Week 5 HW: Protein Design Part2

Part A: SOD1 Binder Peptide Design (From Pranam)
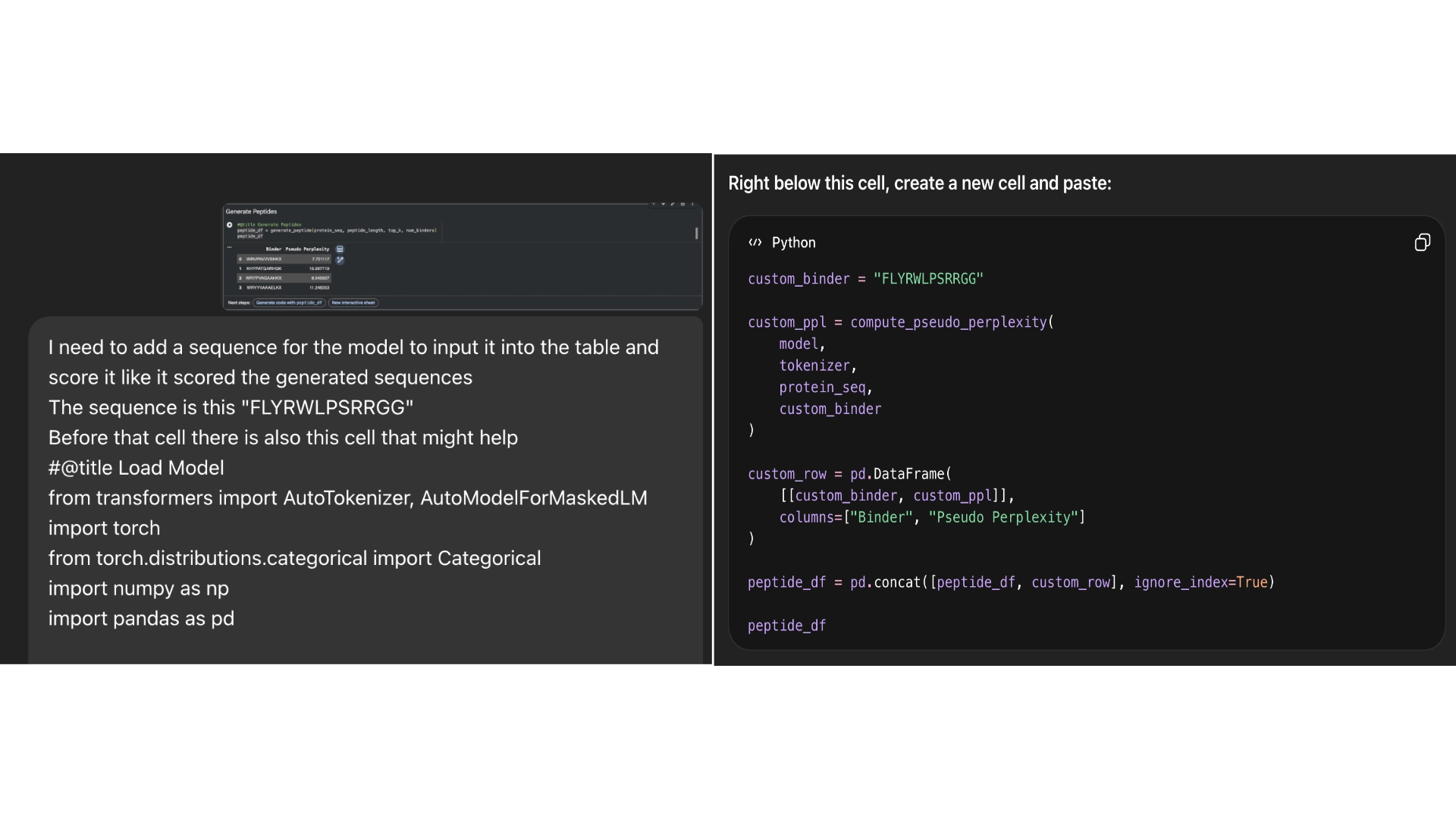
Part 1: Generate Binders with PepMLM
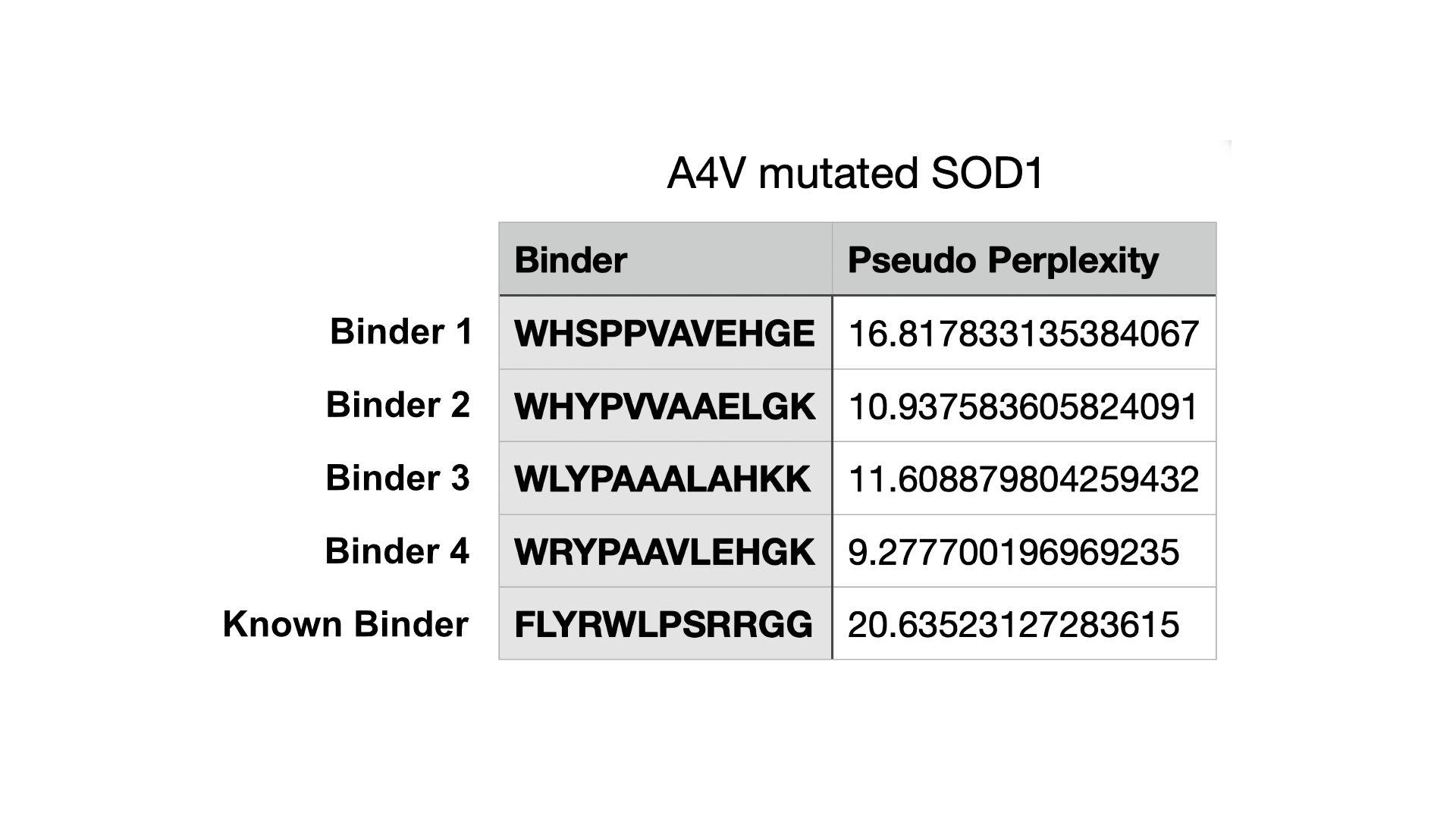

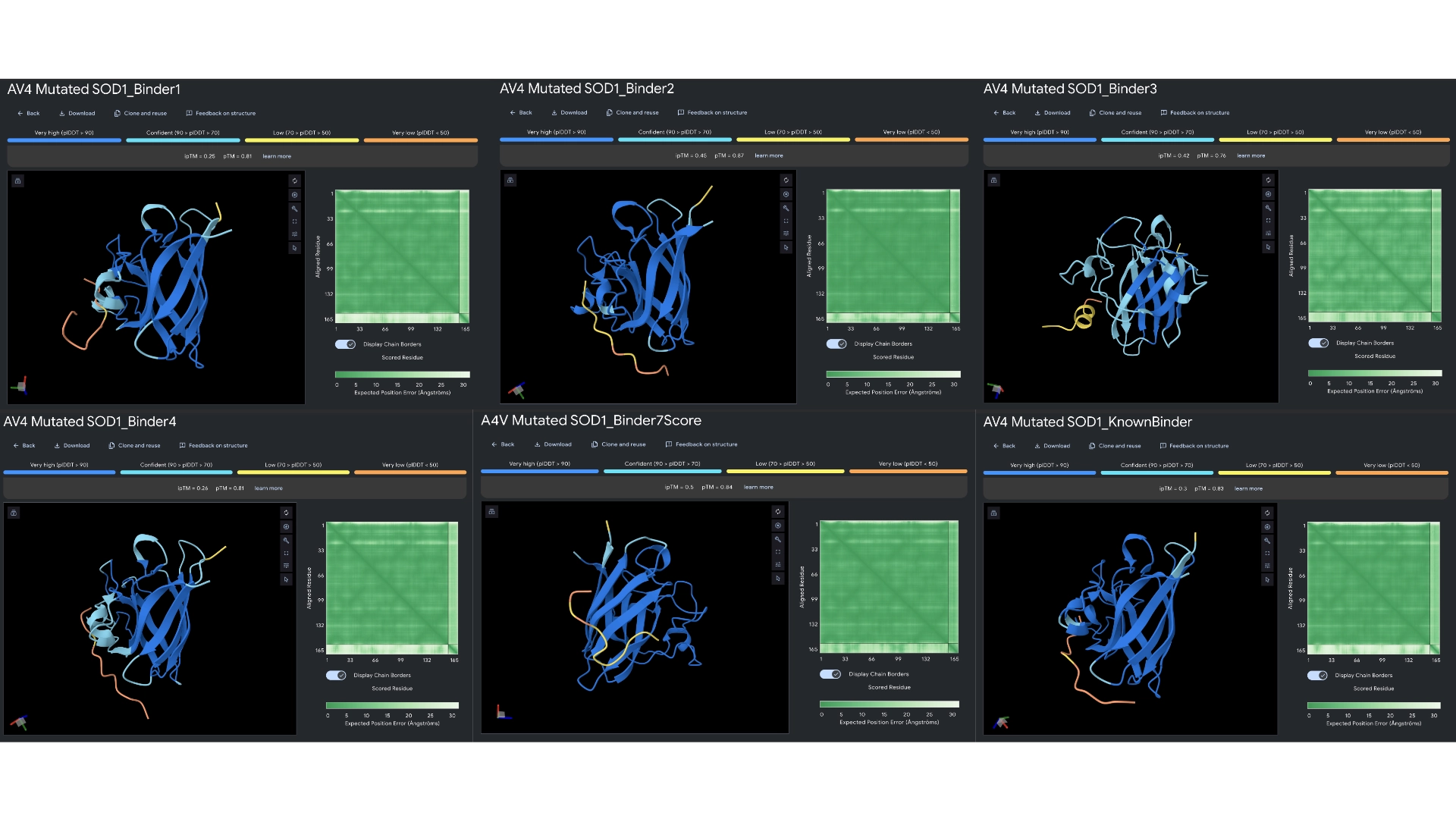
Part 2: Evaluate Binders with AlphaFold3
Part 3: Evaluate Properties of Generated Peptides in the PeptiVerse
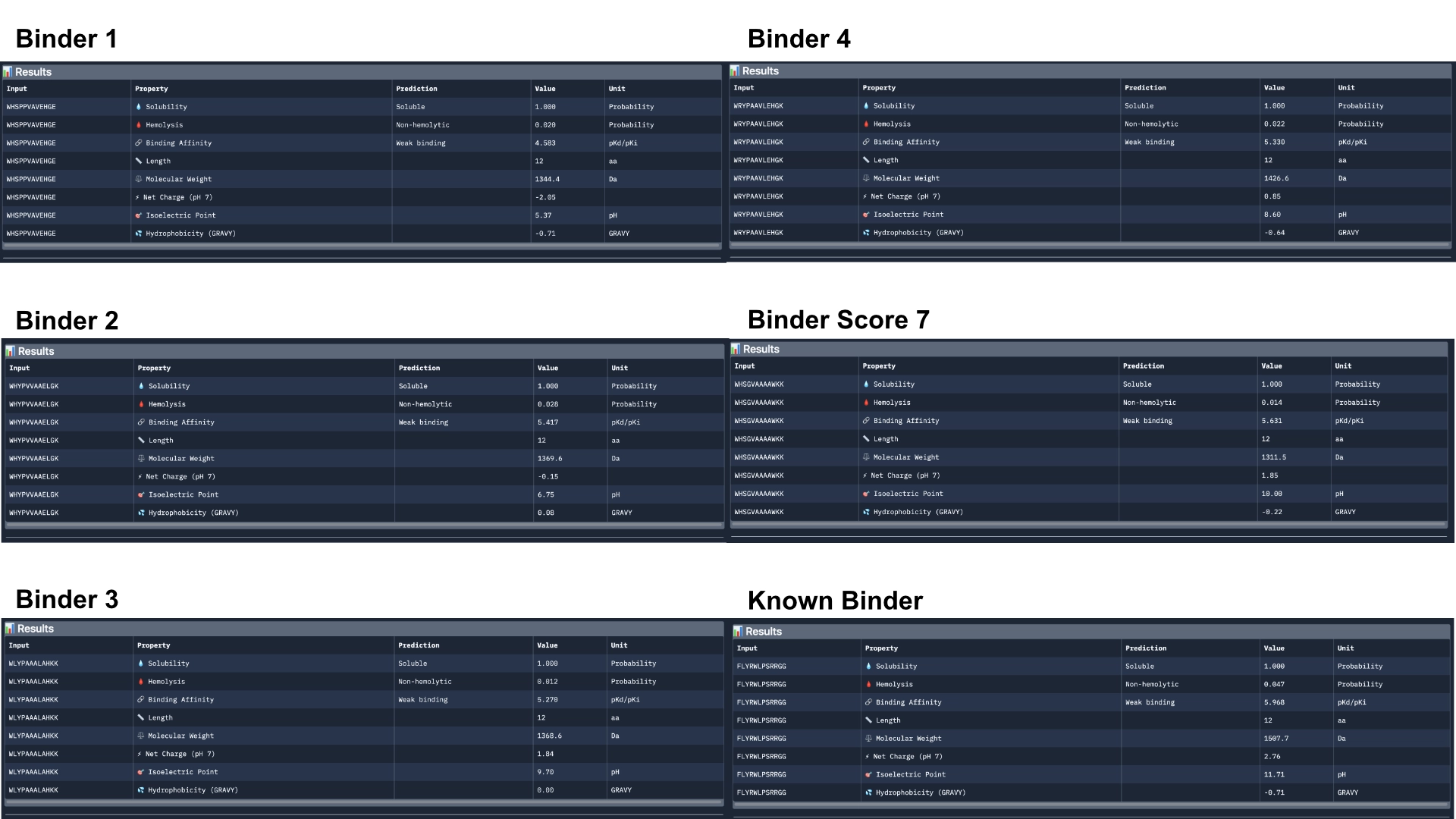

Overall, Peptiverse considers all generated binder and known binder as weak binders, the known binder having the highest score of 5.968, followed by the binder which had the highest ipTM score, with a binding affinity 5.631. However, binders 2 and 3, which surpassed the known binder in ipTM score, were not attributed better binding affinitty, having the scores 5.417 and 5.270 respectively. All predicted