Week 2 HW: DNA Read, Write & Edit
Part 1: Benchling & In-silico Gel Art
Documentation
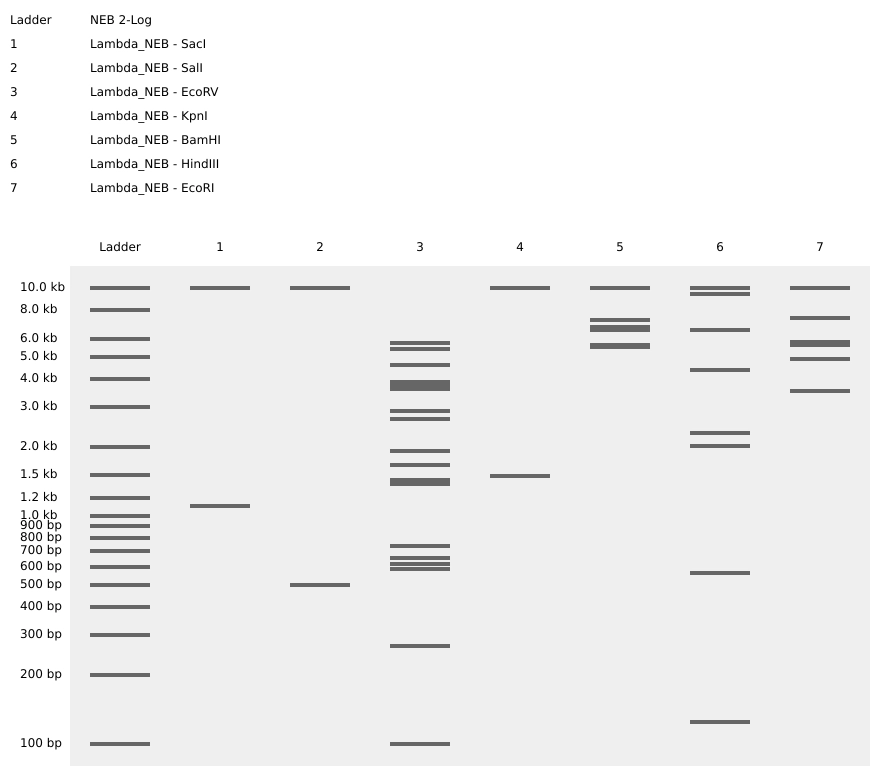


First of all I started by making a digest with a single enzyme at a time.
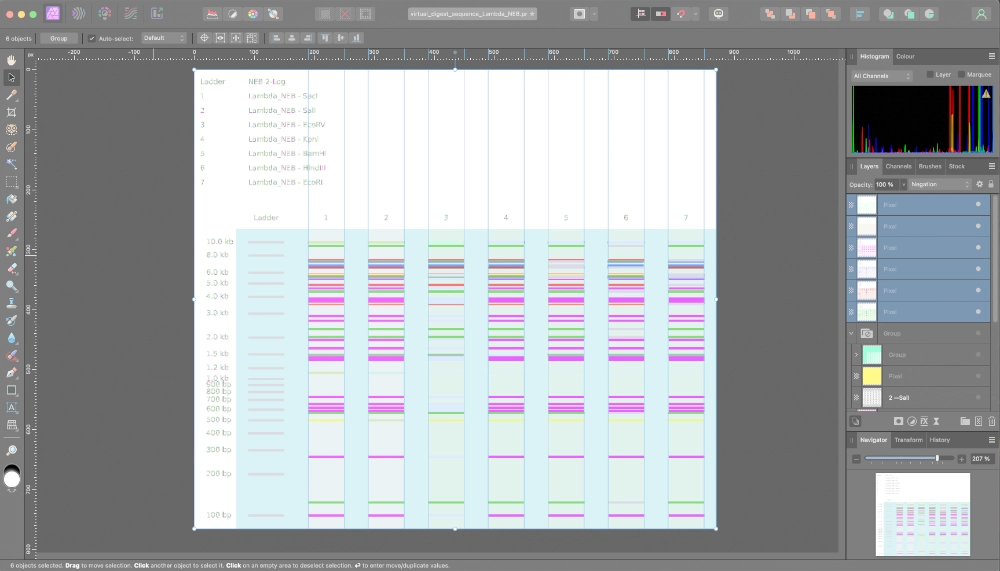

Then tried to color code the result of every enzyme and superimpose them on top of each other so to create a “grid” were I would make my design. I soon understood it would be way too confusing, plus, that the result of using a combination of enzymes doesn’t necessarily correspond to the superimposition of the lines created by each enzyme separately.
