Week 3 HW: Lab Automation


Assignment: Python Script for Opentrons Artwork
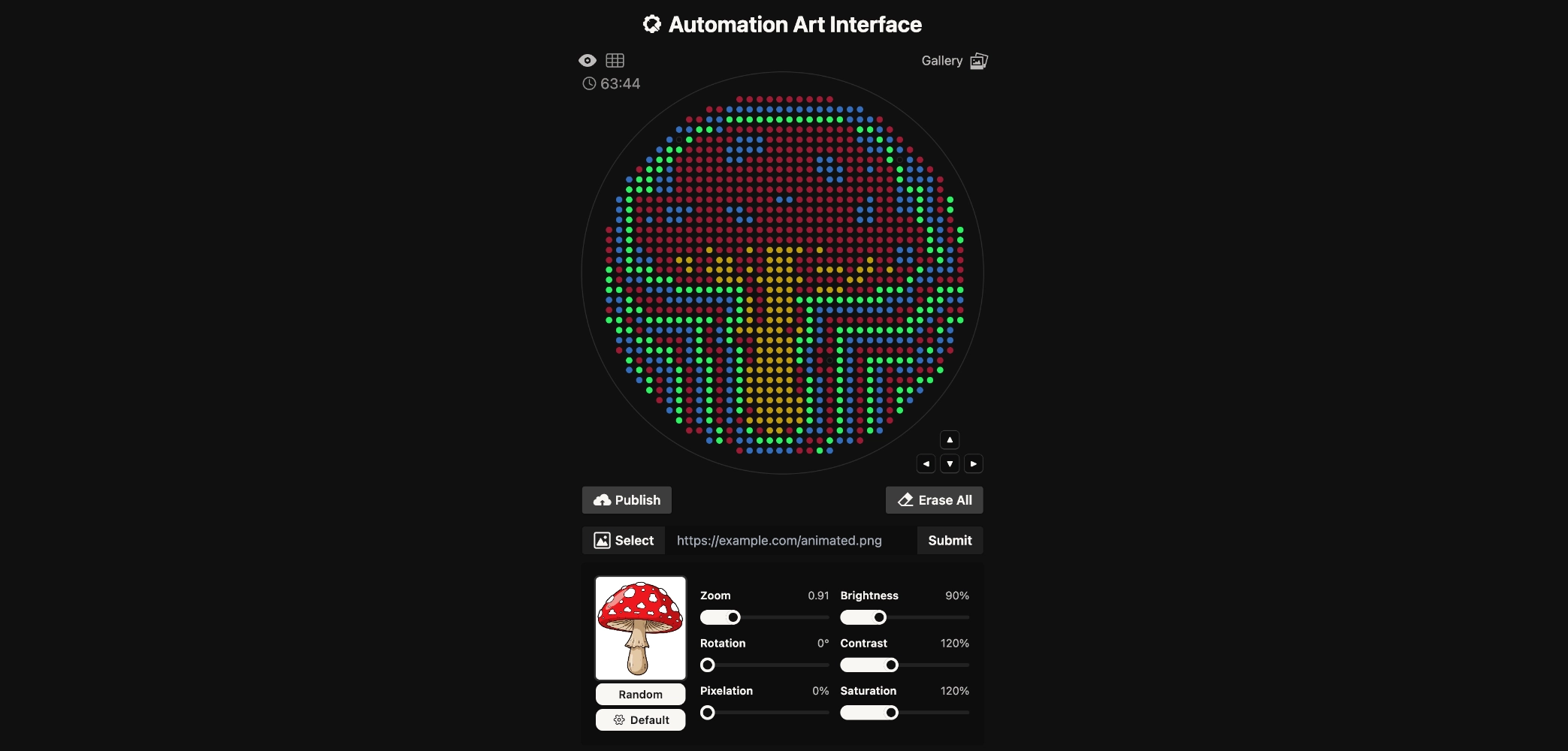
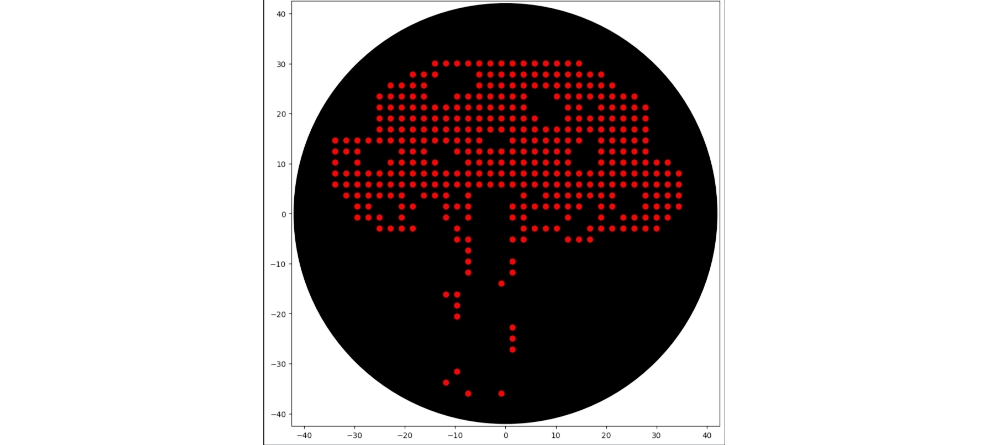
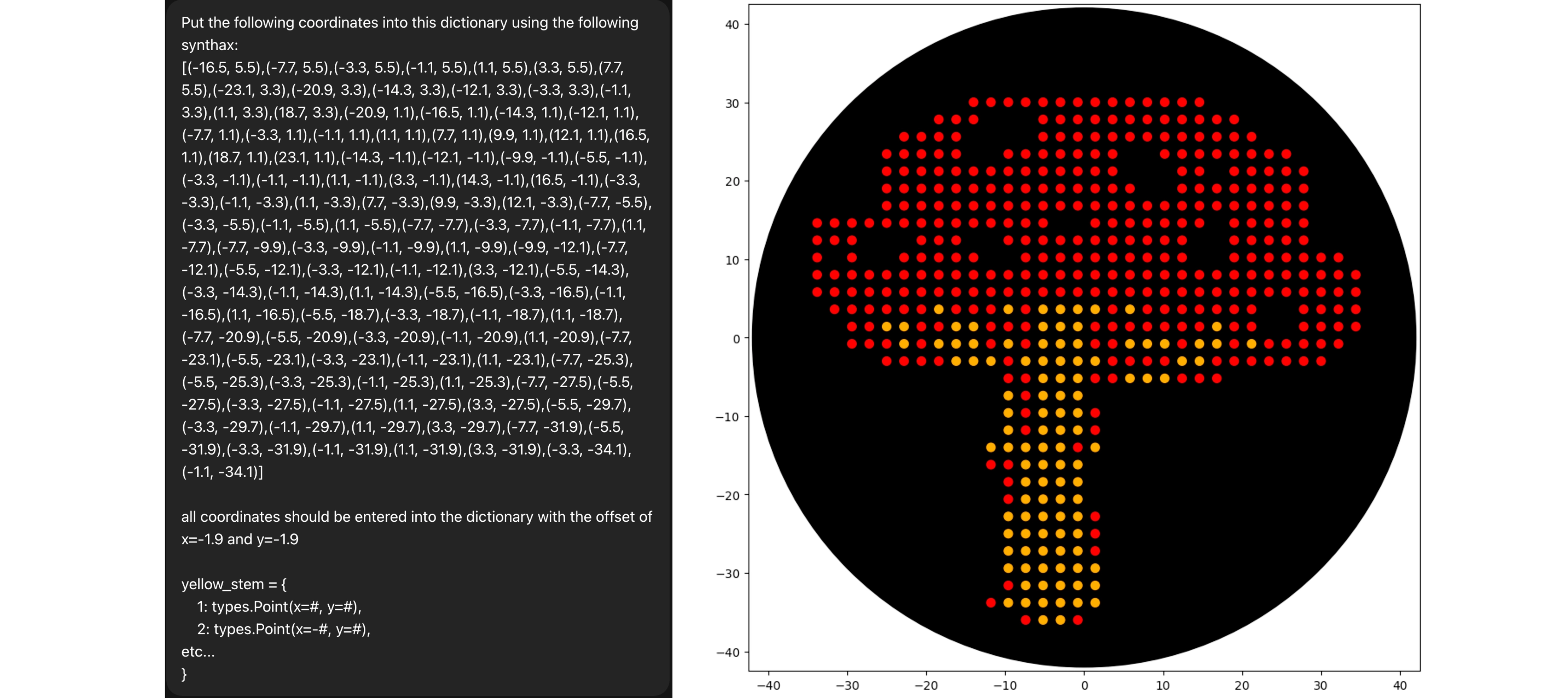
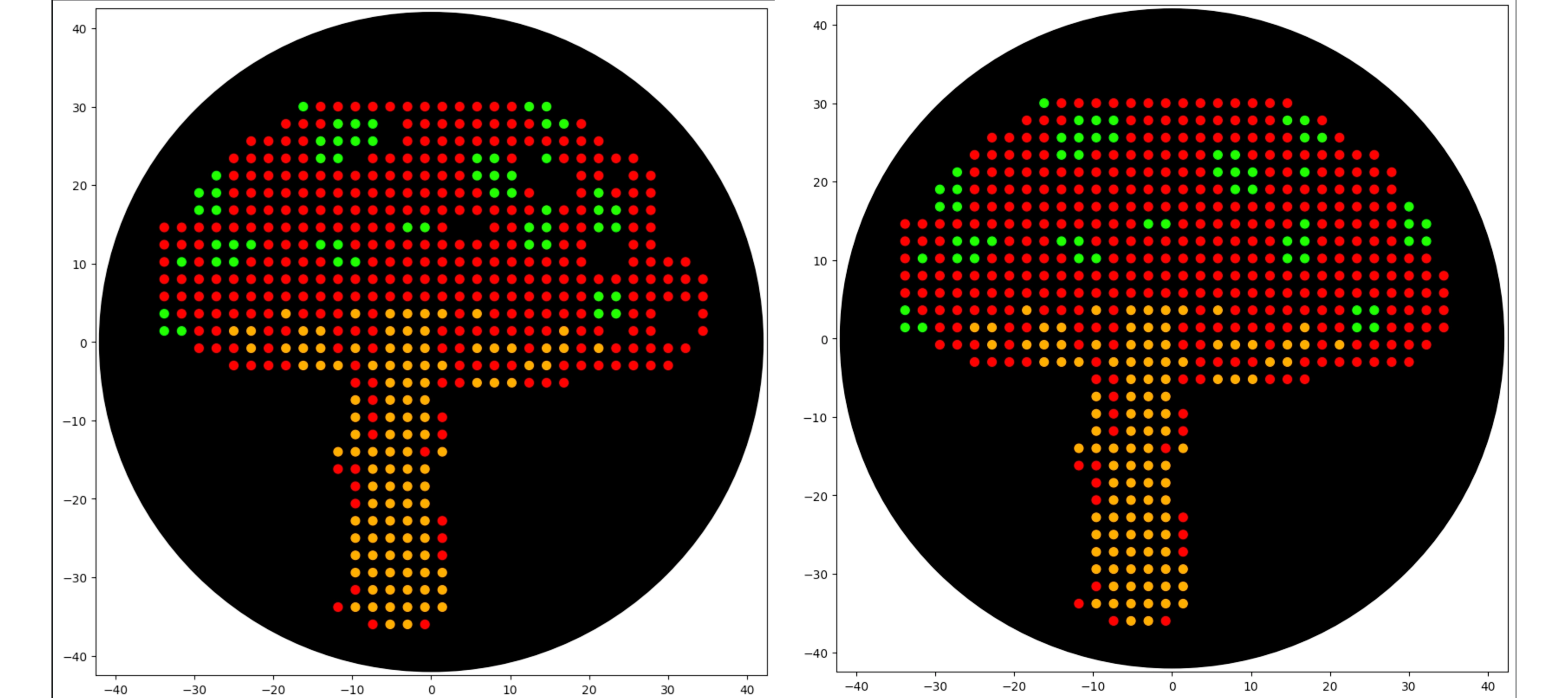
Final Result
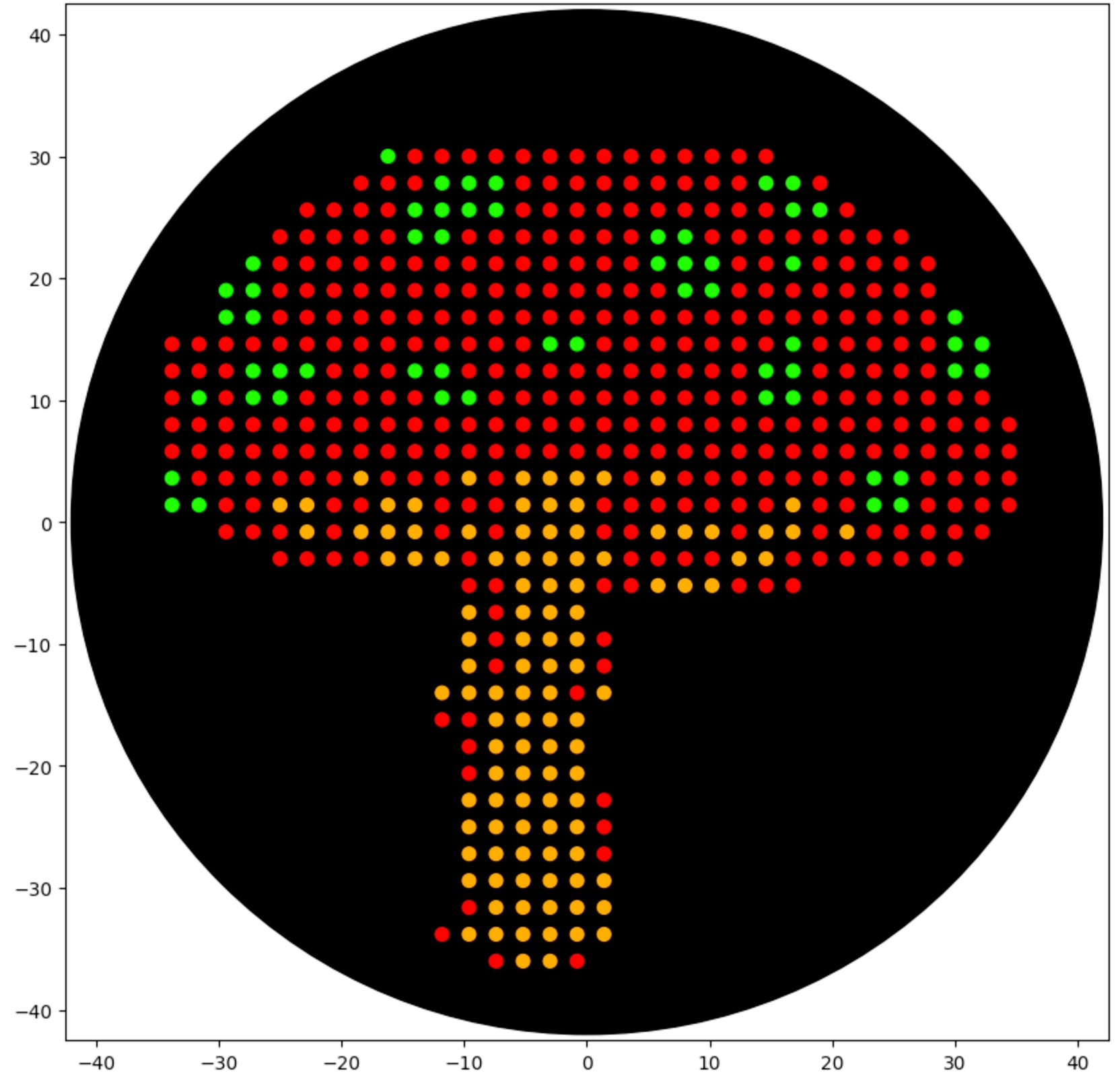