binders had good therapeutic properties scores and none were non-soluble nor hemolytic. The best overall in balancing all aspects was the known binder with 5.968 of predicted binding and 0.047 of hemolytic probability. The best predicted binder, althought, it has a slightly lower hemolytic probability of 0.014, that might not balance out the 0.337 difference in binding affinity (5.631). However, this would probably be the best to advance with, even though it might not exceed the qualities of the already known binder.
Part 4: Generate Optimized Peptides with moPPIt
First of all I generated 3 peptides for the A4V mutated SOD1 with the only parameters being affinity and motif whith the following weights:
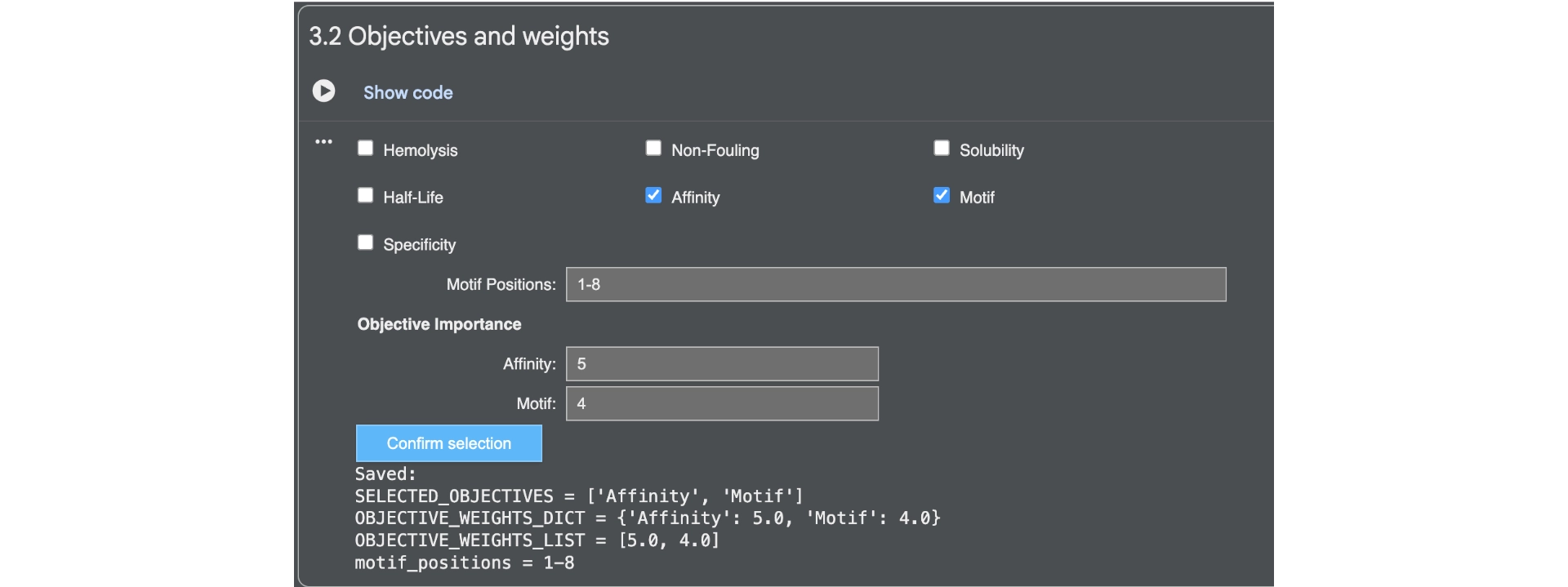

Then analysed the resulting peptides in PeptiVerse.
