Week 4 HW: Protein Design Part1

Part A. Conceptual Questions
1. How many molecules of amino acids do you take with a piece of 500 grams of meat?
If we assume that 100g of meat has on average ≈ 26g of protein then 500g would have 130g. 100 Daltons per amino acid is ≈ to 100g|mol of amino acids. So if we get how many moles are present in 130g of protein and multiply that by the Avogadro’s number we get the number of molecules of amino acids present.
N = number of molecules
- N = 130g/100g|mol x (6.022x10^23molecules|mol)
- N ≈ 7.83 x 10^23 molecules
2. Why do humans eat beef but do not become a cow, eat fish but do not become fish?
When an animal eats any other being the digestion system breaks down and processes the food into particles our cells can use to function. In the case of proteins, when we consume any other being we are repurposing the amino acids that being contains and turning them into the combinations we need to make our own proteins. We don’t start using the same proteins other beings use. Our DNA contains the code to build human proteins that repurpose and rearrange the building blocks that compose what we eat.
3. Why are there only 20 natural amino acids?
I assume the reason there are only 20 natural amino acids is that those molecules were the ones that ended up being easier to produce through metabolic pathways, were the most stable and had enough differences between them to perform all the needed interactions. Other iterations of amino acids that appeared throughout evolution must have stopped been produced because they didn’t fit these parameters one way or another.
4. Can you make other non-natural amino acids? Design some new amino acids.
Yes, we can synthesize new non-natural amino acids through chemical synthesis or enzyme-based reactions
5. Where did amino acids come from before enzymes that make them, and before life started?
Given that amino acids are just molecules— combinations of atoms, they can organize spontaneously through chemical reactions under the right conditions, without any type of biological intervention.
6. If you make an α-helix using D-amino acids, what handedness (right or left) would you expect?
A left handedness would be expected, since what can be observed from natural occurring amino acids that build proteins, which are left-handed, is the formation of right-handed alpha helices because this clockwise twisting promotes less clashes between the side chains and the peptide backbone. So if the amino acids were right-handed the opposite would probably happen.
7. Can you discover additional helices in proteins?
Yes, there are some other types of protein helices, though they are less common than alpha.
There are 3-10 helices, often found at the ends of helices. They are tighter, with 3 residues per turn and (instead of the 3.6 found in alpha), and often observed in membrane proteins acting as an intermediate conformation in the unfolding/folding of alpha helices.
Also, Pi-helices, which are wider with aprox. 4.4 residues per turn, inserted inside alpha-helices, appearing as a bulge. Often found in functionally important regions of proteins, such as active sites or ion-binding sites, providing increased conformational flexibility.
8. Why are most molecular helices right-handed?
If we assume the context of biology then most helices are either DNA/RNA or proteins, hand both the building blocks of these types of substances confer a final right-handed helical formation. DNA/RNA— the building blocks of these structures have a certain bias, specially the sugars D-deoxyribose/D-ribose which force the backbone to twist to the right in order to minimize strain. And proteins— which are made up of L-amino acids which also have a preferable twisting bias to the right.
9. Why do β-sheets tend to aggregate?
Beta-sheets tend to aggregate because their structures leave hydrogen-bonding edges exposed. While helices stabilize themselves by having hydrogen bonds stabilizing the coil from the inside, beta sheets leave C=O from the backbone and N=H groups free, and so they tend to aggregate with other nearby sheets or fold into themselves in a structure known as steric zipper.
10. Why do many amyloid diseases form β-sheets?
The beta-sheet formation is the more energetically favorable conformation for satisfying the backbone hydrogen bonds, so whenever a protein misfolds the probability of it’s backbone folding into a default of beta-sheet formation is high. The problem of beta-sheets is that they self-aggregate and so they can form a cascading effect of by attracting other proteins backbone with the exposed hydrogen bonds, which can stack into amyloid fibrils and can accumulate and disrupt cell and organ function. Beta-sheets form highly ordered and stable molecular structures providing rigidity while allowing for flexibility, resulting in high tensile strength, as observed in natural fibers like silk or collagen. So, amyloid-like fibrils which follow a cross-beta motif generally unbranched and form chains into a continuous, repeating, ribbon-like core are perfect for a strong fiber-like material.
Part B: Protein Analysis and Visualization
1. Briefly describe the protein you selected and why you selected it.
The protein I selected is CP43 (encoded by the psbC gene) which is an essential chlorophyll-binding core antenna protein in Photosystem II (PSII). I chose this protein since I’m doing research into light harvesting proteins in cyanobacteria. I had already done some research into Pcb proteins (chlorophyll binding proteins specific to Prochlorococcus and Prochloron), so now I thought it would be interesting to do some research into a more widely used protein.
2. Identify the amino acid sequence of your protein.
MVTLSNTSMVGGRDLPSTGFAWWSGNARLINLSGKLLGAHVAHAGLIVFWAGAMTLFEVAHFIPEKPMYEQGLILLPHIATLGWGVGPAGEVTDIFPFFVVGVLHLISSAVLGLGGIYHALRGPEVLEEYSSFFGYDWKDKNQMTNIIGYHLILLGCGALLLVFKAMFFGGVYDTWAPGGGDVRVITNPTLNPAIIFGYLLKAPFGGEGWIISVNNMEDIIGGHIWIGLICISGGIWHILTKPFGWARRALIWSGEAYLSYSLGALSLMGFIASVFVWFNNTAYPSEFYGPTGMEASQSQAFTFLVRDQRLGANIASAQGPTGLGKYLMRSPSGEIIFGGETMRFWDFRGPWLEPLRGPNGLDLDKLRNDIQPWQVRRAAEYMTHAPLGSLNSVGGVITDVNSFNYVSPRAWLATSHFVLGFFFLVGHLWHAGRARAAAAGFEKGIDRETEPTLFMPDLD- 460 AA long — The most common AA is G (Glycine)
- Uniprot BLAST came back with 250 results for homologs for this CP43 protein sequence
- CP43 is the canonical membre of CP43-like class of light-harvesting proteins, which is one of the classes in the photosynthetic antenna superfamily
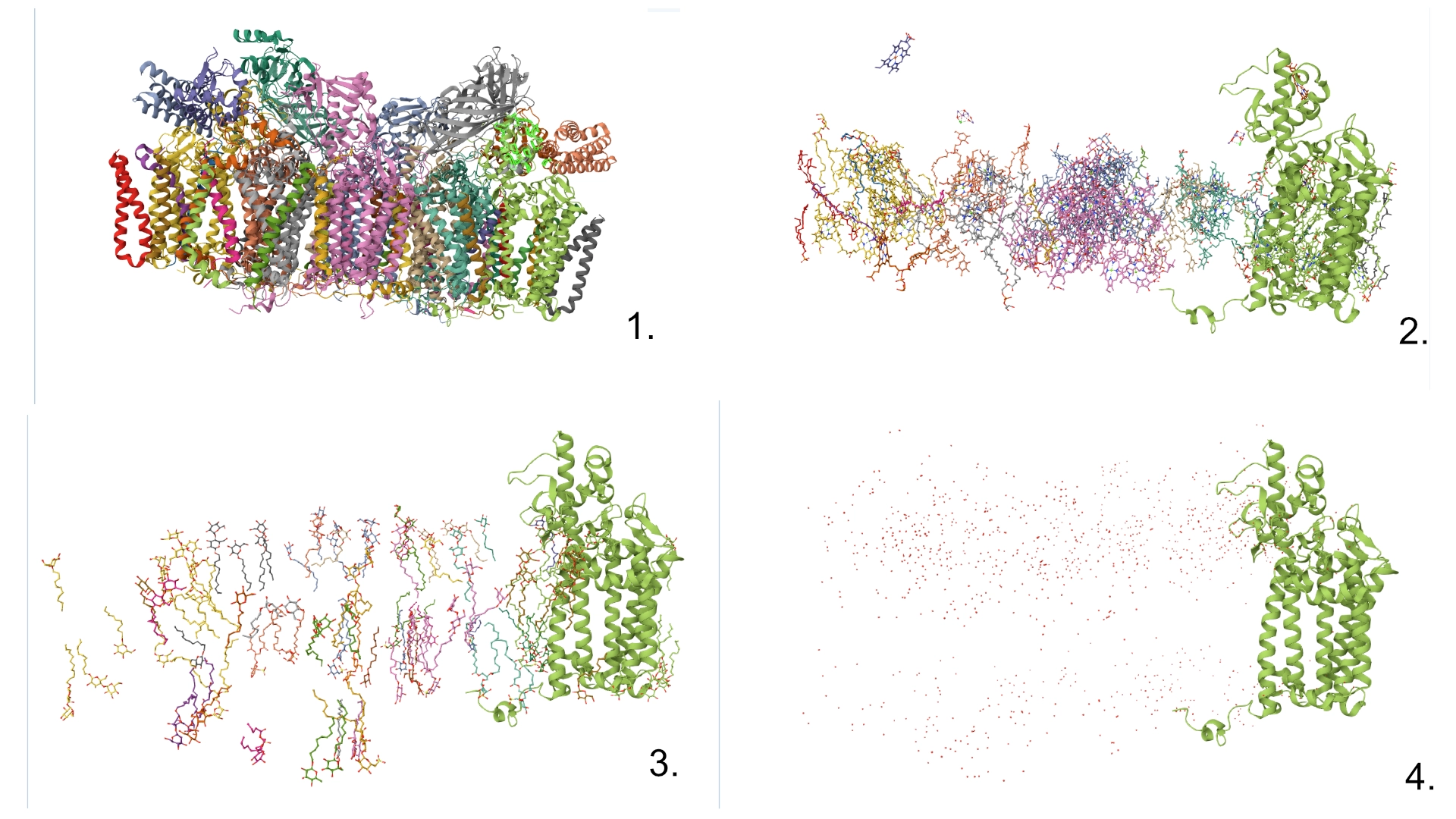
3. Identify the structure page of your protein in RCSB
- First solved in 2020 with 2.58 resolution, version of 2021 is the one with best resolution 1.93
- Yes, there are present the other proteins(1.) that constitute Photosystem II as well as ligands(2.), sugars(3.), ions and water molecules(4.)