Post-Lab Questions
1. Find and describe a published paper
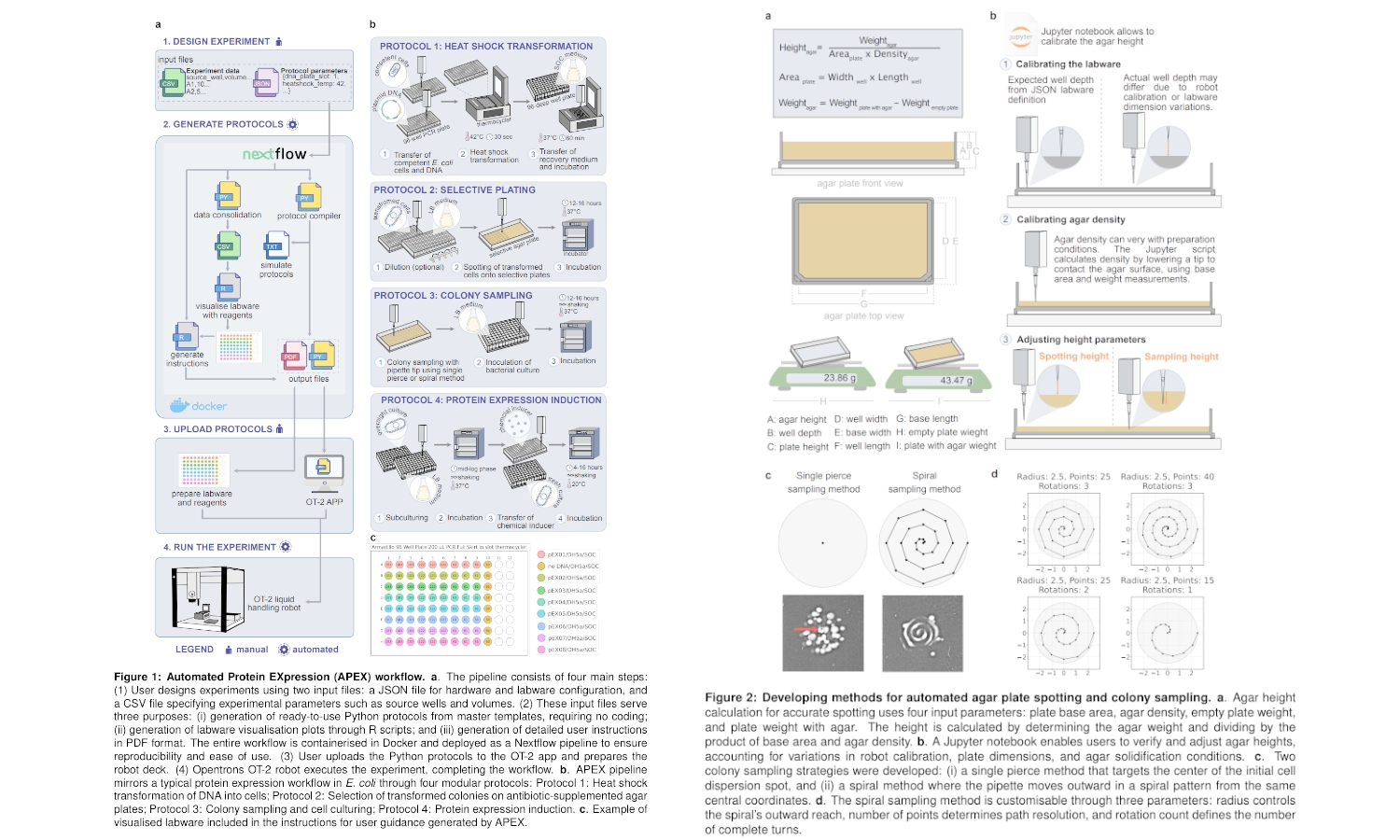
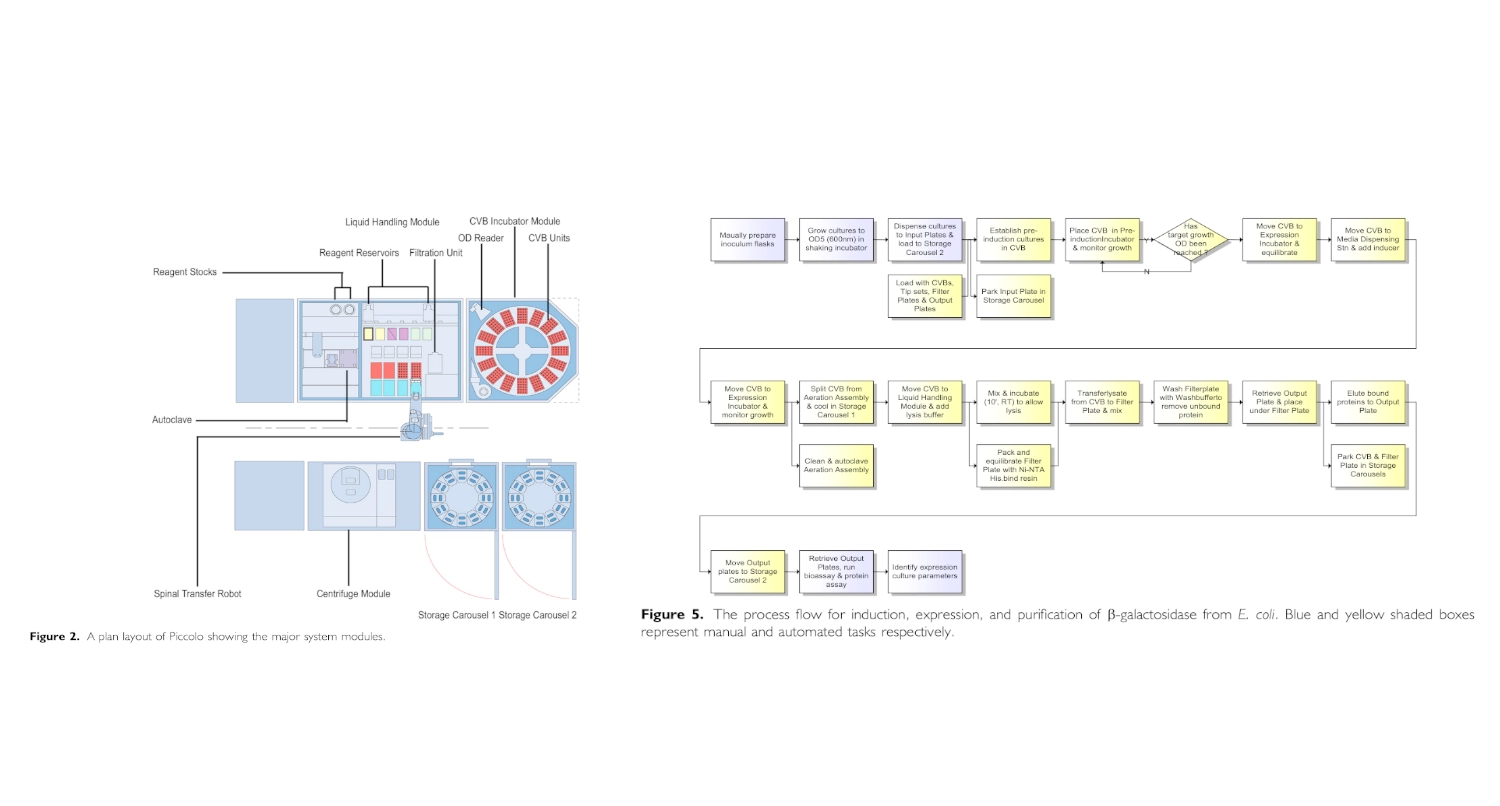
I found and was interested in two articles: The first one “Automation and Optimization of Protein Expression and Purification on a