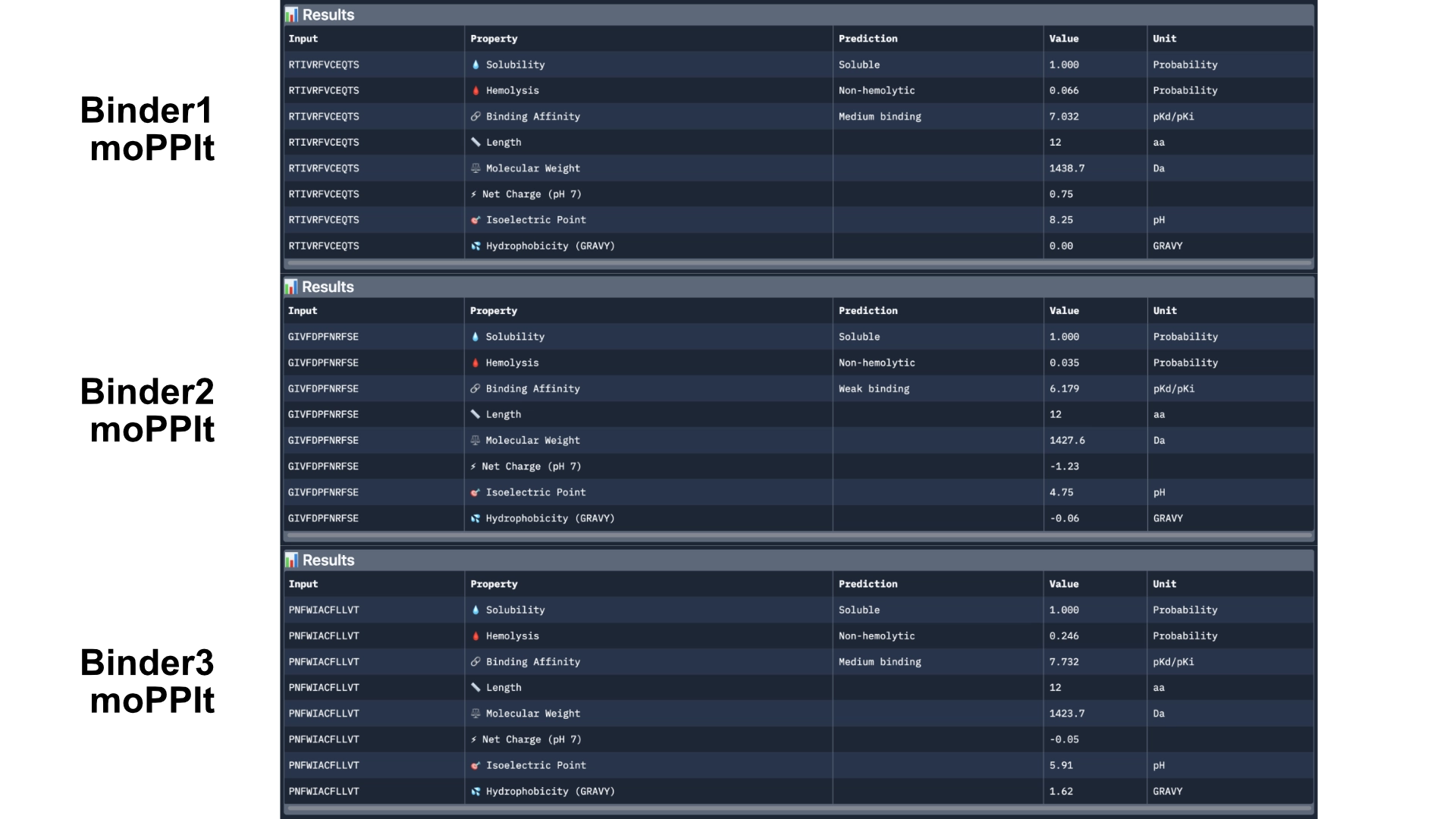

Comparing these to the PepMLM ones, there already was a good improvement as they were all soluble and non-hemolytic and had better affinities with two of them reaching medium binding instead of the previous weak binding levels.
Then generated other 3 peptides with non-hemolytic and solubility objectives to watch how they would differ from the previous ones.
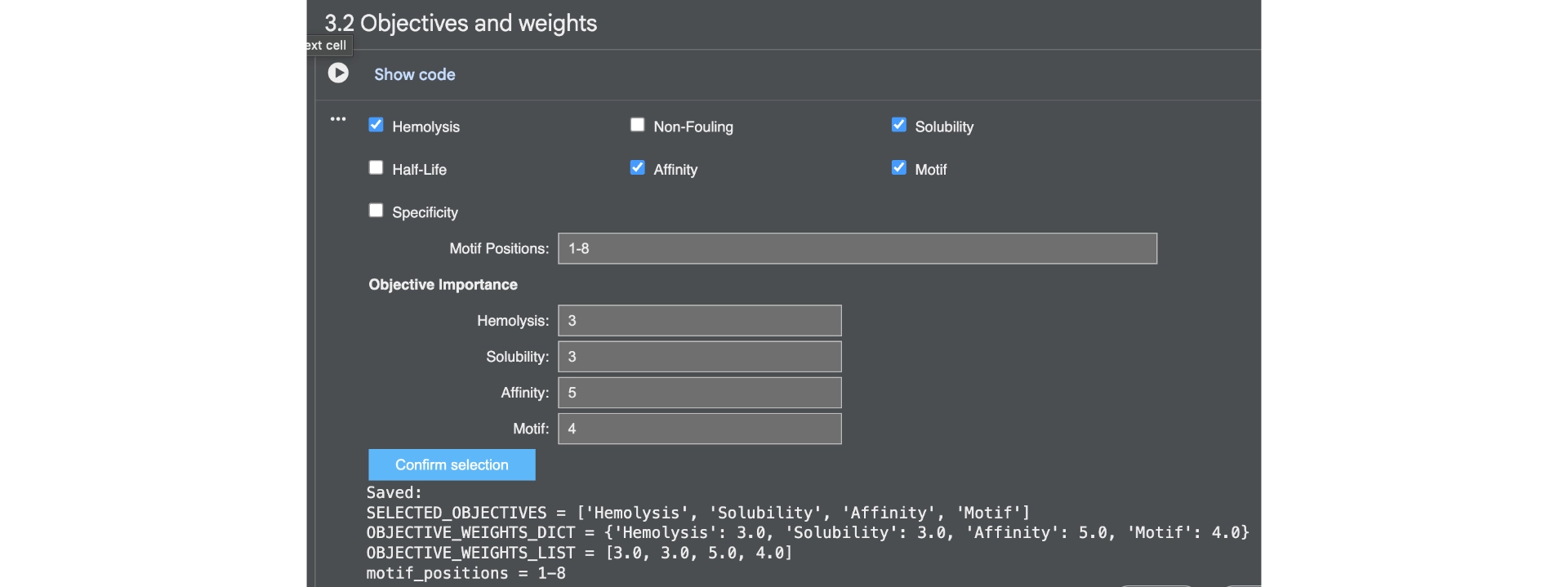

Then analysed with PeptiVerse again.