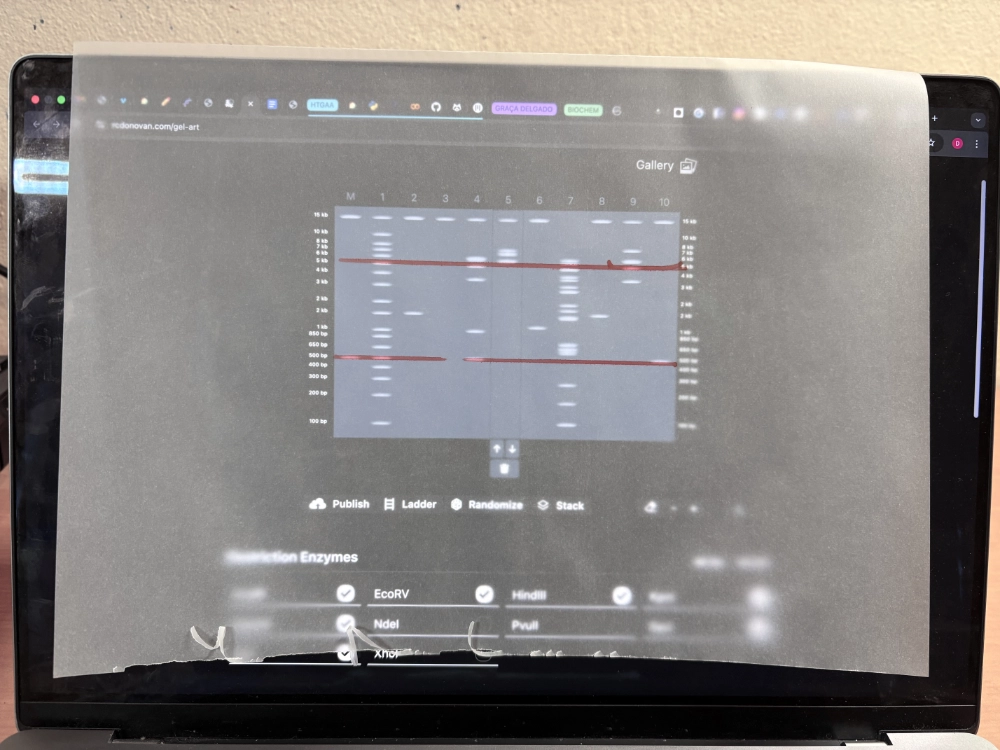

Then I started using Ronan’s website to iterate on combinations of enzymes + using some unconventional techniques.
Final Result — “HTGAA”
.png)
.png)
For the cover image I just edited out some of the space between lines of the same letter to make it more perceptible
- This was a fun exercise that allowed me to visually understand the logic of enzyme’s digests and pay attention to some details I might haved overlooked otherwise.