Novel Robotic Platform” published by Journal of Laboratory Automation (October 2006) that describes an automated robotic system for expression and purification of recombinant proteins grown both in E. coli and other bacterial cells and eukaryotic cells. The second “APEX: Automated Protein EXpression in Escherichia coli” published by ACS Synthetic Biology (September 2, 2025) describes an automated pipeline for recombinant protein production in E. coli, leveraging the open-source Opentrons OT-2 platform to handle microbe culturing and protein expression.

- Although this paper might be outdated since it was published 20 years ago it helped me better understand automation of experimentation in living cells.
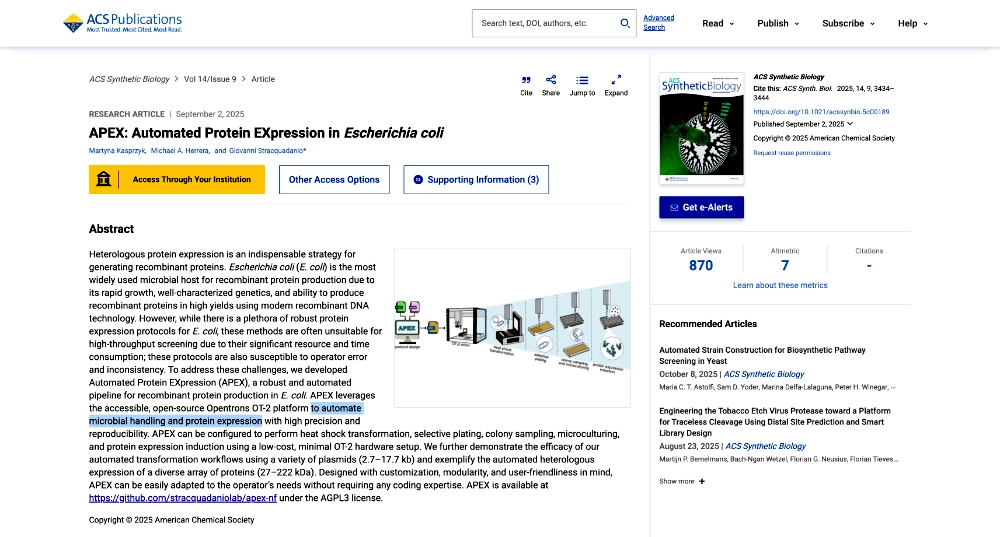

- Write a description about what you intend to do with automation tools for your final project.