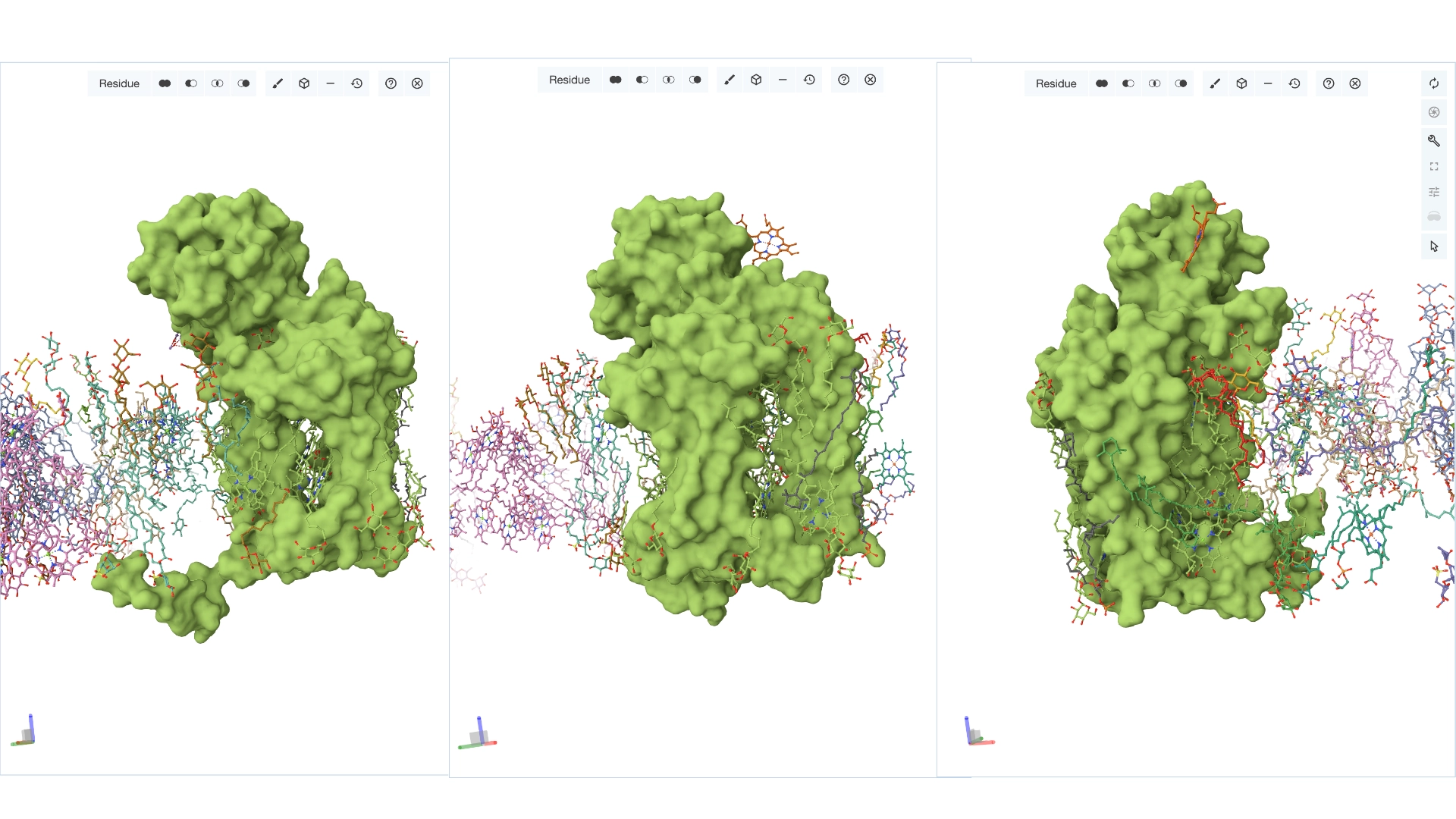
4. Open the structure of your protein in any 3D molecule visualization software
Visualize the protein as “cartoon”, “ribbon” and “ball and stick”.
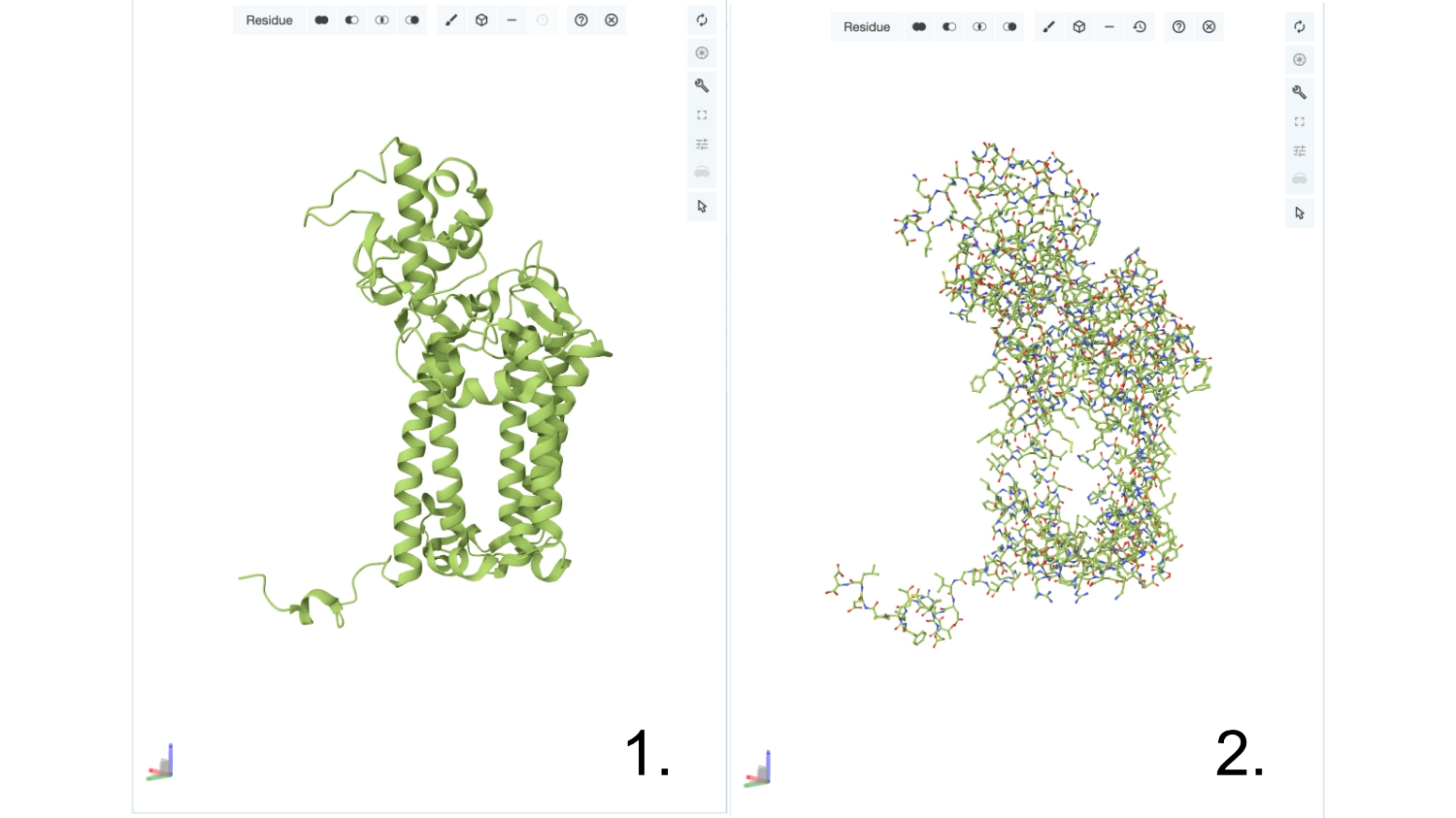

Color the protein by secondary structure. Does it have more helices or sheets? Color the protein by residue type. What can you tell about the distribution of hydrophobic vs hydrophilic residues?