Part 3: DNA Design Challenge
3.1. Choose your protein
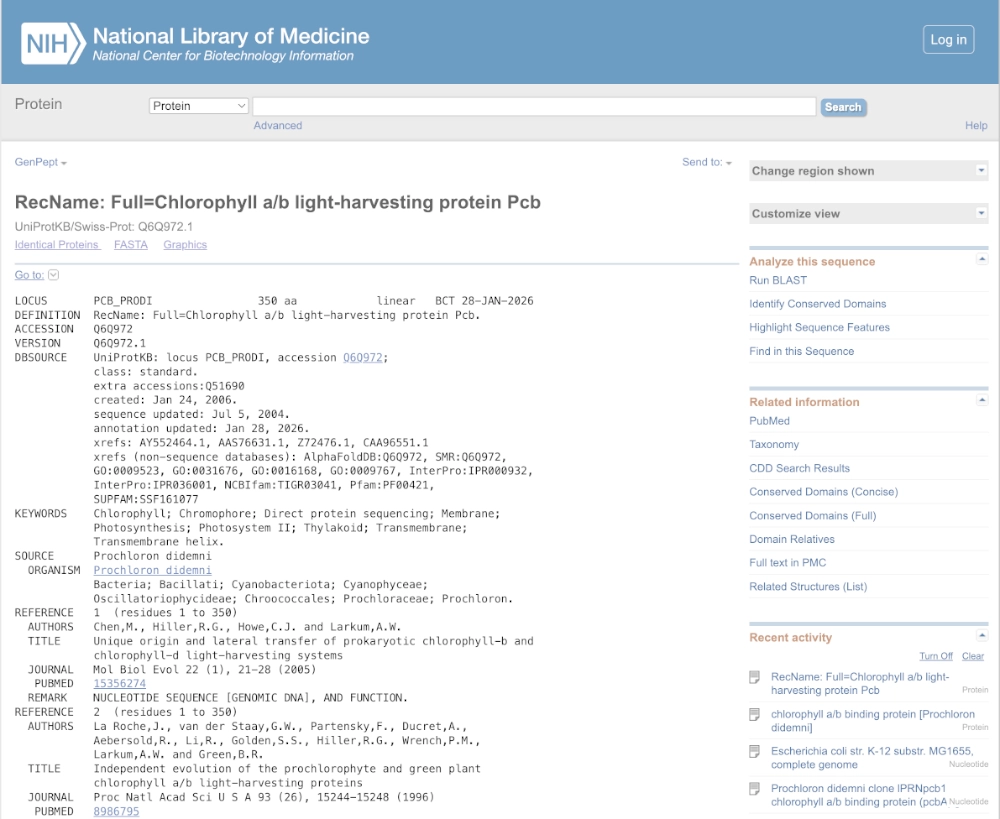
The protein I’d like to work with is the prochlorophyte chlorophyll-binding (Pcb) protein which is the light-harvesting protein (LHP) in prokaryotes that uses only chlorophyll as their photosensitive pigment. A modified version of this protein could be used to efficiently absorb light causing degradation of chlorophyll a, b and d molecules into porphyrin-type derivatives — that can be used to bind iron and create photographic images in a cell-free system.
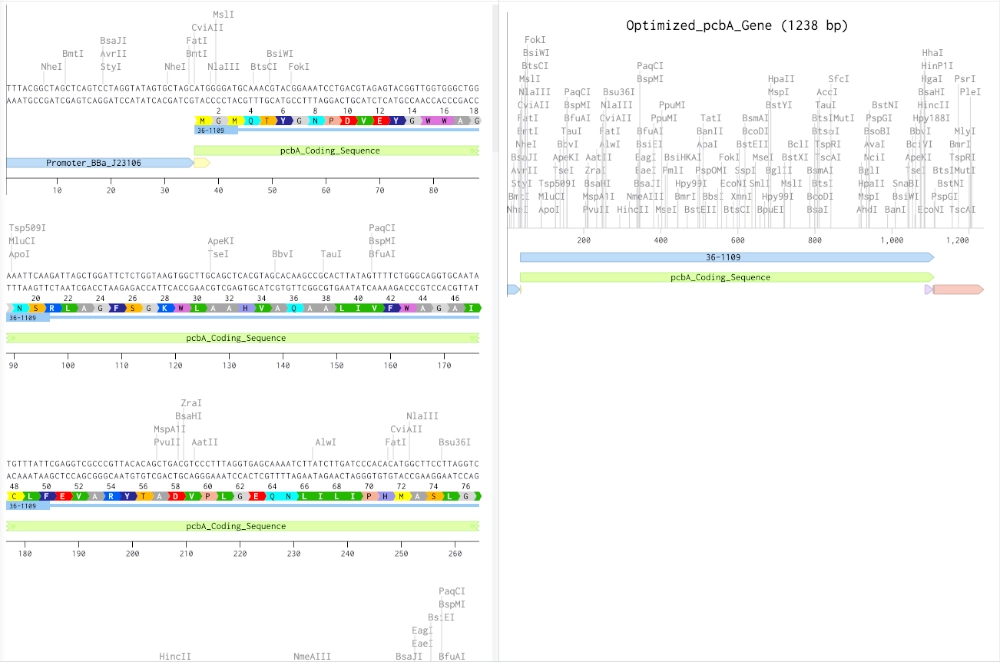
MGMQTYGNPDVEYGWWAGNSRLAGFSGKWLAAHVAQAALIVFWAGAICLFEVARYTADVPLGEQNLILIPHMASLGLGIGEGGQIVDTFPYFAVGVVHLVSSAVIGAGGLYHSLRGPAILKEGPARAPKFDFDWGDGKRLGFILGHHLILLGLGALFLVLWAVFFGIYDPVIGEVRTVTSPTLNPFTIFGYQTHFVETNTLEDLIGGHVYVAIIEISGGLWHIFCPPFKWAQRLIIYSGEGLLAYALGGLAIMGFTAAVYCAFNTLAYPVEFYGPPLDFRFSFAPYFIDTADLPSGQYTARAWLCNVHFFLAFFVLQGHLWHALRTLGFDFKRIPAALGSLSEDVVDAKA(from NCBI chlorophyll a/b binding protein [Prochloron didemni])