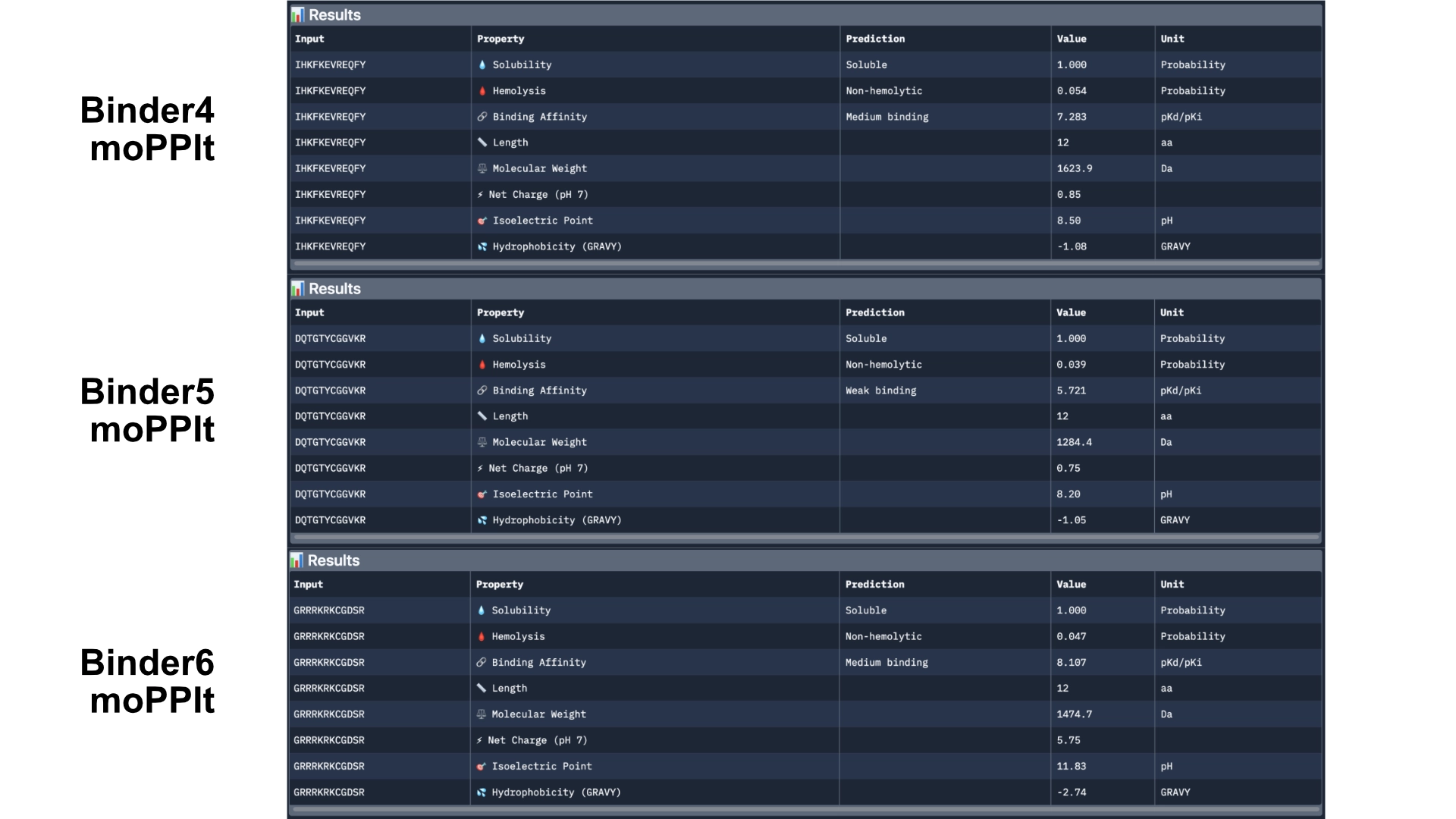

The results came back with low homlytic probabilities, soluble and with good affinity
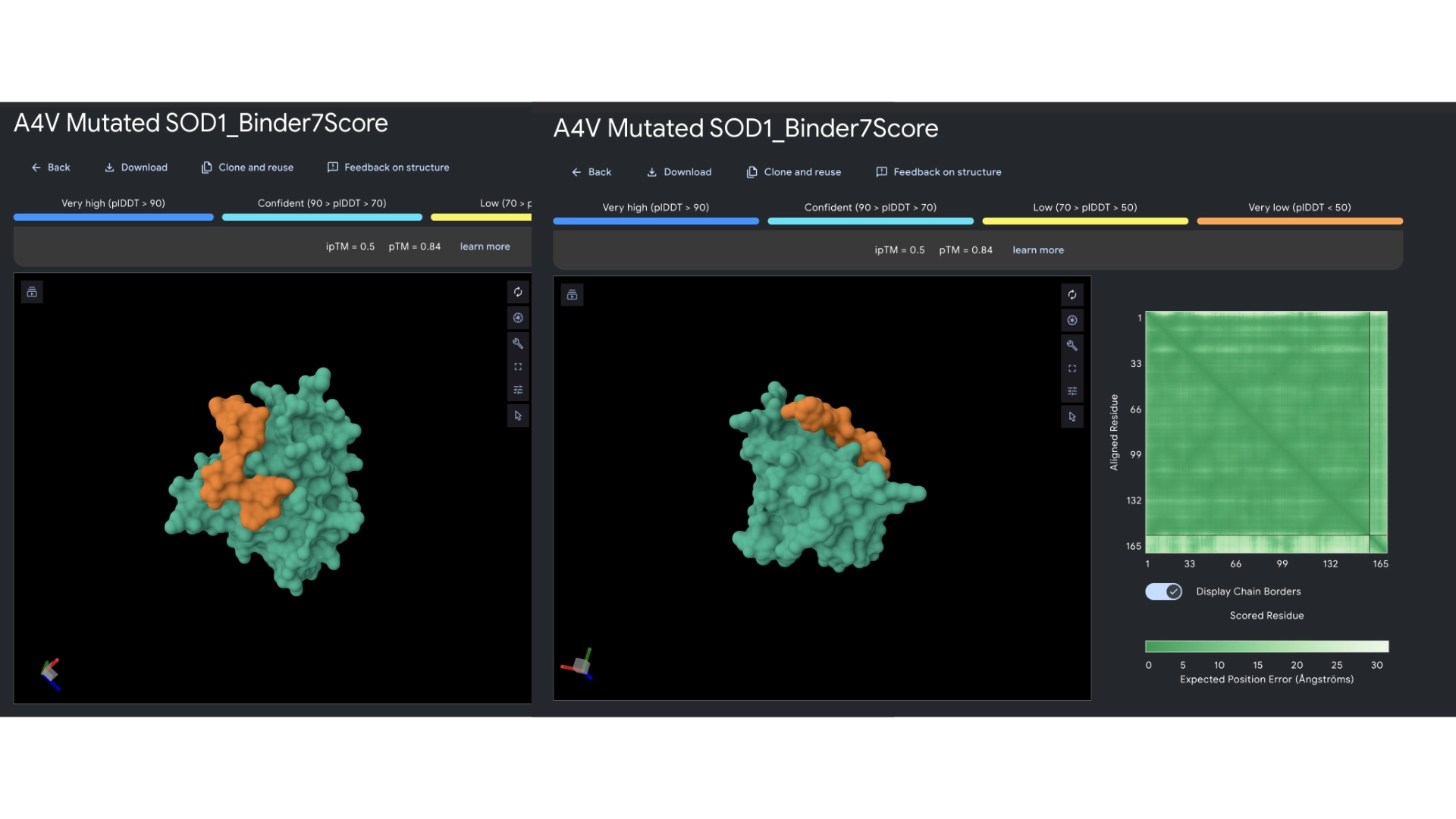
In orther to reach some more conclusions I fed the best 2 peptides— Binders 1 and 6— to Alphafold to see if they would bind closer to the N-terminus which was the motif I had input and see if the ipTM score had gotten any better.
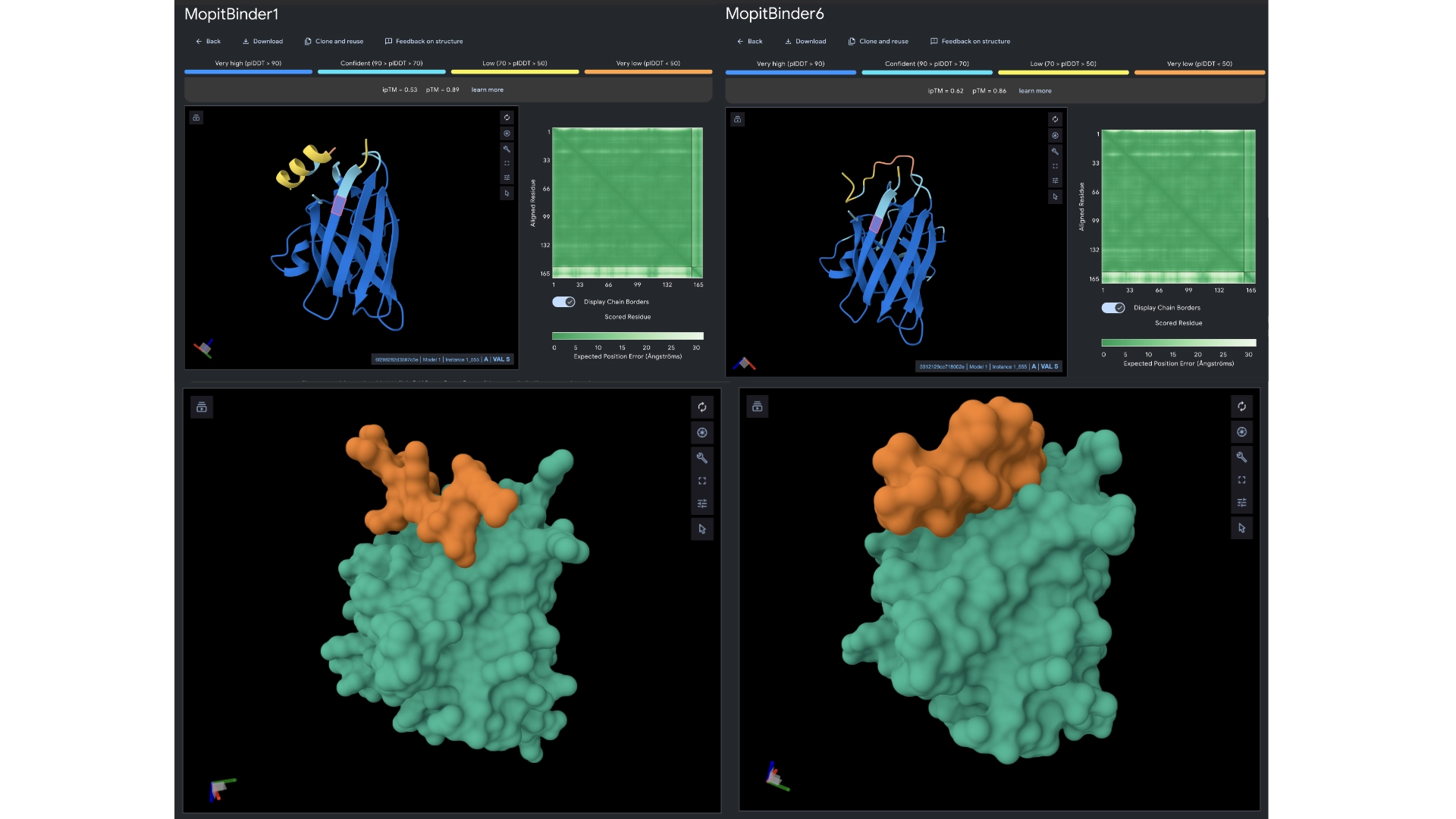

The ipTM scores came back the highest yet, with binder 6 reaching 0.62. They did bind closer to the N-terminus and not to the barrel region. Binder 1 did form a helix in the visualization and this might be interesting since a helical structure is more stable and could be further improved to have really high binding affinity. Then ran them against more therapeutic parameters— non-fouling, halflife and permeability— to see how these would hold up even not having been optimized for those purposes.
