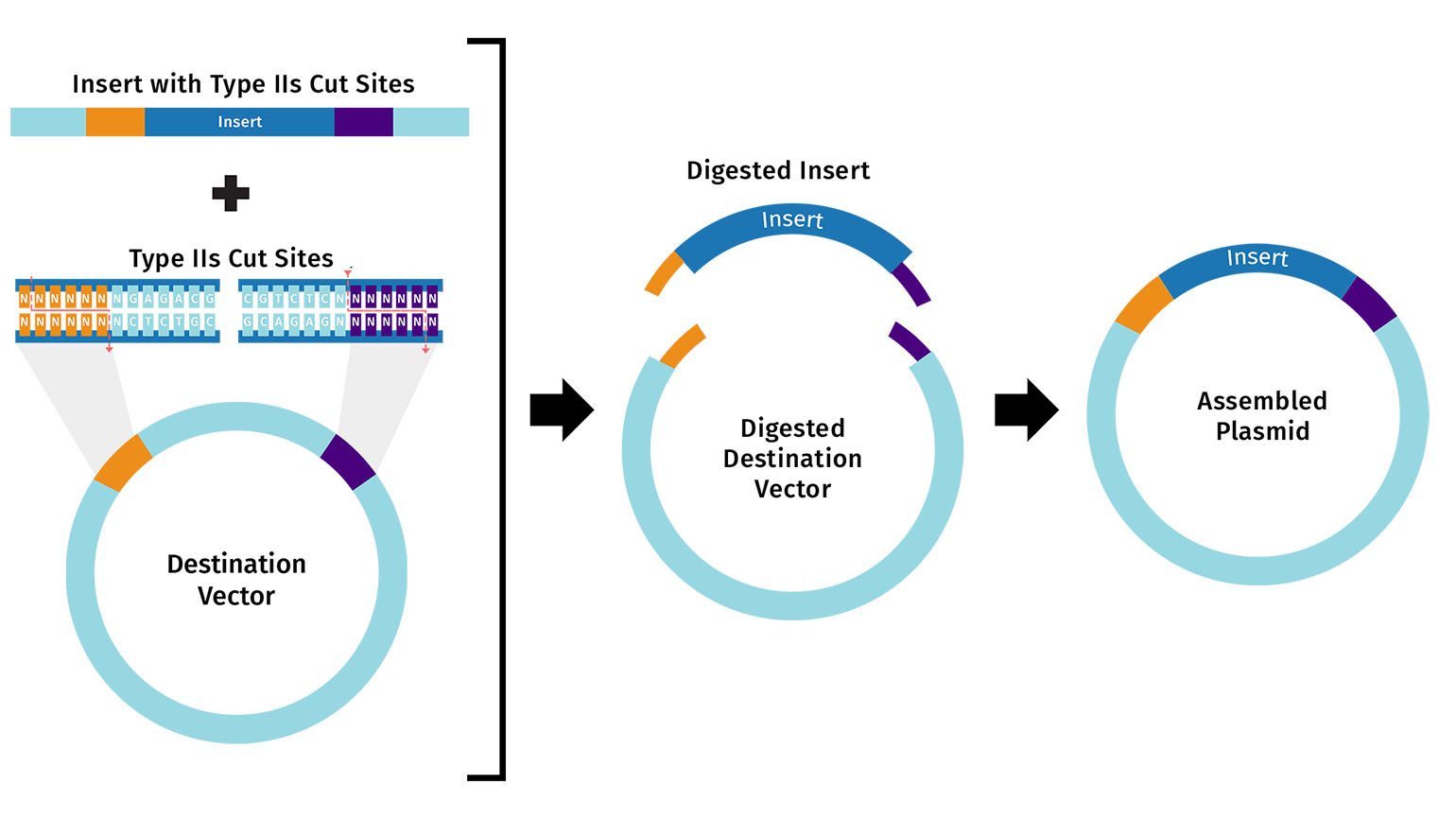
Final project