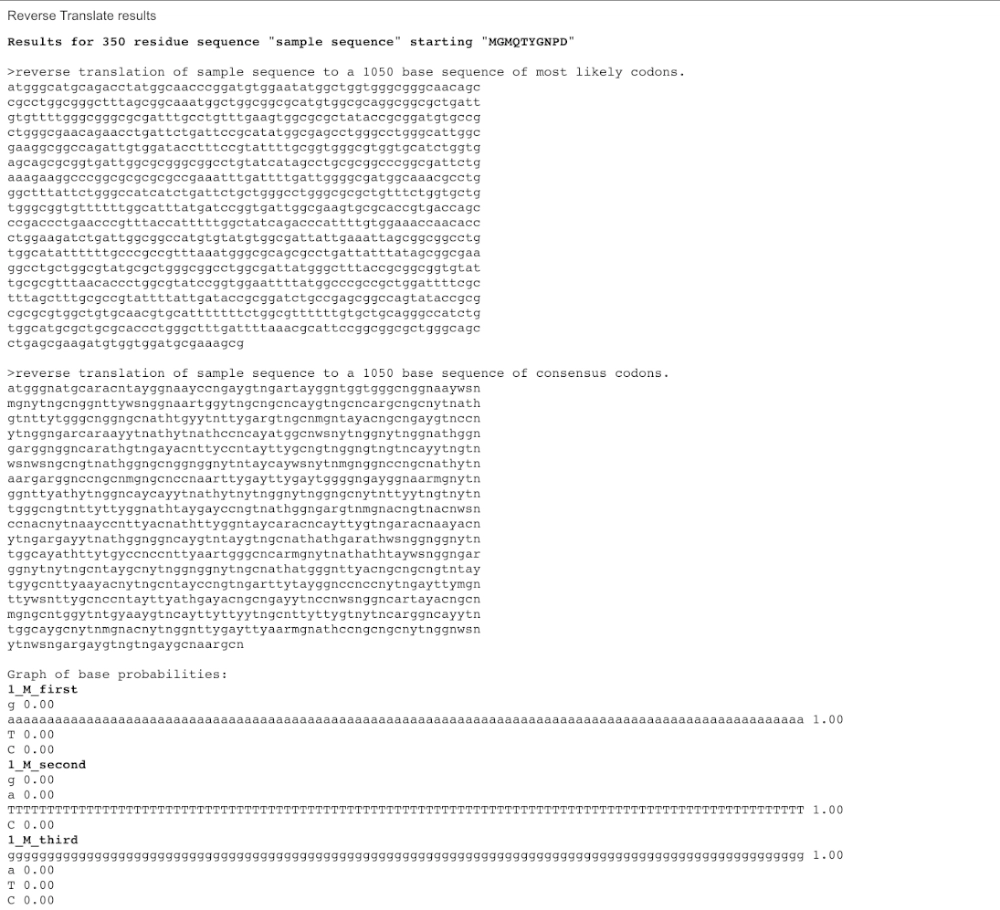
3.2. Reverse Translate — AA to DNA
ATGGGCATGCAGACCTATGGCAACCCGGATGTGGAATATGGCTGGTGGGCGGGCAACAGCCGCCTGGCGGGCTTTAGCGGCAAATGGCTGGCGGCGCATGTGGCGCAGGCGGCGCTGATTGTGTTTTGGGCGGGCGCGATTTGCCTGTTTGAAGTGGCGCGCTATACCGCGGATGTGCCGCTGGGCGAACAGAACCTGATTCTGATTCCGCATATGGCGAGCCTGGGCCTGGGCATTGGCGAAGGCGGCCAGATTGTGGATACCTTTCCGTATTTTGCGGTGGGCGTGGTGCATCTGGTGAGCAGCGCGGTGATTGGCGCGGGCGGCCTGTATCATAGCCTGCGCGGCCCGGCGATTCTGAAAGAAGGCCCGGCGCGCGCGCCGAAATTTGATTTTGATTGGGGCGATGGCAAACGCCTGGGCTTTATTCTGGGCCATCATCTGATTCTGCTGGGCCTGGGCGCGCTGTTTCTGGTGCTGTGGGCGGTGTTTTTTGGCATTTATGATCCGGTGATTGGCGAAGTGCGCACCGTGACCAGCCCGACCCTGAACCCGTTTACCATTTTTGGCTATCAGACCCATTTTGTGGAAACCAACACCCTGGAAGATCTGATTGGCGGCCATGTGTATGTGGCGATTATTGAAATTAGCGGCGGCCTGTGGCATATTTTTTGCCCGCCGTTTAAATGGGCGCAGCGCCTGATTATTTATAGCGGCGAAGGCCTGCTGGCGTATGCGCTGGGCGGCCTGGCGATTATGGGCTTTACCGCGGCGGTGTATTGCGCGTTTAACACCCTGGCGTATCCGGTGGAATTTTATGGCCCGCCGCTGGATTTTCGCTTTAGCTTTGCGCCGTATTTTATTGATACCGCGGATCTGCCGAGCGGCCAGTATACCGCGCGCGCGTGGCTGTGCAACGTGCATTTTTTTCTGGCGTTTTTTGTGCTGCAGGGCCATCTGTGGCATGCGCTGCGCACCCTGGGCTTTGATTTTAAACGCATTCCGGCGGCGCTGGGCAGCCTGAGCGAAGATGTGGTGGATGCGAAAGCGTAA(converted using bioinformatics.org)

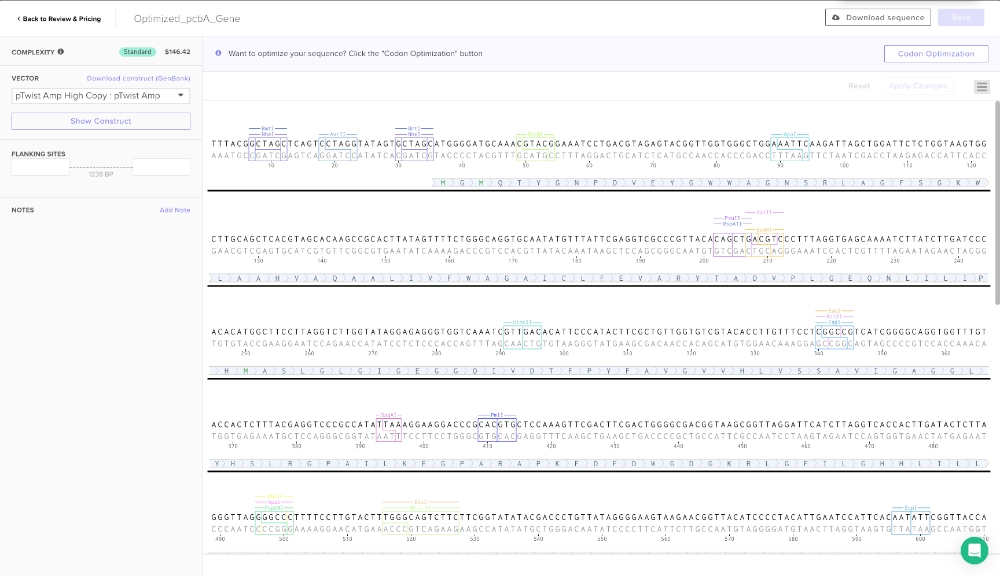
3.3. Codon optimization.
Codon optimization is an important process due to different organisms having and producing different amino acids in different proportions. So, if a gene codes for a rare amino acid, it might slow the translation process and therefore the folding of the protein and might even render the protein non-functional. In this case, the gene should be optimized for e. coli which is probably the best choice since the primary objective is to express a protein that is going to be used in a cell-free system and it is the simplest organism to work with. For this codon optimization I avoided Type IIS enzyme recognition sites for BsaI, BsmBI, and BbsI — these are some enzymes that are useful for ligation with plasmid backone.