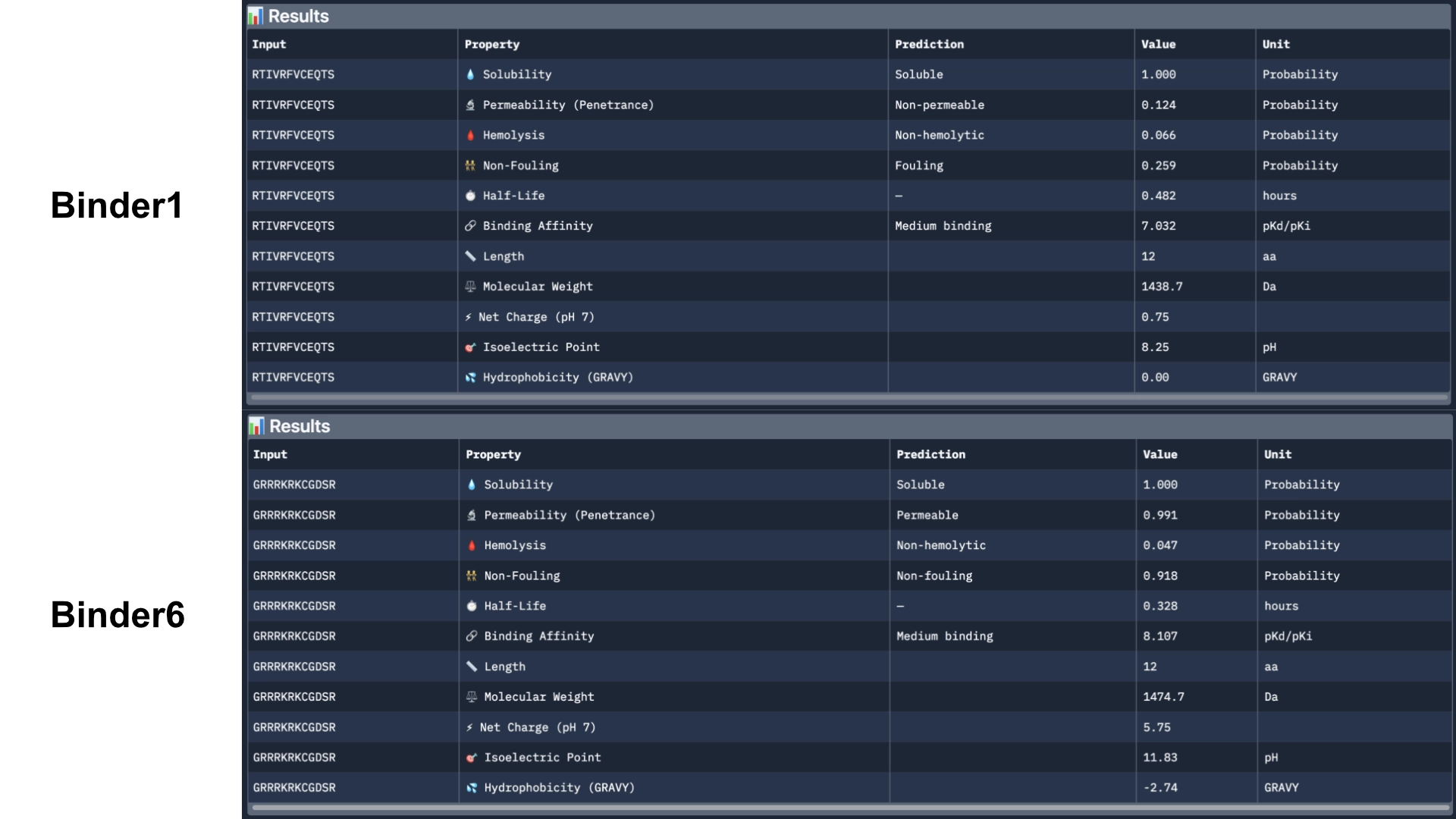

Binder 1 did come back as fouling and non-permeable. However, Binder 6 came back as permeable, non-fouling and with a half-life of 0.328 (which might be on the