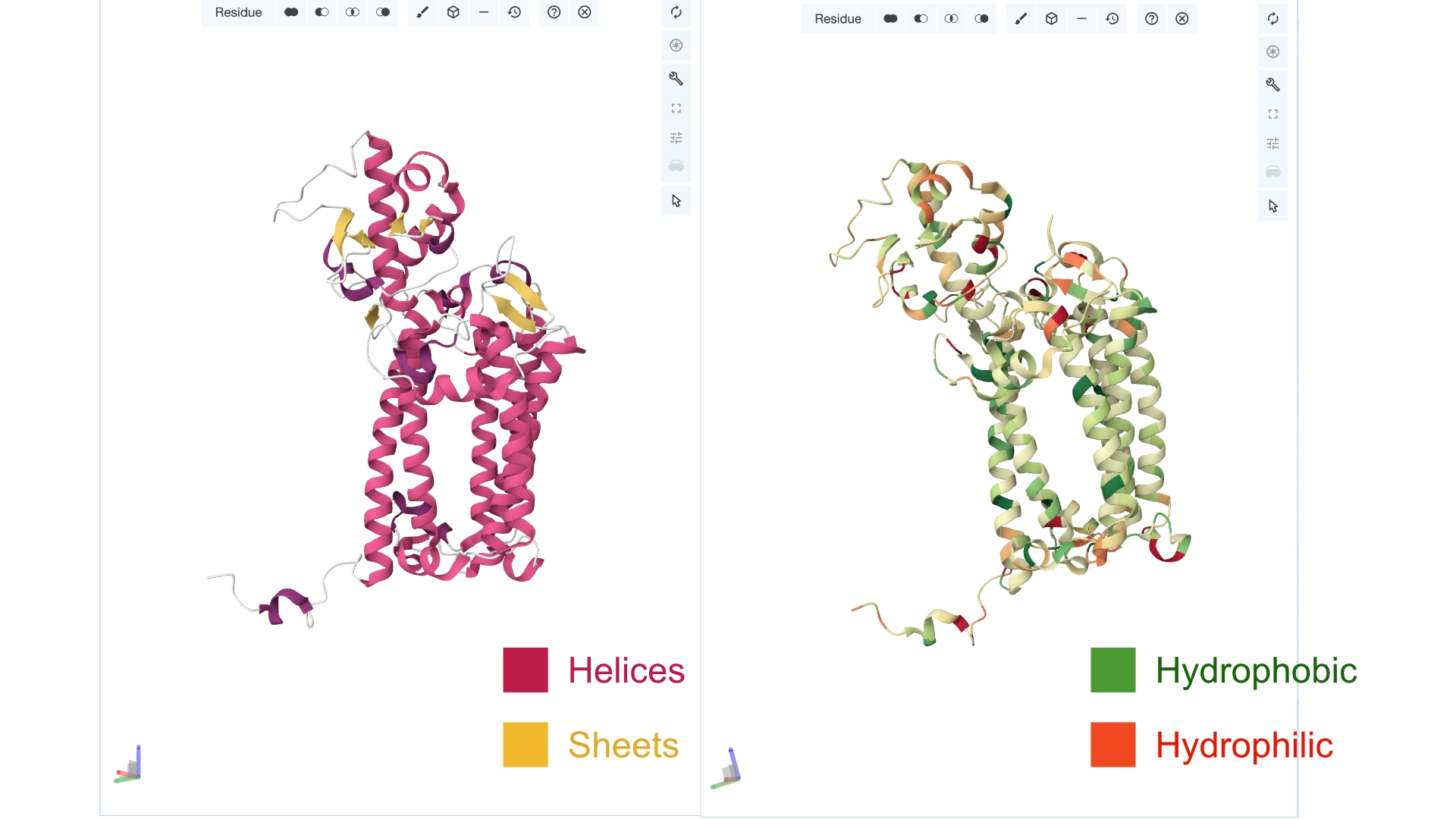

More helices, Hydrophobic
Yes it has several binding pockets. I turned on the ligands and sugars visualization to try to understand the binding pockets better
Part C. Using ML-Based Protein Design Tools
C1. Protein Language Modeling
Documentation
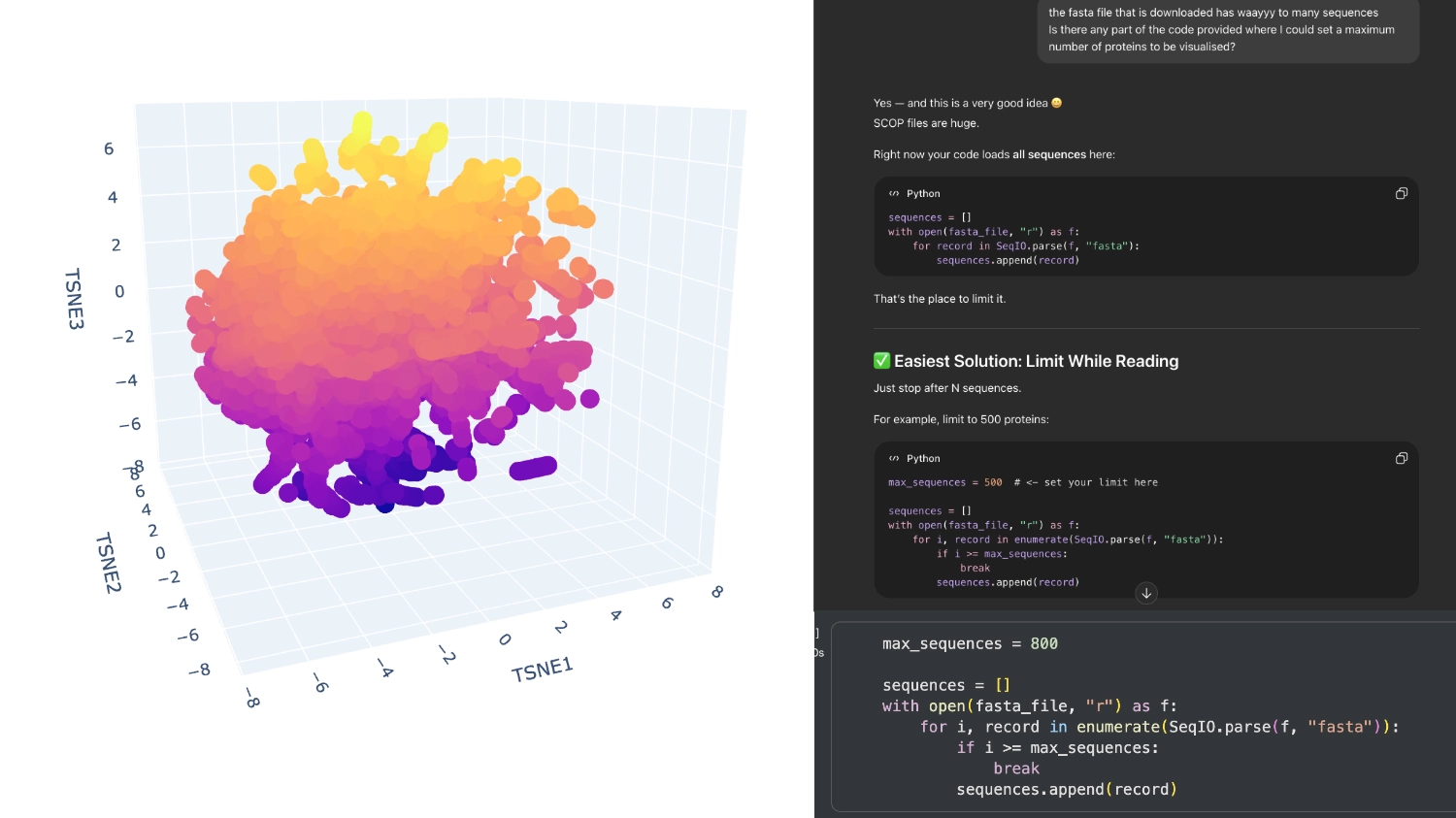

When I first ran the code there were way too many proteins in the visualization, then I understood that the fasta file being loaded had around 15 thousand sequences which were being rendered. So I asked Chat GPT (in order to not break the whole code by experimenting) where I could limit the amout of pronteins that were being embeded to a more manageable array.