ATGGGGATGCAAACGTACGGAAATCCTGACGTAGAGTACGGTTGGTGGGCTGGAAATTCAAGATTAGCTGGATTCTCTGGTAAGTGGCTTGCAGCTCACGTAGCACAAGCCGCACTTATAGTTTTCTGGGCAGGTGCAATATGTTTATTCGAGGTCGCCCGTTACACAGCTGACGTCCCTTTAGGTGAGCAAAATCTTATCTTGATCCCACACATGGCTTCCTTAGGTCTTGGTATAGGAGAGGGTGGTCAAATCGTTGACACATTCCCATACTTCGCTGTTGGTGTCGTACACCTTGTTTCCTCGGCCGTCATCGGGGCAGGTGGTTTGTACCACTCTTTACGAGGTCCCGCCATATTAAAGGAAGGACCCGCACGTGCTCCAAAGTTCGACTTCGACTGGGGCGACGGTAAGCGGTTAGGATTCATCTTAGGTCACCACTTGATACTCTTAGGGTTAGGGGCCCTTTTCCTTGTACTTTGGGCAGTCTTCTTCGGTATATACGACCCTGTTATAGGGGAAGTAAGAACGGTTACATCCCCTACATTGAATCCATTCACAATATTCGGTTACCAAACTCACTTCGTAGAGACTAATACGCTTGAGGACTTAATCGGTGGTCACGTTTACGTCGCCATCATCGAGATCTCCGGCGGGTTGTGGCACATCTTCTGTCCCCCATTCAAGTGGGCACAACGATTGATCATATACTCAGGTGAGGGGTTGCTTGCCTACGCATTGGGTGGTCTCGCTATAATGGGTTTCACTGCCGCAGTCTACTGTGCCTTCAATACGCTTGCCTACCCTGTAGAGTTCTACGGTCCACCTTTAGACTTCCGTTTCTCATTCGCACCATACTTCATCGACACAGCCGACTTGCCGTCCGGGCAATACACAGCCCGAGCCTGGTTGTGTAATGTTCACTTCTTCTTAGCTTTCTTCGTATTGCAAGGGCACCTTTGGCACGCATTACGTACGCTTGGTTTCGACTTCAAGCGTATCCCCGCAGCATTAGGTTCCCTCTCTGAGGACGTTGTTGACGCCAAAGCGTAA(made with Codon Optimization Tool | Twist Bioscience)

3.4. You have a sequence! Now what?
This gene sequence could be synthesized through chemical synthesis on silicon chips and assembled into a vector backbone— bacterial plasmid— then, put into e. coli. This allows for the use of the bacteria’s own cellular machinery to first transcribe this DNA sequence into mRNA (which would be identical to this coding DNA strand, except for “U”), and finally the bacteria’s ribosomes would translate the resulting mRNA into the final amino acid sequence. This AA sequence would be the pcbA protein in its apoprotein form— since e. coli lacks the machinery to produce chlorophyll molecules that play as a co-factor in the folding of this protein— which would be later combined with chlorophyll extract to render it functional.
3.5. How does it work in nature/biological systems?
1. Describe how a single gene codes for multiple proteins at the transcriptional level.
In biological systems, a single gene can code for multiple proteins at the transcriptional level through the process of alternative splicing (in eukaryotes)— a process where different combinations of exons from the same pre-mRNA molecule are joined together. This process happens inside the nucleus during the processing of pre-mRNA, leading to the synthesis of multiple protein isoforms, which are related forms of the same protein, but with different structural or functional properties. Another process that allows for a single gene to code for multiple proteins (both in eukaryotes and prokaryotes) is the action of alternative promoter genes that create different initiation sites, affecting which exons are included in transcription.