lower side). I would further optimize these 2 best peptides for the different theurapeutic qualities where they are weakest— fouling and permeability and half-life for binder 1 and half-life for binder 6— using a MOG-DMF model and run them against those parimeters using PeptiVerse.
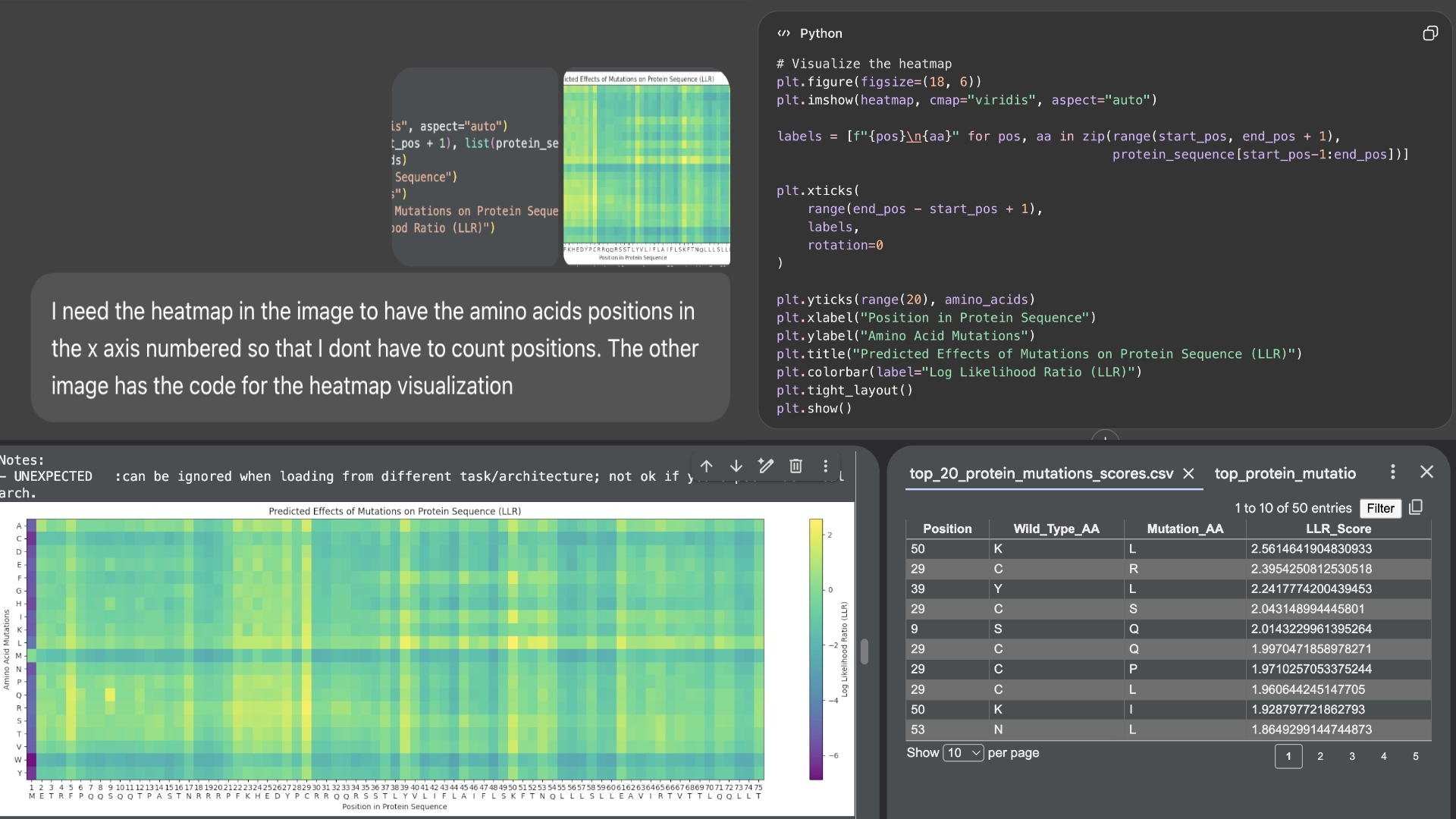
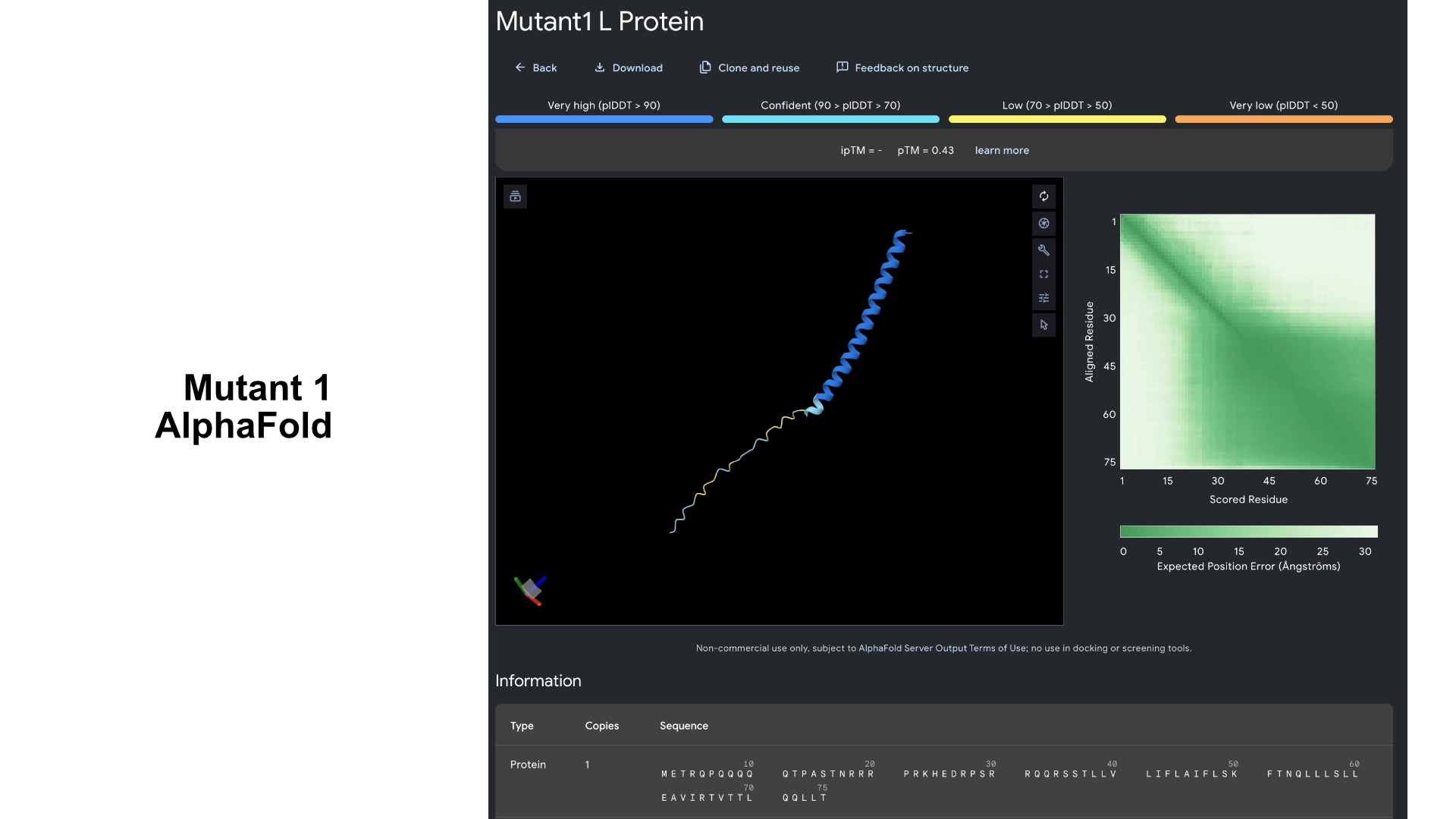
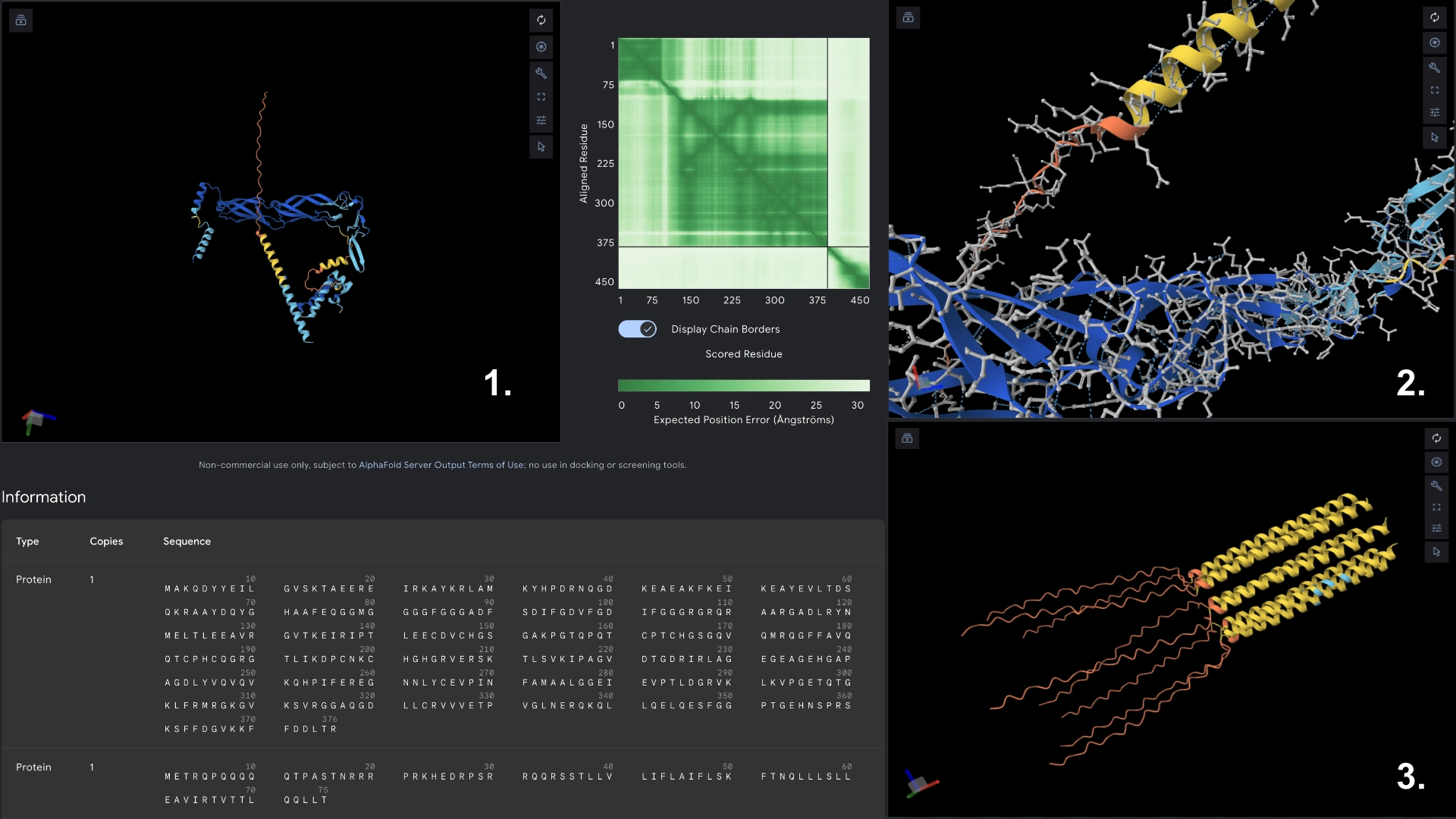