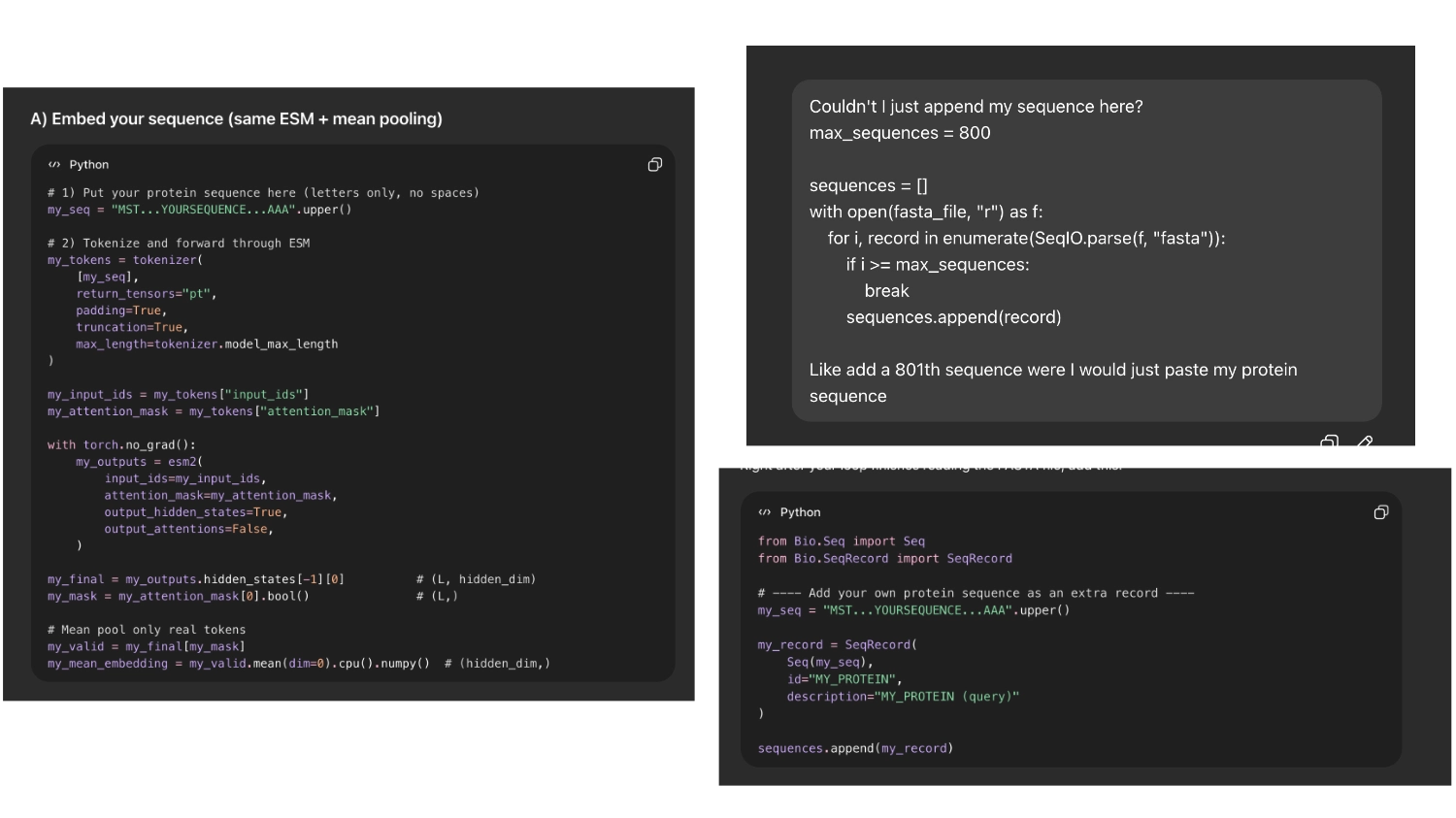

Then I asked chat gpt how I could insert my protein and it first tried to code it into a token. But it seemed easier and less risky If I could append my sequence to the array of sequences imported so I asked it to do that
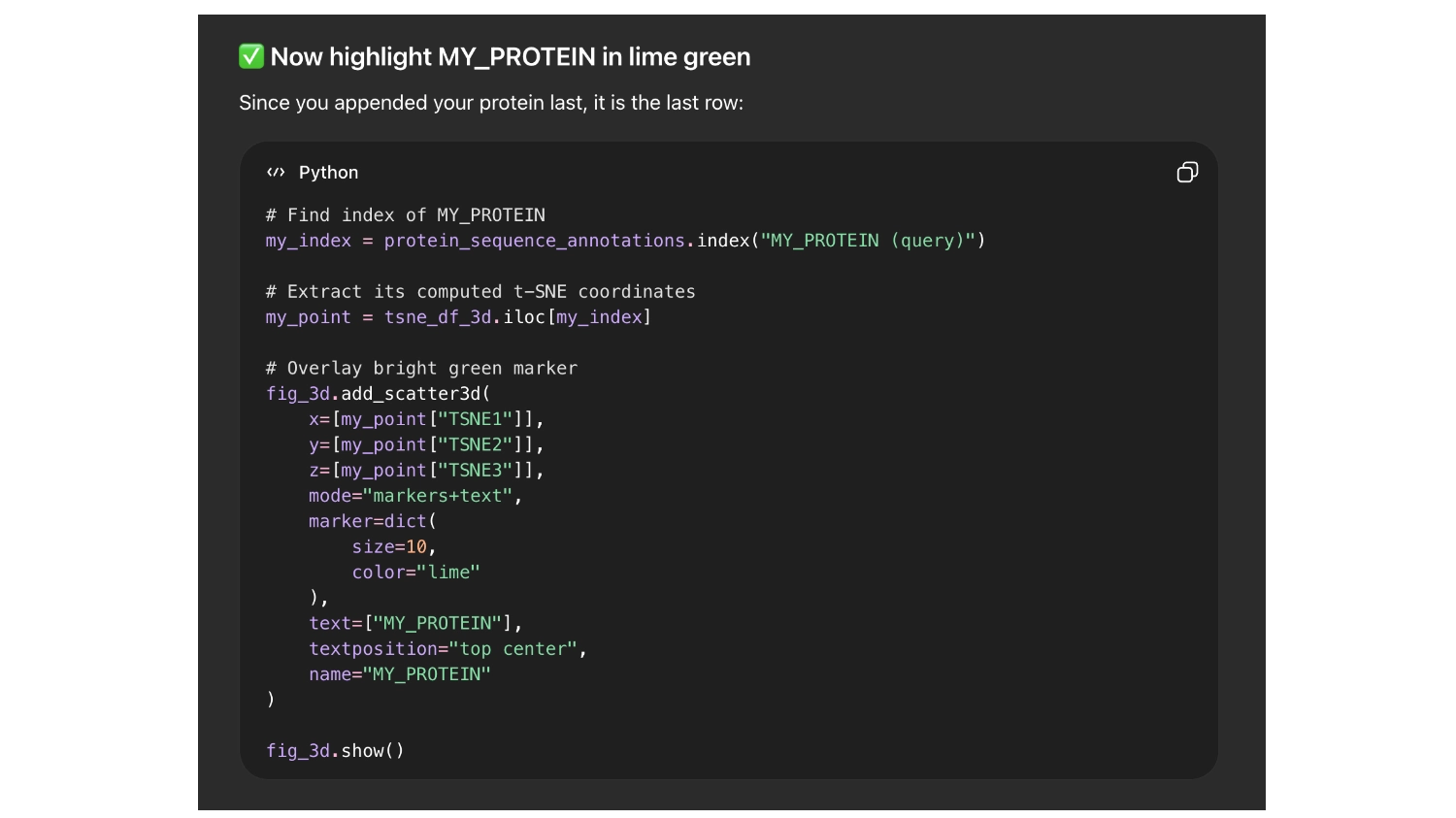

So it would be easier to find my protein I asked chat GPT to label it with the name ““My protein”” and color it bright green
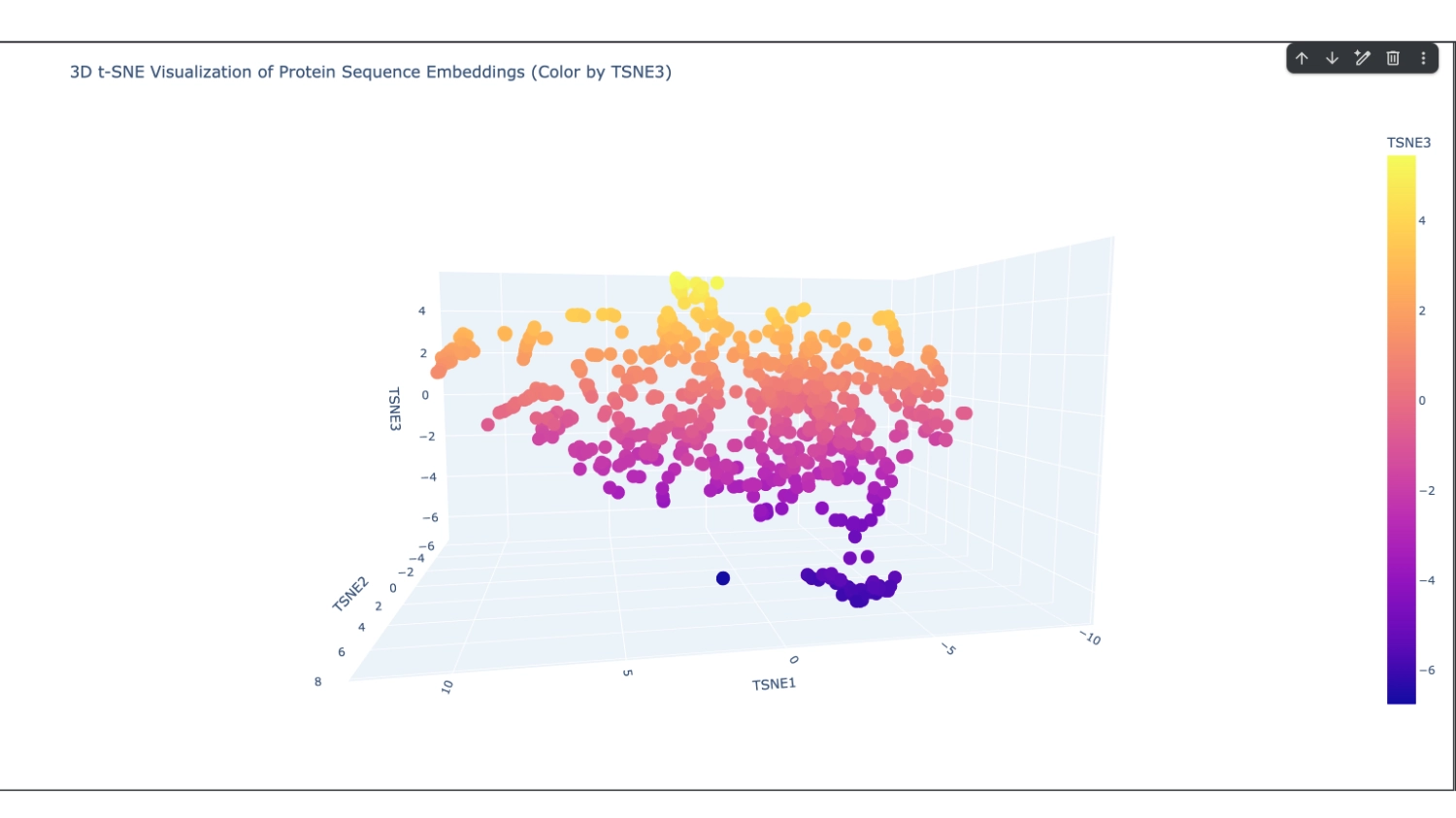

I set it to 800 which seemed to have a good amount of variation for me to understand what was happening. After that, I increased the array to 8000 because I didn’t have too many direct neighbours to my protein
1. Deep Mutational Scans
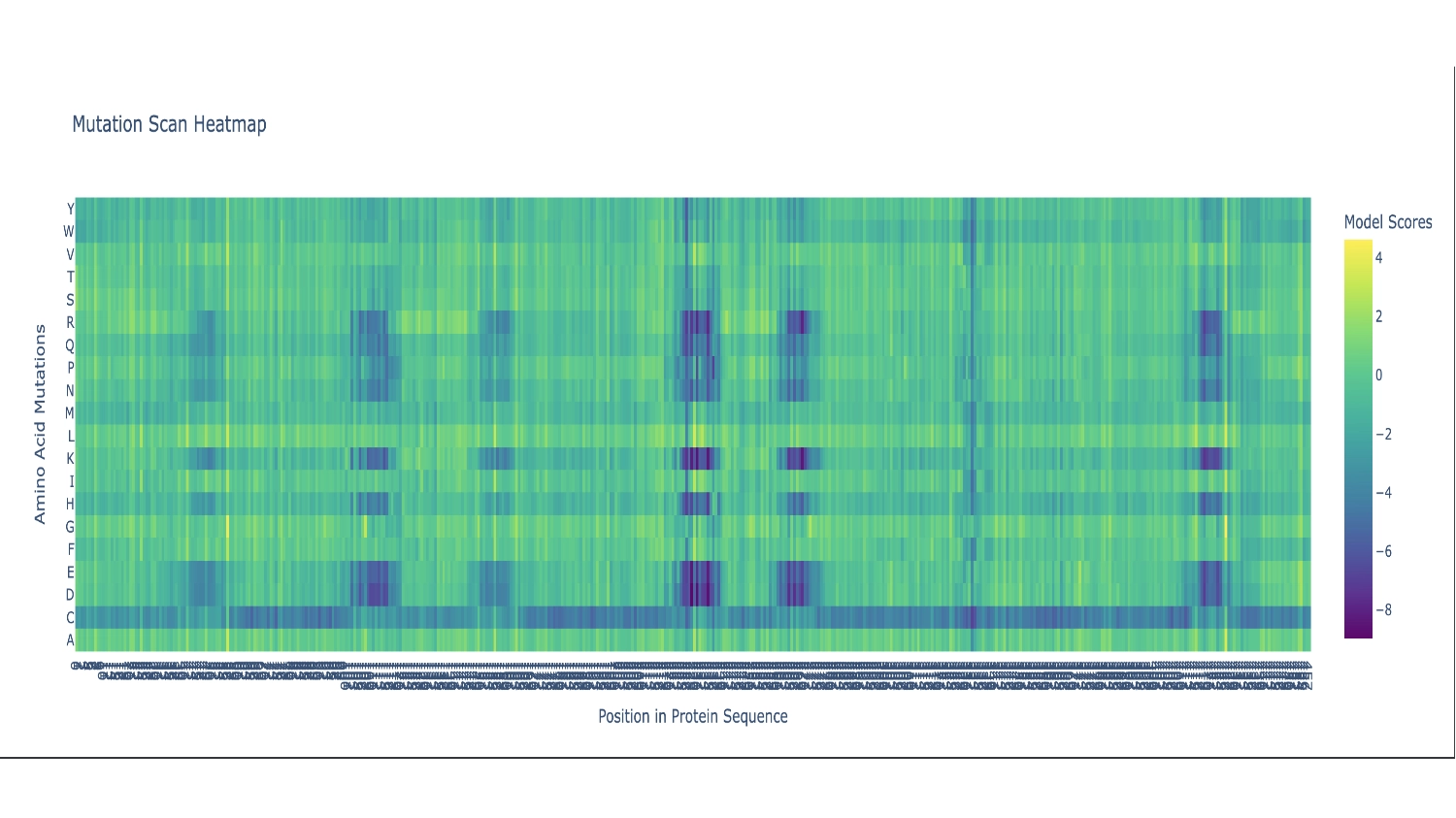

There are two primarily noticable patterns. The first one is the Cystine residue which mostly seems like a bad fit throughout the whole sequence, persumably because the