2. Try aligning the DNA sequence, the transcribed RNA, and also the resulting translated Protein!!!
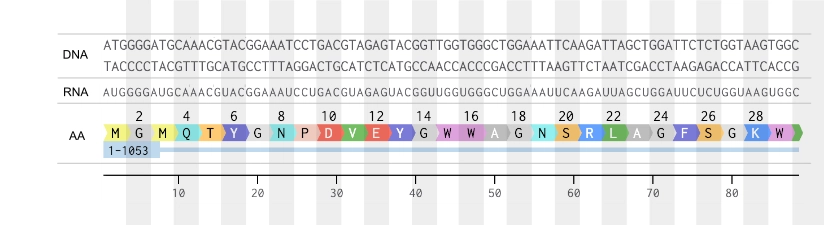

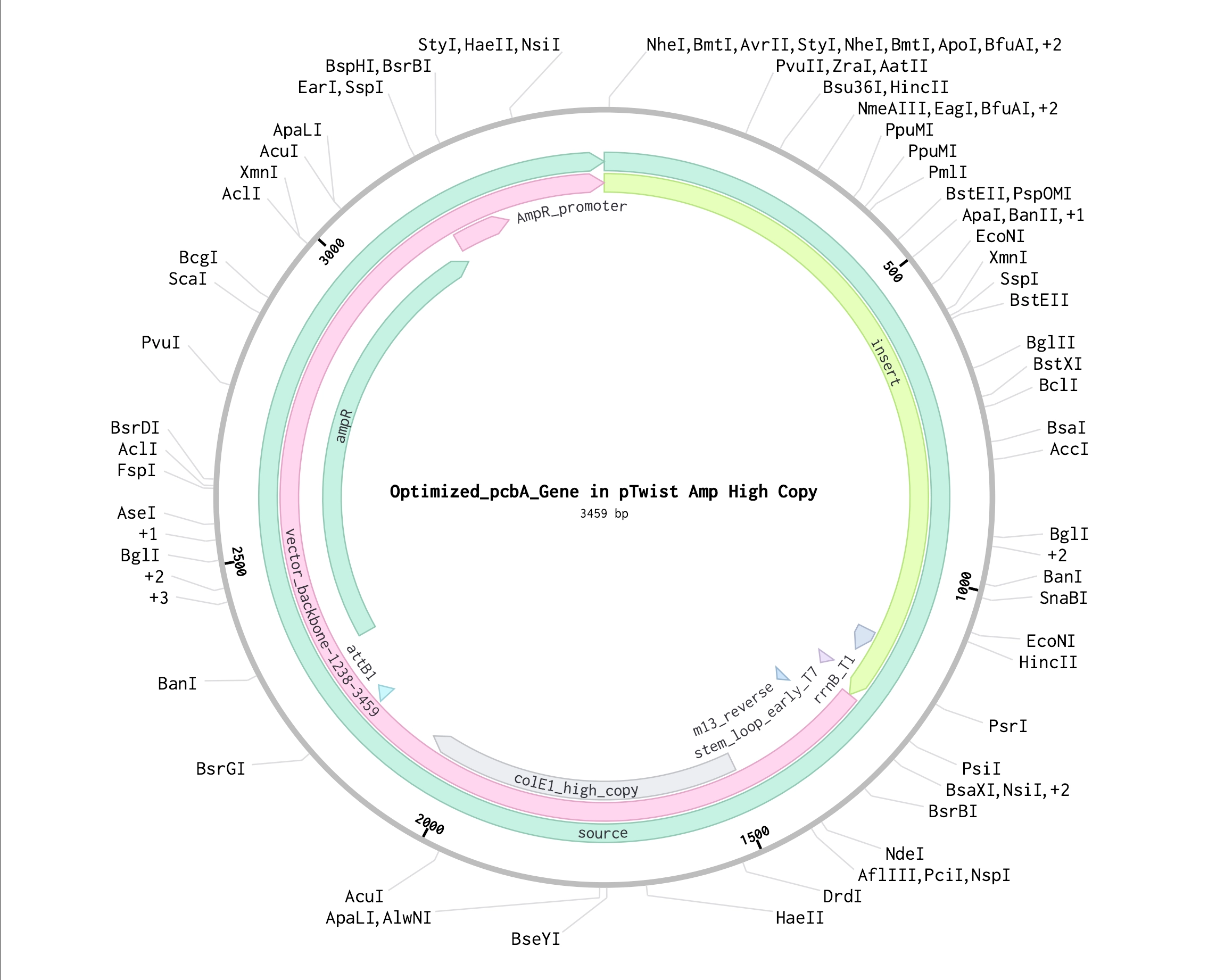
Part 4: Prepare a Twist DNA Synthesis Order
Part 5: DNA Read/Write/Edit
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why?
