Week 10: Advanced Imaging & Measurement Technology
Homework: Final Project
- Please identify at least one (ideally many) aspect(s) of your project that you will measure. It could be the mass or sequence of a protein, the presence, absence, or quantity of a biomarker, etc.
Expression of the Reporter Gene LacZ
- Please describe all of the elements you would like to measure, and furthermore describe how you will perform these measurements
β-galactosidase hydrolyzes chromogenic substrates, producing a colored product that can be visually detected or quantified spectrophotometrically. LacZ will be measured using a colorimetric assay with ONPG, enabling both spectrophotometric quantification and visual detection
- What are the technologies you will use (e.g., gel electrophoresis, DNA sequencing, mass spectrometry, etc.)? Describe in detail
Colorimetric β-Galactosidase Assay To quantify the final output of the biosensor—LacZ expression—which reflects the presence of the DENV pathogen
Homework: Waters Part I — Molecular Weight
- Based on the predicted amino acid sequence of eGFP (see below) and any known modifications, what is the calculated molecular weight?
The molecular weight of eGFP with His-purification tag and a linker is 5.90 / 28006.60
- Calculate the molecular weight of the eGFP using the adjacent charge state approach described in the recitation. Select two charge states from the intact LC-MS data (Figure 1) and:
Peaks: 933.7848 and 903.7844
Determine z for each adjacent pair of peaks (n,n +1) z=30,1
Determine the MW of the protein using the relationship: (Peak 1 and Z) MW=27983,3
Cal culate the accuracy of the measurement using the deconvoluted MW from 2.2 and the predicted weight of the protein from 2.1 Accuracy=8,3x10(4)
- Can you observe the charge state for the zoomed-in peak in the mass spectrum for the intact eGFP? If yes, what is it? If no, why not?
No, the charge state cannot be determined from the enlarged peak, because the peak does not show adjacent peaks or sufficient isotopic resolution to infer the charge
Homework: Waters Part II — Secondary/Tertiary structure
- Based on learnings in the lab, please explain the difference between native and denatured protein conformations. For example, what happens when a protein unfolds? How is that determined with a mass spectrometer? What changes do you see in the mass spectrum between the native and denatured protein analyses (Figure 2)?
Native proteins retain their folded, 3D structure, stabilized by interactions such as hydrogen bonding, hydrophobic interactions, and salt bridges. Denatured proteins have lost this folded structure due to disruption of these interactions. The protein becomes unfolded or extended.
When a protein unfolds: Lys, Arg, His become exposed, the protein accept more protons during electrospray ionization (ESI) and this leads to higher charge states (larger z values)
- Zooming into the native mass spectrum of eGFP from the Waters Xevo G3 QTof MS (see Figure 3), can you discern the charge state of the peak at ~2800 M/Z? What is the charge state? How can you tell?
It’s approximately. z=10
Homework: Waters Part III — Peptide Mapping - primary structure
- How many Lysines (K) and Arginines (R) are in eGFP? Please circle or highlight them in the eGFP sequence given in Waters Part I question 1 above.
26
- How many peptides will be generated from tryptic digestion of eGFP?
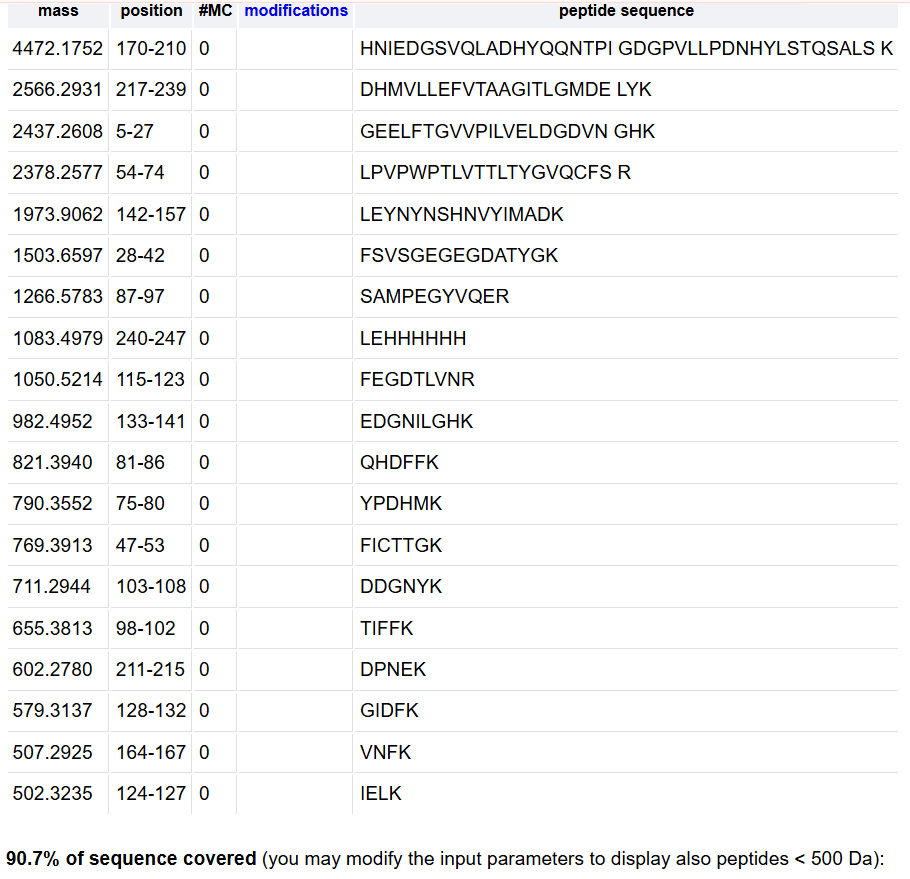

- Based on the LC-MS data for the Peptide Map data generated in lab (please use Figure 5a as a reference) how many chromatographic peaks do you see in the eGFP peptide map between 0.5 and 6 minutes? You may count all peaks that are >10% relative abundance
21
- Assuming all the peaks are peptides, does the number of peaks match the number of peptides predicted from question 2 above? Are there more peaks in the chromatogram or fewer?
Not, there are fewer
- Identify the mass-to-charge (m/z) of the peptide shown in Figure 5b. What is the charge (z) of the most abundant charge state of the peptide (use the separation of the isotopes to determine the charge state).
- The dominant peak cluster is centered at m/z≈525.76
- charge state: z=2+
- Neutral peptide mass: ≈1049.5Da
- Mass of [M + H]+:1050,5 m/z
- dentify the peptide based on comparison to expected masses in the PeptideMass tool. What is mass accuracy of measurement?
- Peptide: DLGEEYVQAFK (GFP)
- Mt= 1049.53 Da
- Mexp= 1049.52 Da
- Error= 9.5 ppm
- What is the percentage of the sequence that is confirmed by peptide mapping? (see Figure 6)
88%
Homework: Waters Part IV — Oligomers
- Identify where the following oligomeric species are on the spectrum shown below from the CDMS
- 7FU Decame (3,4 MDa) -> Peak 3
- 8FU Didecamer (8,3 MDa) -> Peak 5
- 8FU 3-Decamer (12,67 MDa) -> Peak 6
- 8FU 4-Decamer (16 MDa) -> No peak
Homework: Waters Part V — Did I make GFP?
Molecular weight Theoretical = 26.9 kDa Molecular weight Intact LC-MS = 27.0 kDa PPM Mass Error ~3 700 ppm