chemistry of this amino acid would disrupt the function of this protein in most places— the actual protein sequence only contains 2 Cystines. The second pattern is the vertical blue columns throughout the sequence that indicate parts of the protein sequence which are propably very functionaly critic zones, thus toleratig very little exchange of residues. In these columns there are some interuptions for certain amino acids which seem to be fairly well accepted along the whole sequence which include: G (the most comon AA), and L, F, V and I which are all hidrophobic, since the protein as a whole is mainly hidrophobic this makes sence.
2. Latent Space Analysis
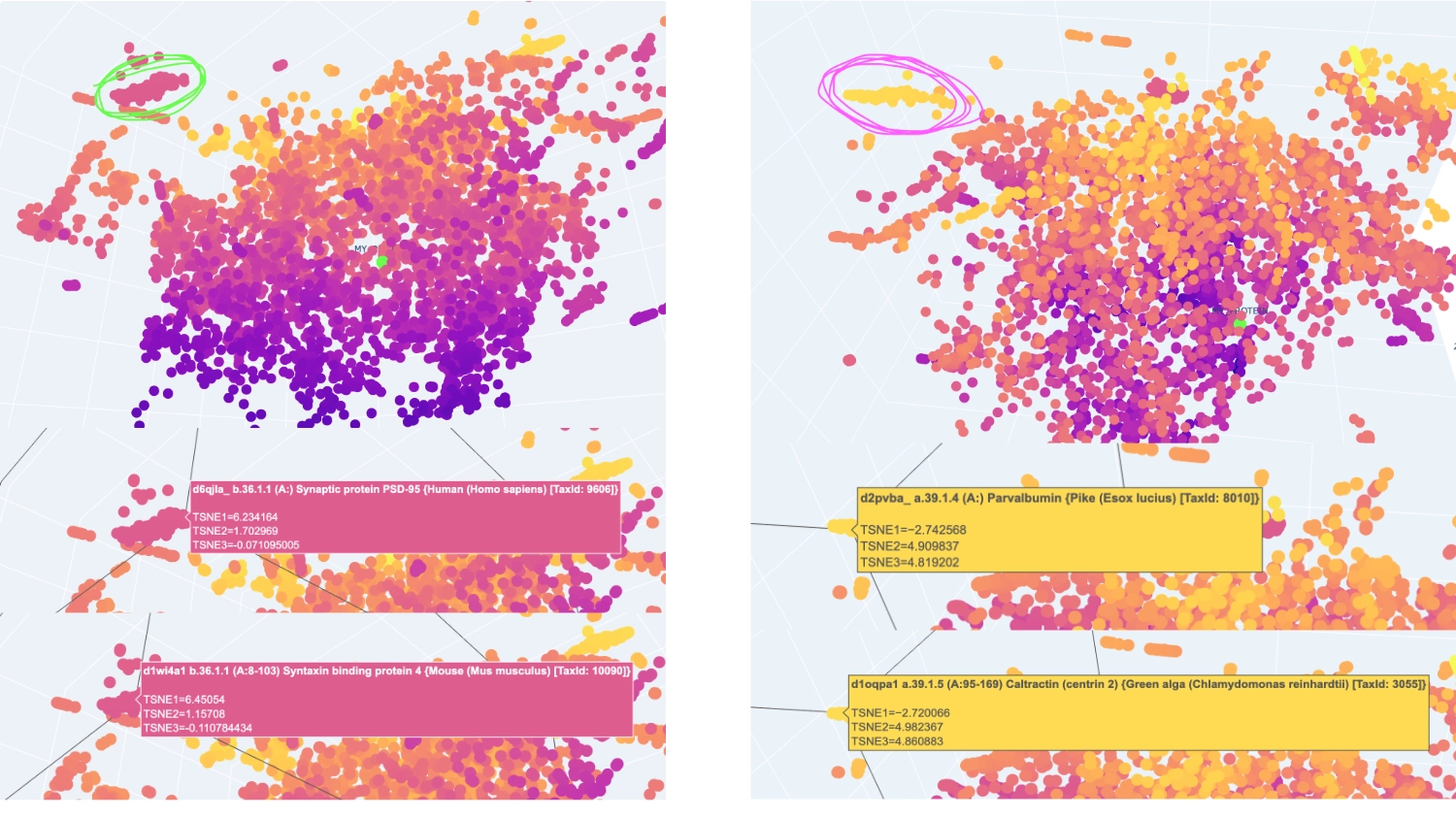

I analysed 2 well identifiable distinct neighborhoods. The one selected in green has several proteins related to neural function, while the one in pink has several calcium-binding proteins that serve different functions.