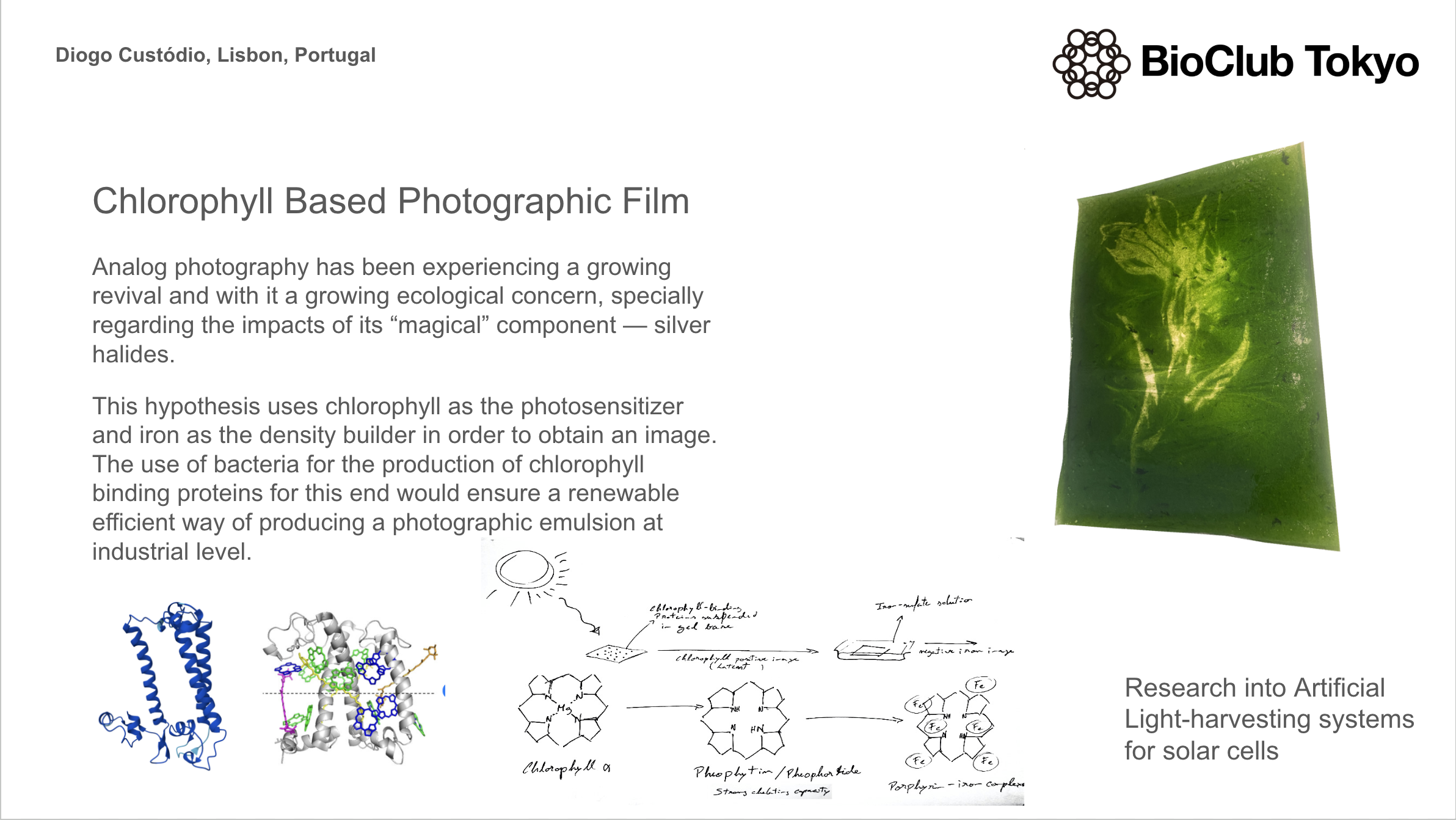
The development on photographic emulsions has an added difficulty of them being light-sensitive. So having an automated workflow to produce iterations of different possibilities would largely make the process more efficient
- Automation for culturing of e coli and synthesis of an array of modified and non-modified chlorophyll binding proteins.
- Lipid-induced folding of those different proteins by combining with chlorophyll extract
- Combination of the different final proteins with an agarose base to allow dispersion onto a base surface
- 3d printed holder for the base supports for the chlorophyll protein emulsion to be dispersed on
- Drying of the emulsion in dark conditions
For the purpose of this exercise, I tried to create a comprehensible workflow for the following operations that I think would be essential for this project:
Workflow for expression of chlorophyll-binding proteins
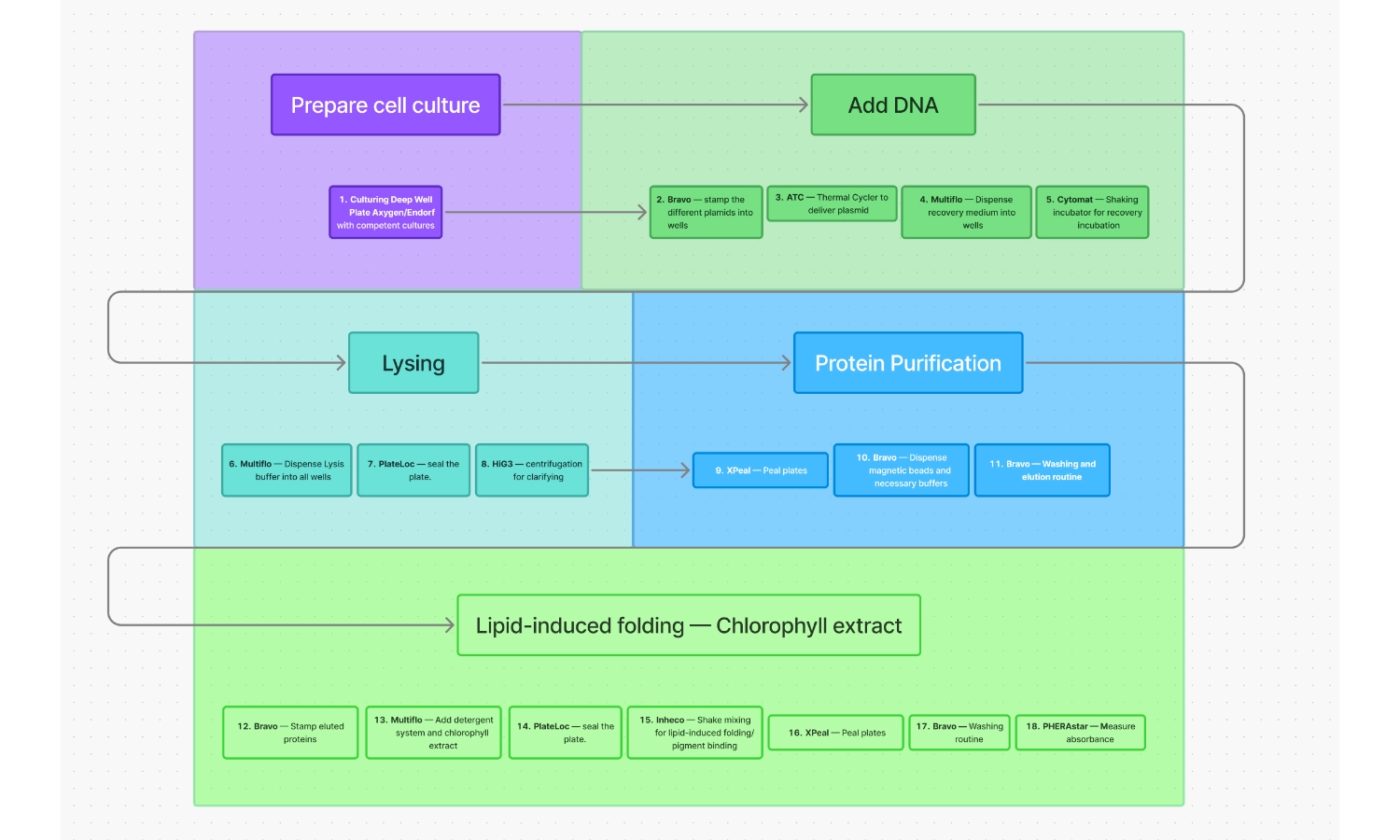
- Culturing Deep Well Plate Axygen/Endorf with competent cultures