The DNA I’m interested in sequencing and further understanding is cyanobacteria’s genes for the production of Chlorophyll LHP (Light-Harvesting proteins) which is the nature’s way of organizing chlorophyll molecules in order to get the most light absorption out of them. This biological way of organizing light sensitive pigments could be the answer to a new generation of analog photography media and can also be used to engineer more efficient solar cells to produce energy.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
For this type of application, the most adequate sequencing technology would probably be Sanger sequencing using a device like Sanger-ABI, a 1st generation technology that has been around since 1977 but would be more than enough for things like reading single protein coding sequences. This would be a small-scale project needing only to analyze relatively small nucleotide sequences, it wouldn’t demand the comparison of more complex genes like comparing/analyzing whole genomes. For this method I would make a DNA extraction from cyanobacteria cells and purify it, followed by designing primers specific for the sequence I want to analyze and amplify it through PCR, then remove excess primers and dNTPs. Next step would be to perform a cycle sequencing reaction “chain terminator PCR” using single primers, DNA polymerases, dNTPs and fluorescently labeled ddNTPs— these fluorescent ddNTPs act as chain terminators, stopping synthesis randomly at every possible length to create labeled fragments. Clean up residual dye labeled ddNTPs to prevent noise during read and submit these tagged fragments to Sanger-ABI capillary electrophoresis, which separates the fragments by length and then makes the read by exciting the label of each fragment and detecting the color emitted— then using software to translate the fluorescence signals into a chromatogram, revealing the sequence of the DNA sample.
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why?
I would need to synthesize a bacterial plasmid (for e. coli) with the insert for the Chlorophyll LHP gene, in this case the PcbA protein. The application for this would be to develop a way to keep these proteins functioning in a cell free system in order to create a novel biomaterial that would serve as photographic emulsion for analog film.
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
Probably the most affordable and effective way to synthesize this kind of DNA would be the through clonal gene chip-based chemical synthesis and assembly of the DNA sequence into a plasmid vector through Golden Gate Assembly using type IIS restriction enzymes and T4 DNA ligase. Due to the small-scale nature of this project and standard difficulty of synthesizing this kind of DNA, I don’t think there would be significant limitations speed, accuracy and scalability wise.
5.3 DNA Edit
(i) What DNA would you want to edit and why?
For the objective of this project the DNA that would be interesting to edit would range from the genes coding for the chlorophyll synthesis pathway— in order to develop a modified version of chlorophyl that would be more optimized for photographic purposes— and the sequences coding for the LHP if there is the need to modify the natural occurring proteins, either by decreasing the protection these proteins confer against chlorophyll degradation or by potentially improving on their ability to maximize chlorophyll’s light absorption qualities.
(ii) What technology or technologies would you use to perform these DNA edits and why?
CRISPR-based genome editing would probably be the best choice for this purpose since Chlorophyll biosynthesis involves multiple genes, often with regulatory fine-tuning rather than simple on/off behavior. CRISPR systems allow gene-specific, locus-specific edits, making them well suited for altering enzyme functionality in the chlorophyll synthesis pathway; modifying regulatory regions that affect pigment ratios (e.g. chlorophyll a, b or d) and engineering specific amino-acid changes in LHPs. CRISPR edits DNA by using an RNA guide to bring a DNA-cutting enzyme (nuclease) to a specific genomic site which is cut and where the cell’s own DNA-repair machinery makes the final change utilizing a DNA template (single stranded or double stranded for larger edits) which is delivered together with the Cas9 enzyme. For this end, the first steps would be to define an objective precisely like make chlorophyll more sensitive to light or more prone to degradation under certain conditions, or reduce photoprotective quenching of the LHP and identify genes, regulatory regions or domains that are relevant for those functions. After that, decide what type of edit strategy is needed for a specific site (gene knockout, single or few nucleotide changes or sequence replacement) and design a guide RNA to bind to the target DNA of that specific site, and if a sequence rewrite is needed, design the DNA template for that repair which must have homologous endings that match those surrounding the cut site. To perform the actual edit the gRNA, Cas9 nuclease and template DNA are combined into a plasmid vector or ribonucleoprotein and introduced into the cells via heat shock, electroporation or lipofection. The limitations of this method might include off-target effects by binding and editing similar but not the exact sites; in a cell culture it might happen that not all cells be edited resulting in a mixture of edited and unedited cells, which can make it difficult to achieve a uniform result, and the HDR might have a low efficiency compared to the non-homologous end joining (NHEJ) pathway, not resulting in the intended edit.
.webp)