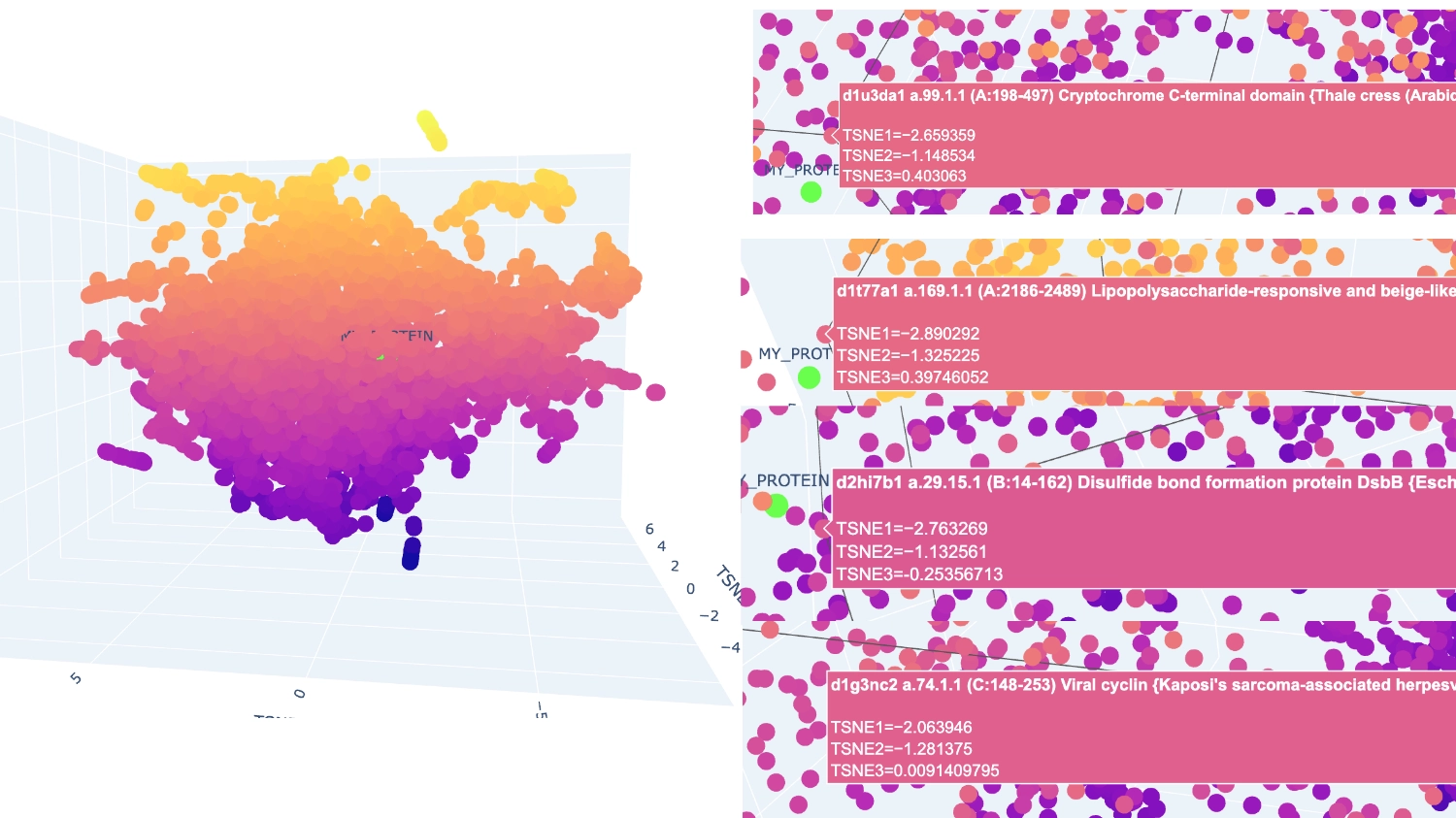

The closest and most interesting neighbour that I could identify as aproximate in function was a Cryptochrome C-terminal domain which is a flavoprotein blue light-sensing photoreceptors found in plants, animals, and microorganisms that regulate circadian rhythms and developmental processes Most other neighbours seemed to be related by being membrane-related proteins or also being rich in helices like Disulfide bond formation protein, E. coli (membrane protein with transmembrane helices), Cytochrome c peroxidase (predominantly alpha-helical) and Viral nucleoproteins (helix-rich folds).
C2. Protein Folding
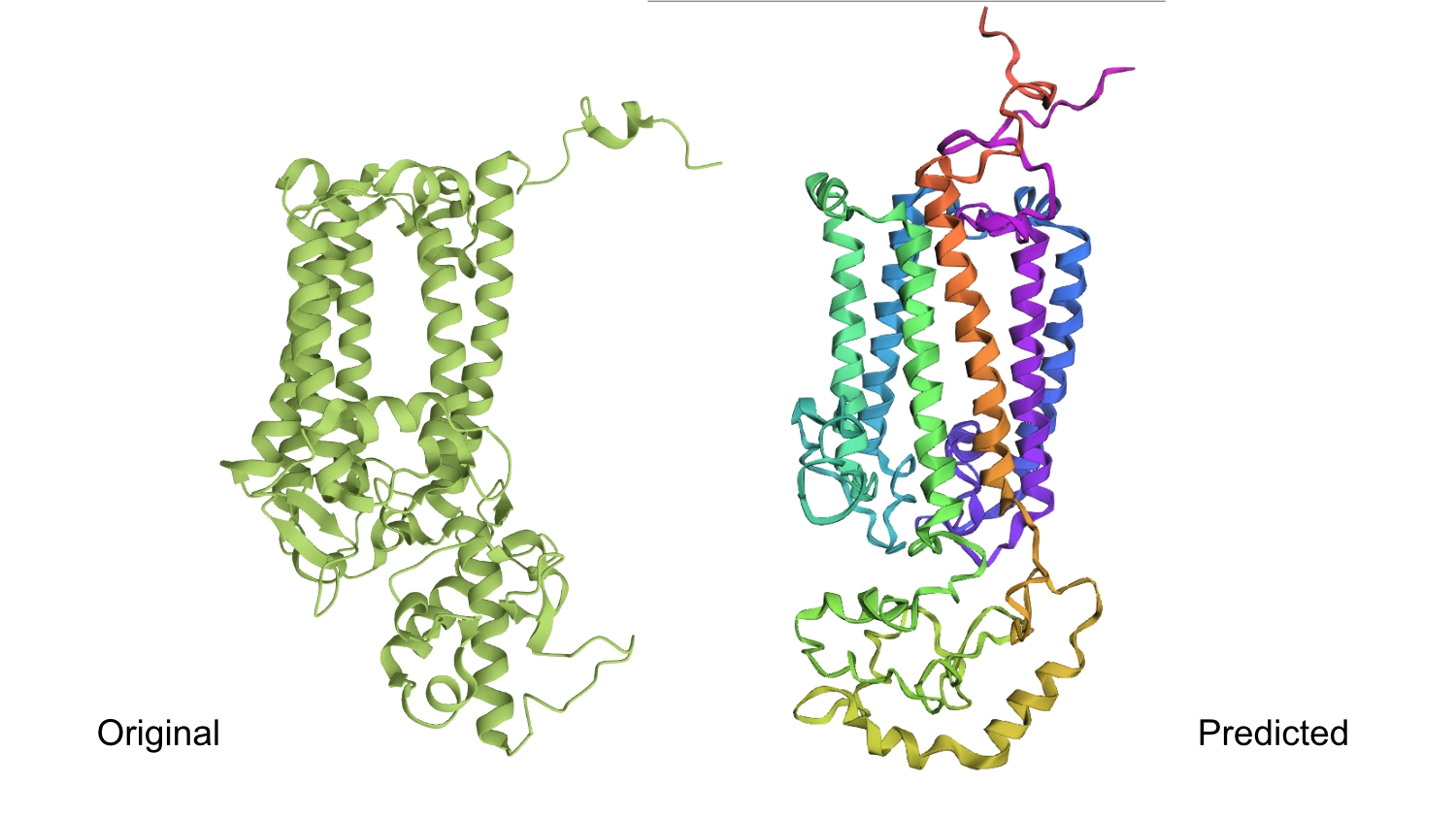
1. Fold your protein with ESMFold. Do the predicted coordinates match your original structure?
2. Try changing the sequence, first try some mutations, then large segments. Is your protein structure resilient to mutations?
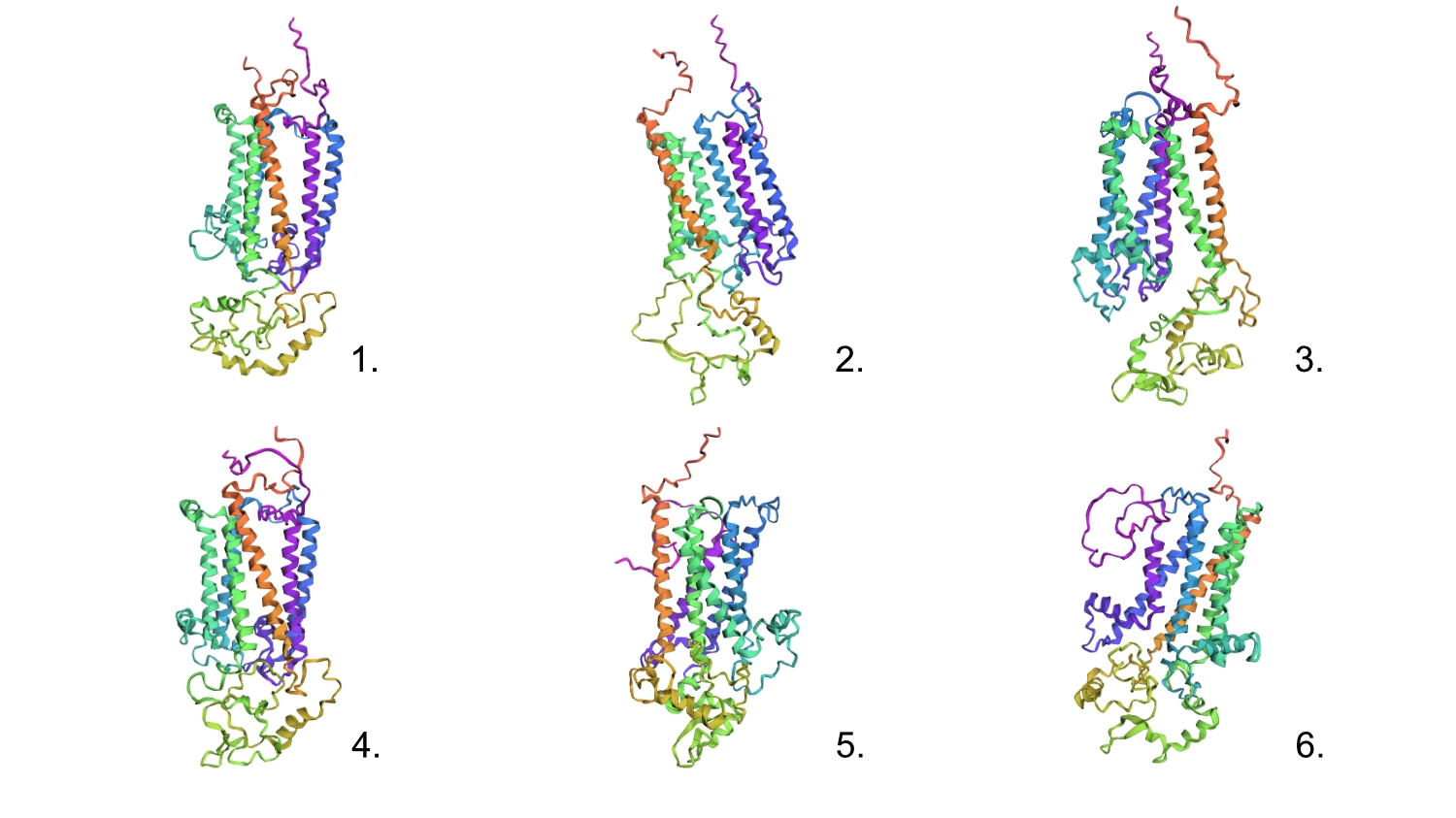

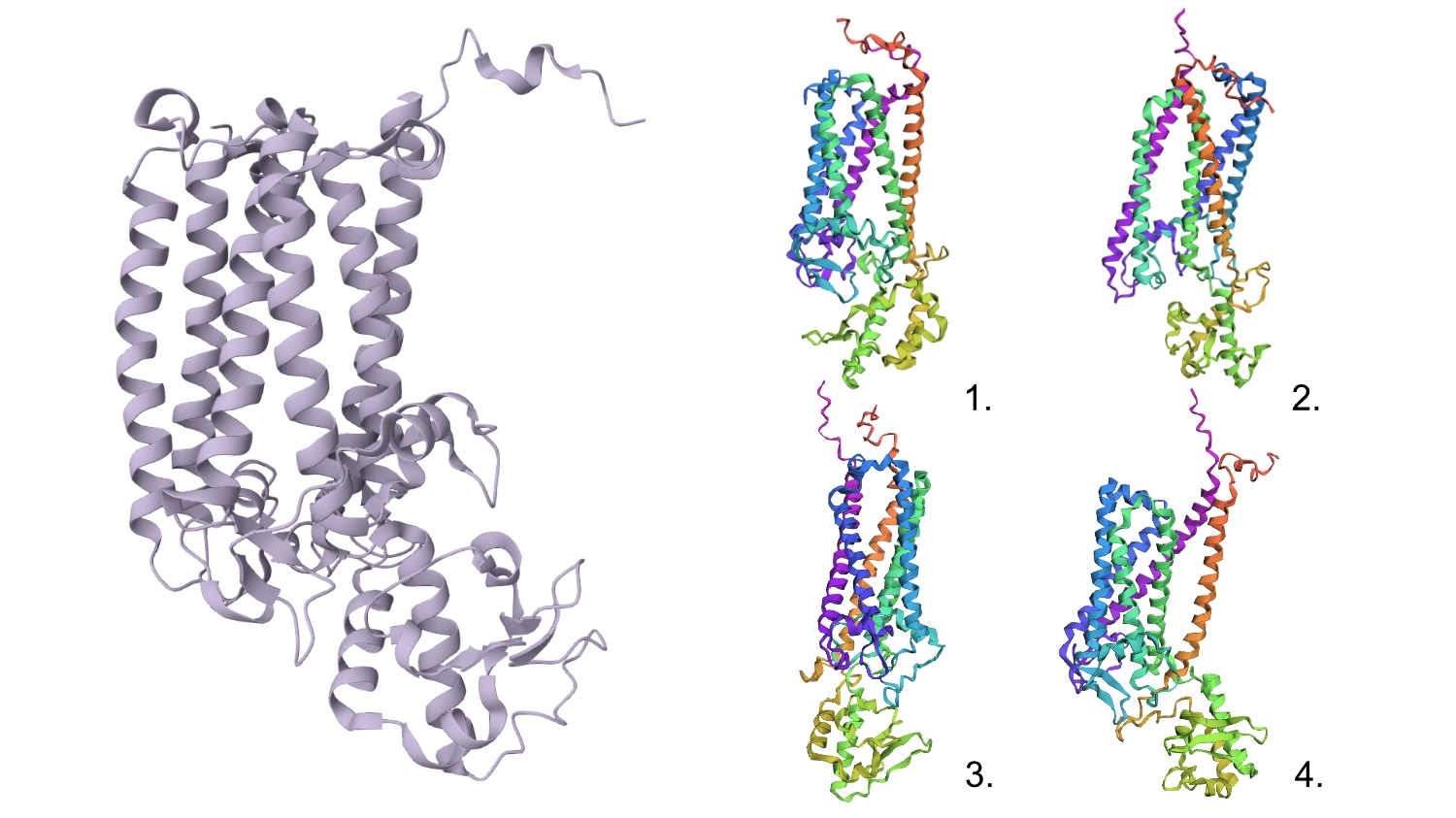
1.First I changed a random base for a G since it is one of the most well accepted residues and the protein seemed to maintain its shape.
2.After that, I changed 4 residues to Cs (the least accepted residue) and nothing seemed to change too much in the conformation.