- Bravo — Stamp the different plasmids into wells
- ATC — Thermal Cycler to deliver plasmid via thermo shock
- Multiflo — Dispense recovery medium into wells
- Cytomat — Shaking incubator for recovery incubation
- Multiflo — Dispense Lysis buffer into all wells
- PlateLoc — Seal the plate
- HiG3 — Centrifugation for clarifying
- XPeal — Peal plates
- Bravo — Dispense magnetic beads and necessary buffers
- Bravo — Washing and elution routine
- Bravo — Stamp eluted proteins
- Multiflo — Add detergent system and chlorophyll extract
- PlateLoc — Seal the plate
- Inheco — Shake mixing for lipid-induced folding/pigment binding
- XPeal — Peal plates
- Bravo — Washing routine
- PHERAstar — Measure absorbance
- After completing this part of the homework, I realized that CFPS might be a better bet, using an automated system, for the expression and testing of this particular kind of protein since the open system nature of this method would allow the direct addition of chlorophyll into the reaction mixture and allow for instant protein folding. Further along the progression of this project, when the chemistry of the proteins is optimized, living cells — like cyanobacteria — could be used to express them in larger quantities
3D printing of a holder for base supports for the chlorophyll emulsion For the chlorophyll proteins (suspended in some kind of gelling agent like agar) to be dispersed on, so field tests could be performed with pinhole cameras (for the testing of an array of photographic emulsion iterations). Bellow a quick sketch of what it could be.
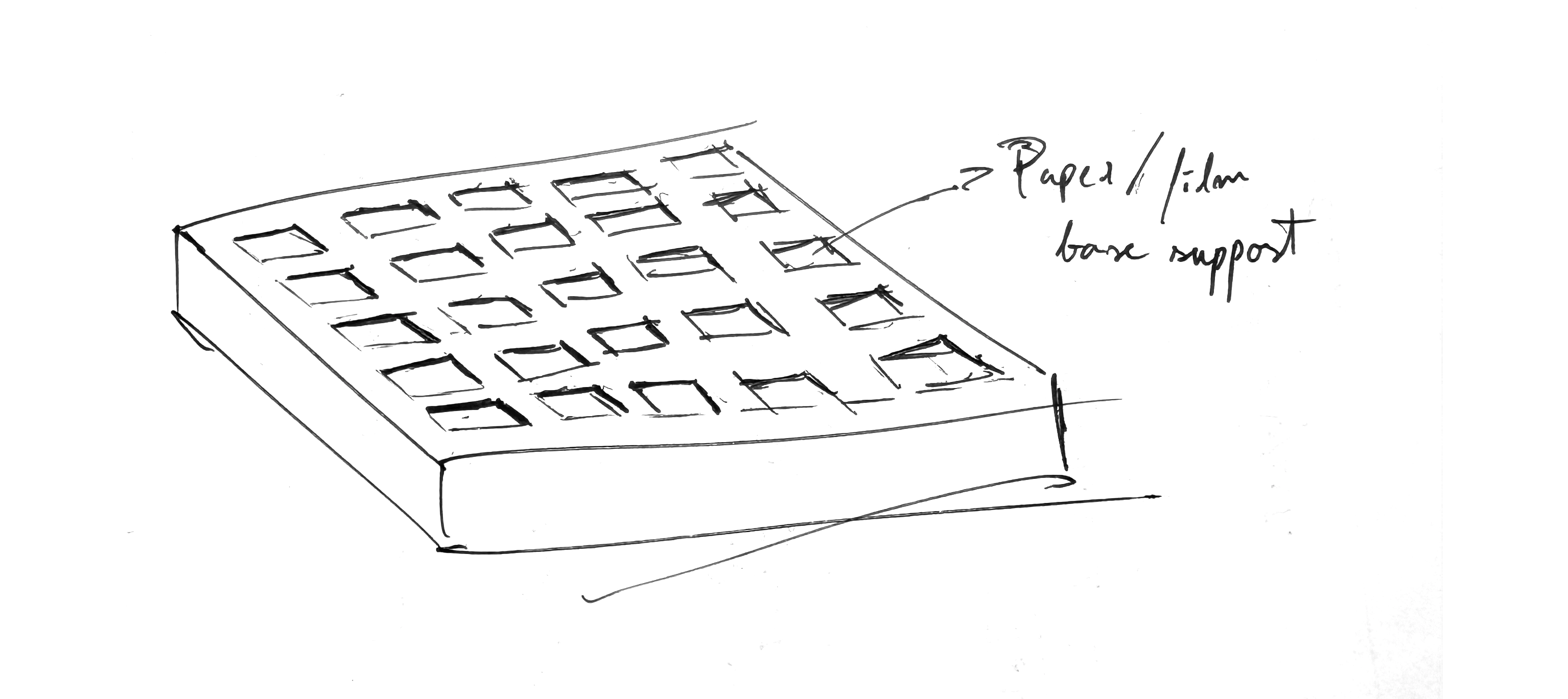
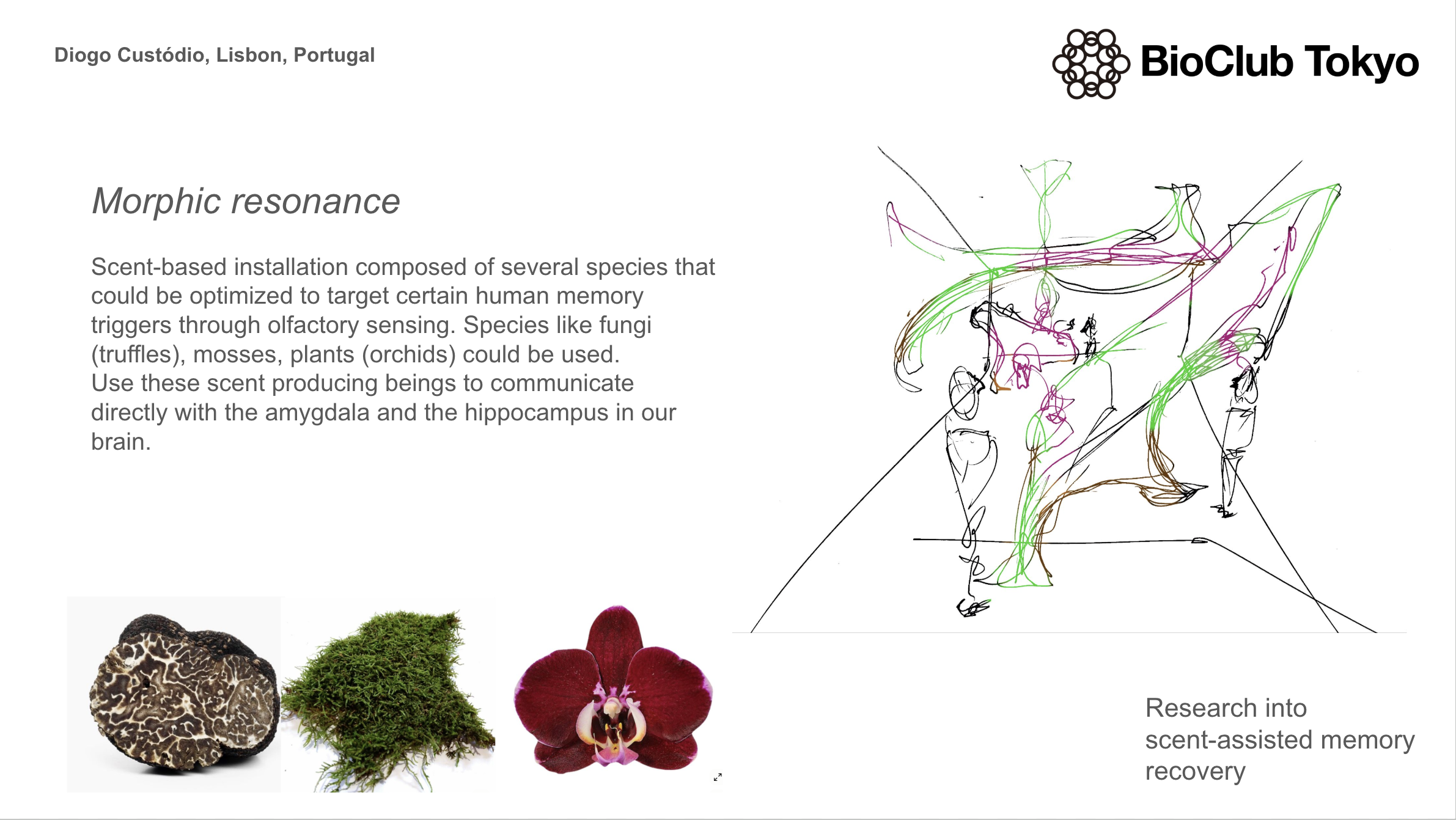
Final Project Ideas
References
APEX: Automated Protein EXpression in Escherichia coli
Automation and Optimization of Protein Expression and Purification on a Novel Robotic Platform
Bacterial Transformation Workflow
Folding in vitro of light-harvesting chlorophyll a/b protein is coupled with pigment binding
High-throughput, Microscale Magnetic Bead Protein Purification … Ginkgo … (RAC) Platform
Combining In vitro Folding with Cell-Free Protein Synthesis for Membrane Protein Expression