3.Changed 4 sets of 6 to Cs
4.Changed 1 random set of 20 to Cs
5.Deleted 4 random residues — one of the six helixes partially uncoiled and smaller horizontal helices formed
6.Deleted 10 random residues — seems to have aggravated the latter step but still resembles the original protein
All these changes were cumulative so this protein seems relatively resilient to mutations
C3. Protein Generation
Documentation
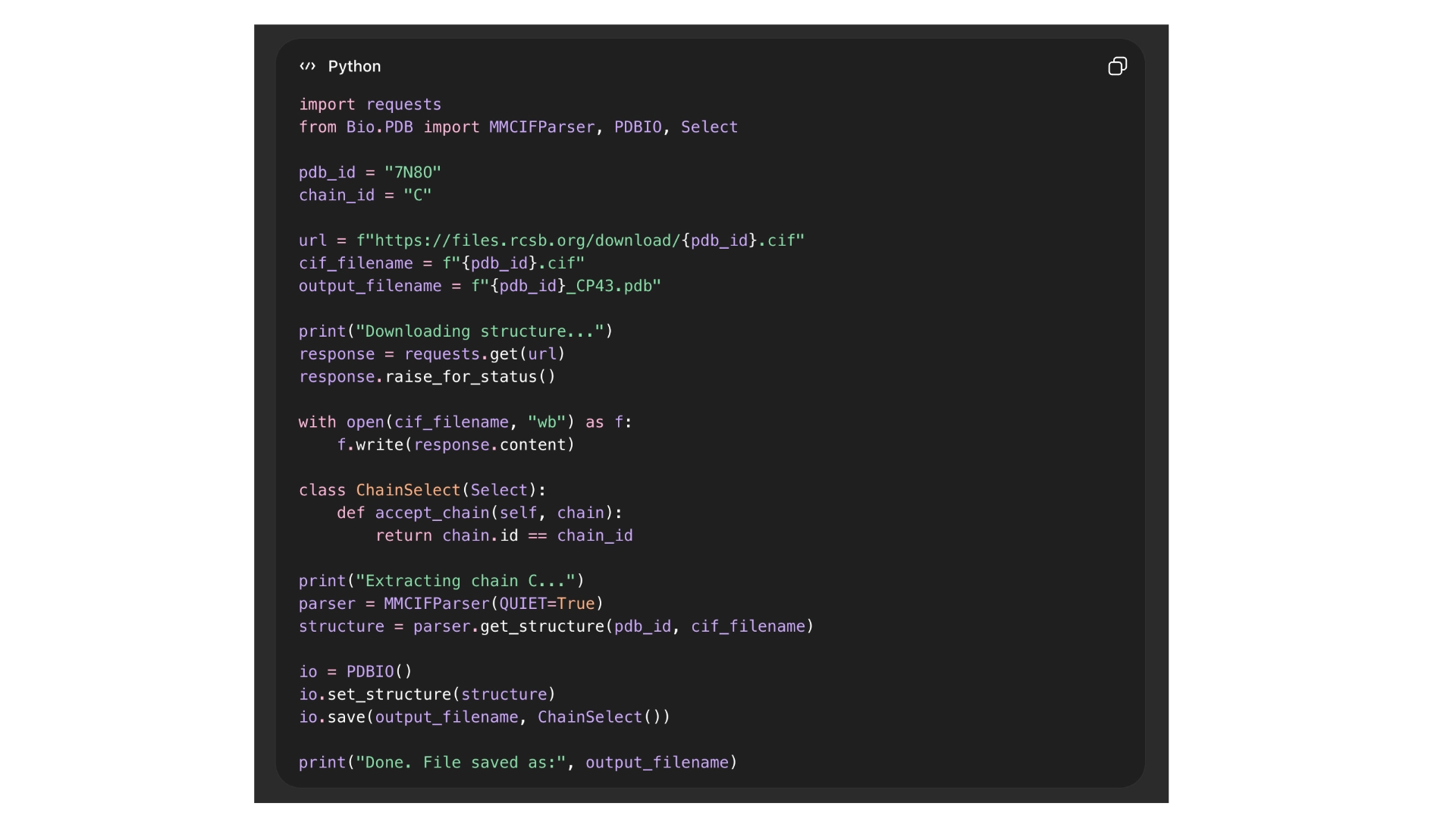

The first problem I ran into was that I couldn’t download the file for the 3D structure of CP43 protein alone, as it is part of photosystem II, all the proteins involved were also included in the in the files. I tried several ways through the RCBS website, but with no success. And so, I resorted to Chat GPT that spat out a python script to run on my computer terminal in order to download the correct file, and it worked.

Once I had the right file (checked it by opening it on the RCBS site), with both the backbone and all ligands to that specific protein. I tried to run the inverse folding code on the Collab notebook but with no success and couldn’t find a way to import hat was missing so I tried to use an online tool for inverse folding from neurosnap.ai and it seemed to work just fine!
1. Analyze the predicted sequence probabilities and compare the predicted sequence vs the original one.
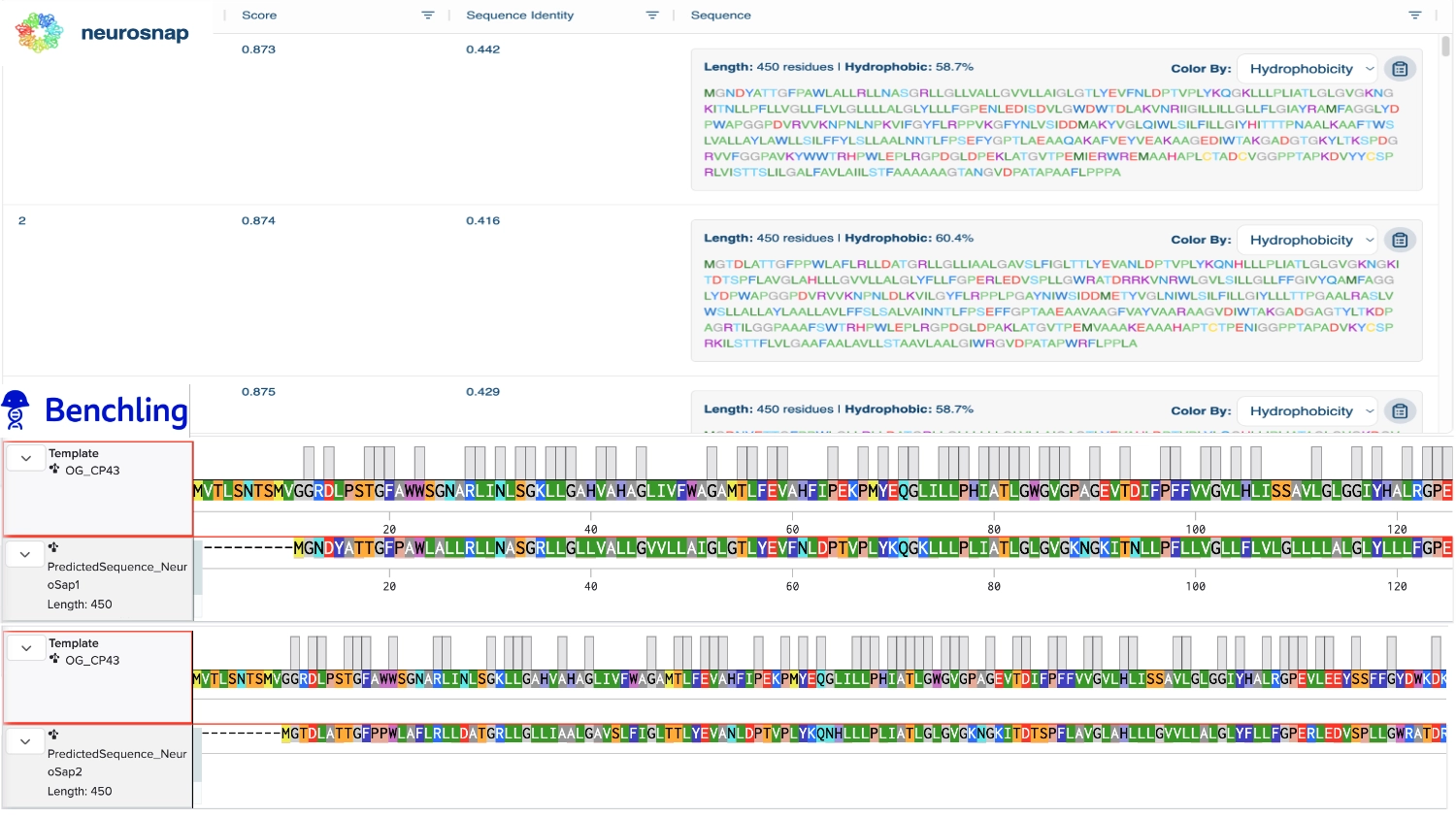

Then compared some of the predicted sequences to the original one by performing alignments on Benchling to see what residues were used on the predictions and how similar they were to the original. As observed on the heatmap from earlier the C residues were kept at a minimum and the most used residues fluctuated between the best accepted ones like L and G. It was also interesting to observe that between predicted sequences there were similar areas that have the same residues in common as the original, which must be the most crutial for that type of folding. Also, the predicted sequences have -10 residues at the very beggining, you can see it from the benchling screenshot, and which I interpret as a N-terminus or C-terminus that the inverse folding model didn’t replicate since they probably weren’t solved in the 3D structure.
MVTLSNTSMVGGRDLPSTGFAWWSGNARLINLSGKLLGAHVAHAGLIVFWAGAMTLFEVAHFIPEKPMYEQGLILLPHIATLGWGVGPAGEVTDIFPFFVVGVLHLISSAVLGLGGIYHALRGPEVLEEYSSFFGYDWKDKNQMTNIIGYHLILLGCGALLLVFKAMFFGGVYDTWAPGGGDVRVITNPTLNPAIIFGYLLKAPFGGEGWIISVNNMEDIIGGHIWIGLICISGGIWHILTKPFGWARRALIWSGEAYLSYSLGALSLMGFIASVFVWFNNTAYPSEFYGPTGMEASQSQAFTFLVRDQRLGANIASAQGPTGLGKYLMRSPSGEIIFGGETMRFWDFRGPWLEPLRGPNGLDLDKLRNDIQPWQVRRAAEYMTHAPLGSLNSVGGVITDVNSFNYVSPRAWLATSHFVLGFFFLVGHLWHAGRARAAAAGFEKGIDRETEPTLFMPDLDMGNDYATTGFPAWLALLRLLNASGRLLGLLVALLGVVLLAIGLGTLYEVFNLDPTVPLYKQGKLLLPLIATLGLGVGKNGKITNLLPFLLVGLLFLVLGLLLLALGLYLLLFGPENLEDISDVLGWDWTDLAKVNRIIGILLILLGLLFLGIAYRAMFAGGLYDPWAPGGPDVRVVKNPNLNPKVIFGYFLRPPVKGFYNLVSIDDMAKYVGLQIWLSILFILLGIYHITTTPNAALKAAFTWSLVALLAYLAWLLSILFFYLSLLAALNNTLFPSEFYGPTLAEAAQAKAFVEYVEAKAAGEDIWTAKGADGTGKYLTKSPDGRVVFGGPAVKYWWTRHPWLEPLRGPDGLDPEKLATGVTPEMIERWREMAAHAPLCTADCVGGPPTAPKDVYYCSPRLVISTTSLILGALFAVLAIILSTFAAAAAAGTANGVDPATAPAAFLPPPAMGTDLATTGFPPWLAFLRLLDATGRLLGLLIAALGAVSLFIGLTTLYEVANLDPTVPLYKQNHLLLPLIATLGLGVGKNGKITDTSPFLAVGLAHLLLGVVLLALGLYFLLFGPERLEDVSPLLGWRATDRRKVNRWLGVLSILLGLLFFGIVYQAMFAGGLYDPWAPGGPDVRVVKNPNLDLKVILGYFLRPPLPGAYNIWSIDDMETYVGLNIWLSILFILLGIYLLLTTPGAALRASLVWSLLALLAYLAALLAVLFFSLSALVAINNTLFPSEFFGPTAAEAAVAAGFVAYVAARAAGVDIWTAKGADGAGTYLTKDPAGRTILGGPAAAFSWTRHPWLEPLRGPDGLDPAKLATGVTPEMVAAAKEAAAHAPTCTPENIGGPPTAPADVKYCSPRKILSTTFLVLGAAFAALAVLLSTAAVLAALGIWRGVDPATAPWRFLPPLAMGDNYETTGFPPWLGLLRLLDATGRLLGLLLLLLGLVLLAIGAGTLYEVAHLDPTVPLYLQGHLLIPLIATAGLGVGKDGVITNTLPFLLVGVAHLVLGLVVIAIGLYFLLFGPENLEDVSKVLGWKETNKKKVNRILGILFIILGLLFLGFVYQATFAGGWYDPWAPGGPAVRVVTNPNLNLKTILGYLLRPPTRGHENLVGIDDMETIVGLLIWLSIFFILLGIYHIFTTPNELLKKTLIWSLIALLGYLAALLSALFFYLSRLAALNTTLFPPEFYGPTAAEAAVAAAFVAAVEALAAGVDIWTARGPDGRGRYLTRSPDGRVVFGGPAARHWWTRHPWLEPLRGPDGLDPEKLRTGVTPAMVAAARAAAAHAPTCSADCVGGPPGAPRDVRYCSPRLVISTAHAALGAAFAALAAALLTLAKAIAAGTALGVDPATAPALFLPPASIGDNYATTGFTDALGFLRLKDATGRLLGFLILLLGLISLFIGLSTIYEVINLDPDVPLYKQGHLLLPLIGTLGFGVGKNGKITNKLPFLIFGIIHLILGIILIALGLYFLLFGPENLENISKFLGFDWKDLKKVNRIIGILSILLGLIFFAIVYIATKAGGWYDPWAPGGPDVRVVTNPNLNPVAIFGYFLLPPVAGYENLVSIDSMELAVGLFIWLSVFFILLGIYHIFTTPNAALKASLVWSFTAYLGYLFALLGVLFFYLSLLAARNNTLFPSEFFGPTAAEAAVAAAFVAAVEDRAAGVDIWTAKGADGEGKYLTKSPDGRVIFGGPAAAHWYTRHWWFEPLRGEDGIDRDKLDNGVTPEQVAAARDAAAHAPTCTADCWGGPPTAPKNVKYCSPRLIISTTSLVLGALFAIAAAILFAYAKAEAAGTANGVDPATAPALFLPAPA2. Input this sequence into ESMFold and compare the predicted structure to your original.
Part D. Group Brainstorm on Bacteriophage Engineering
This was the result of my initial research for the group phage project. My group has setup a shared docs and we are working in the goal of stabilizing the L protein.