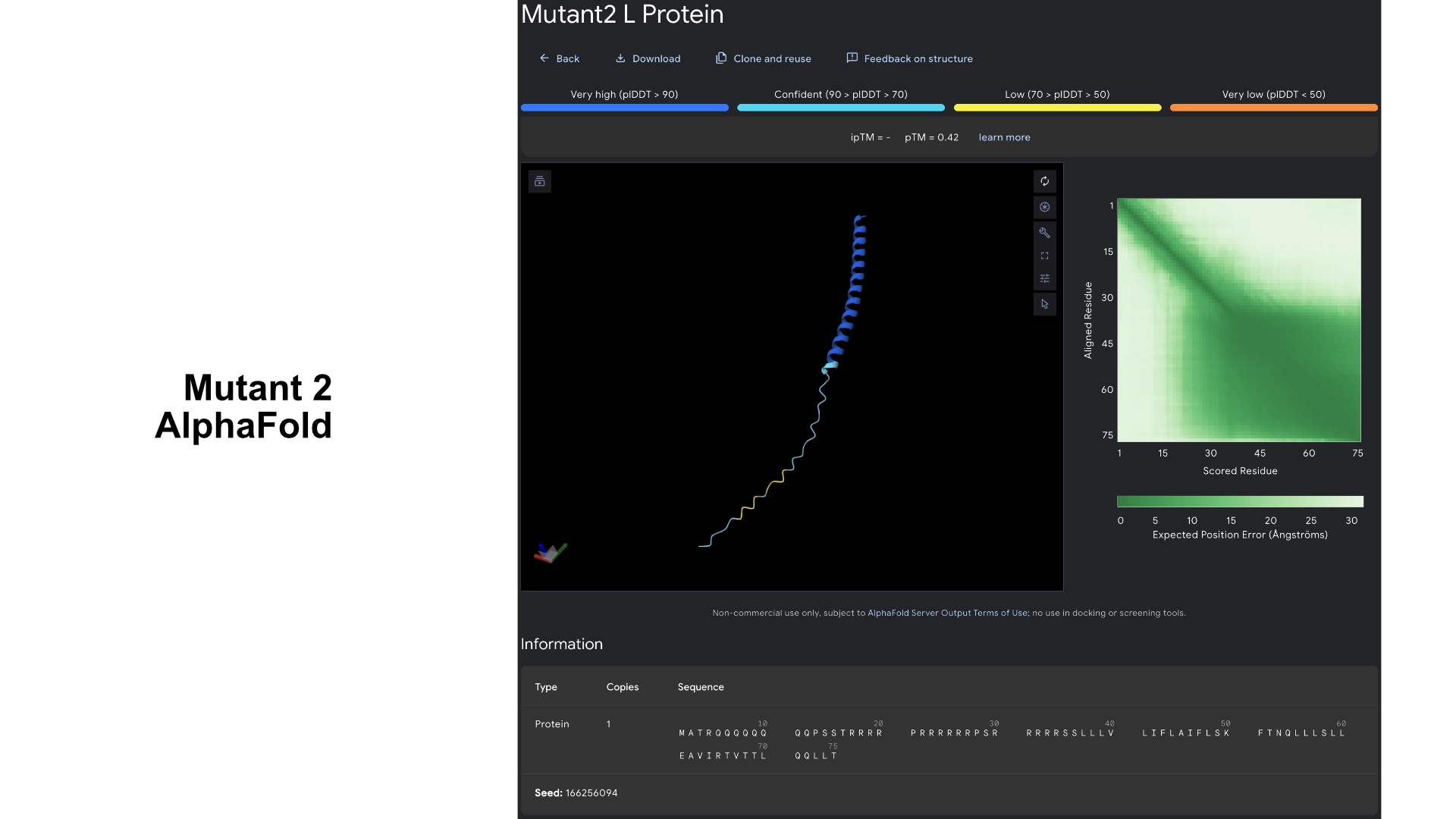
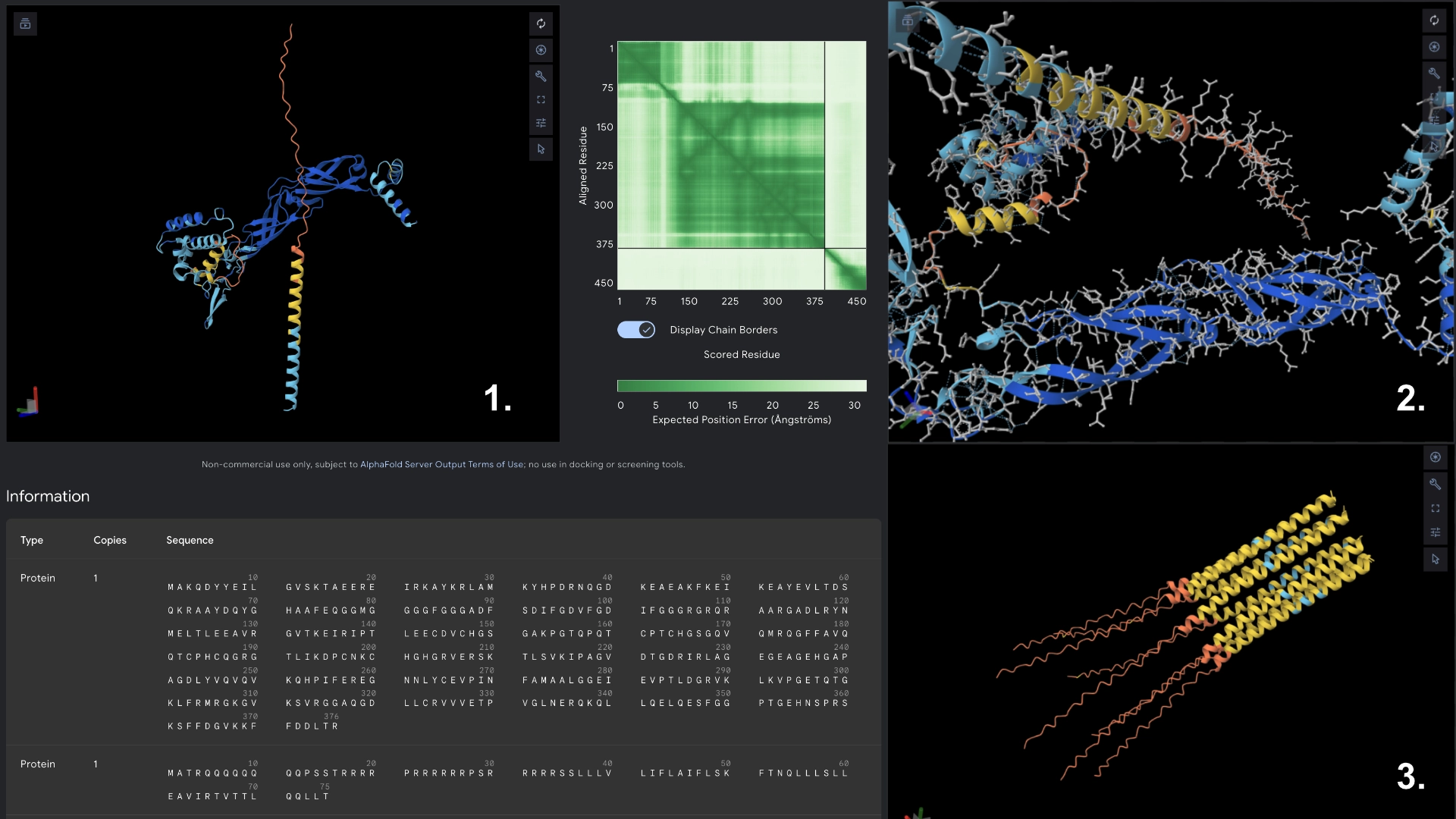
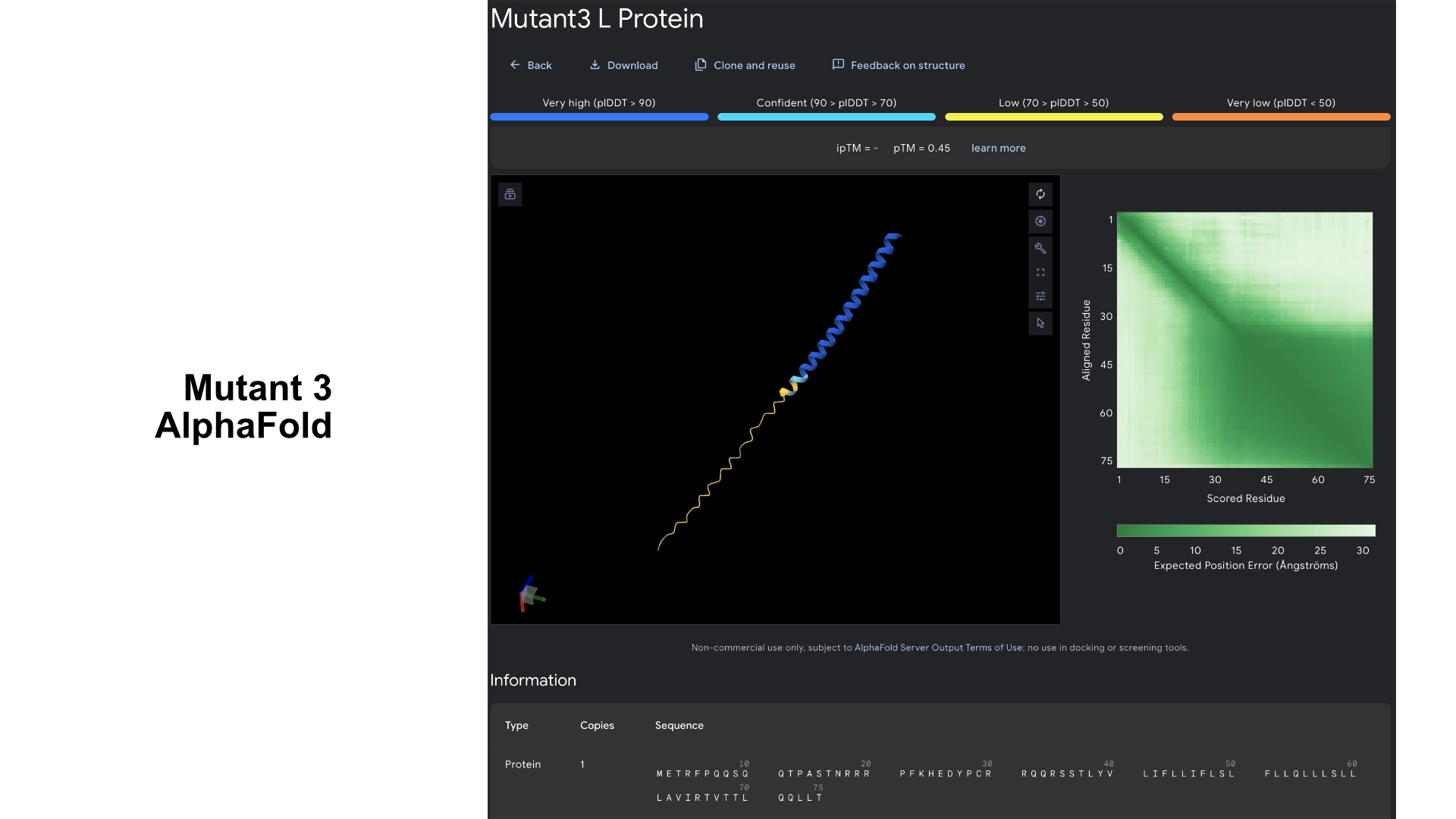
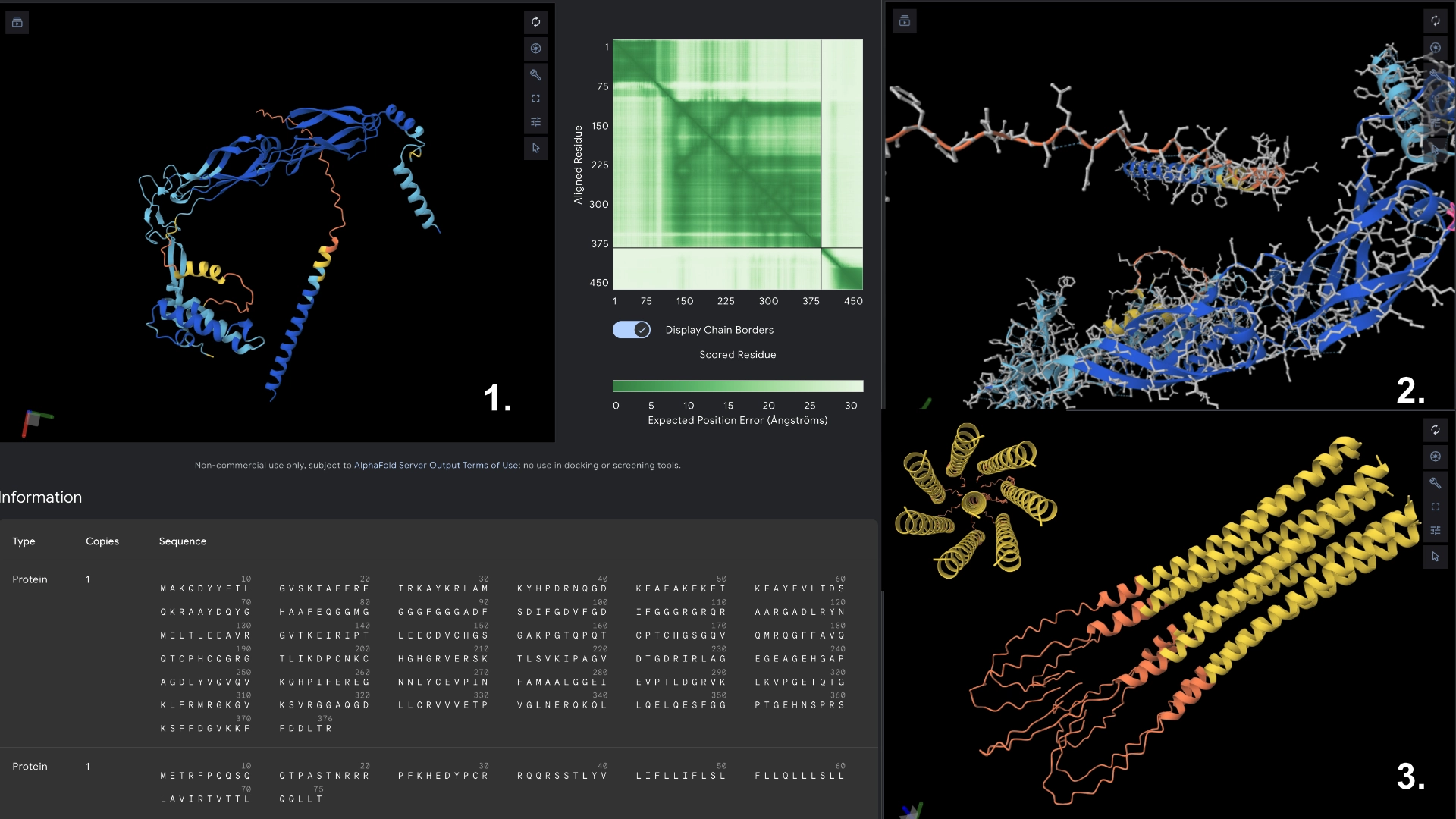
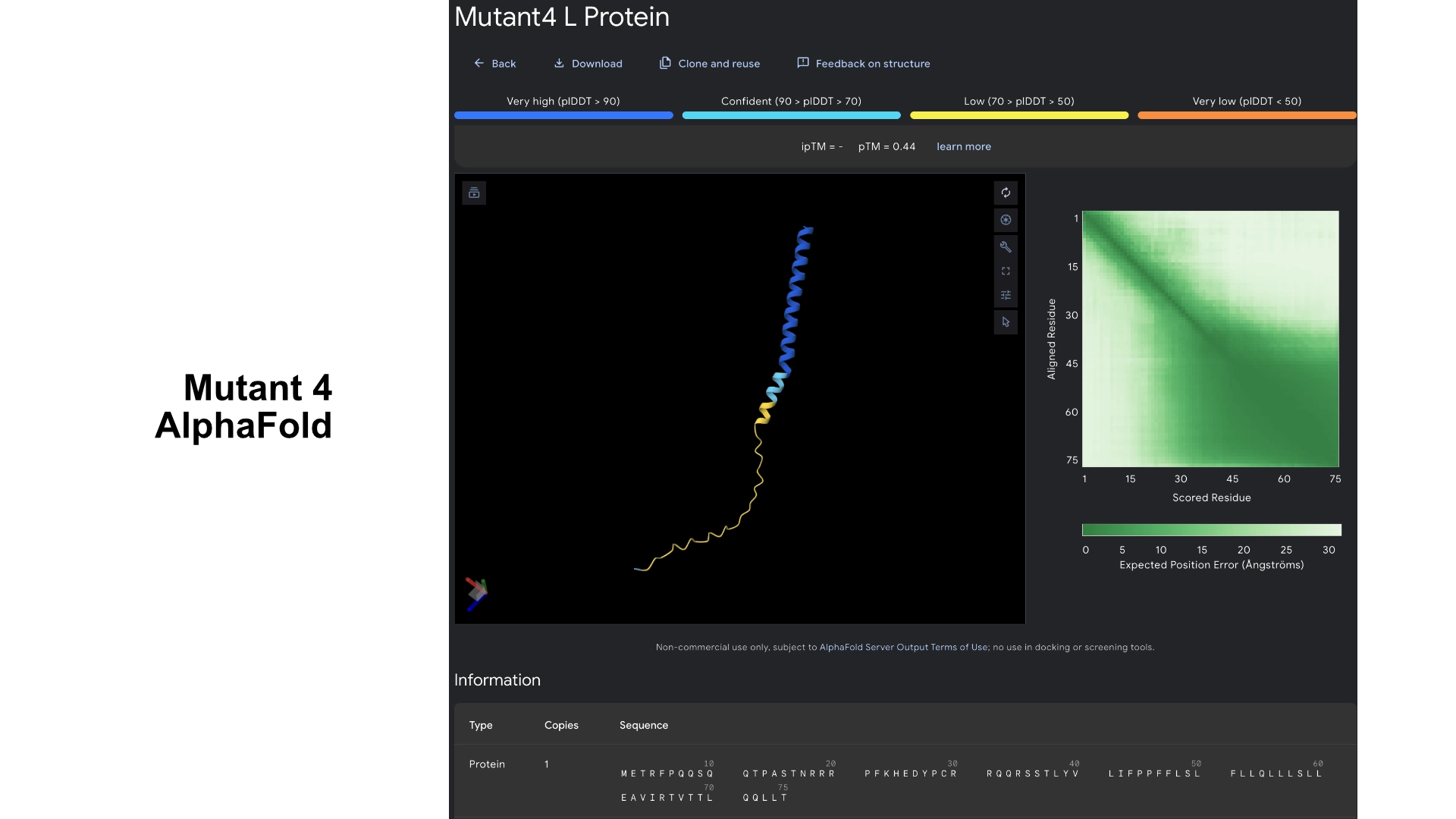
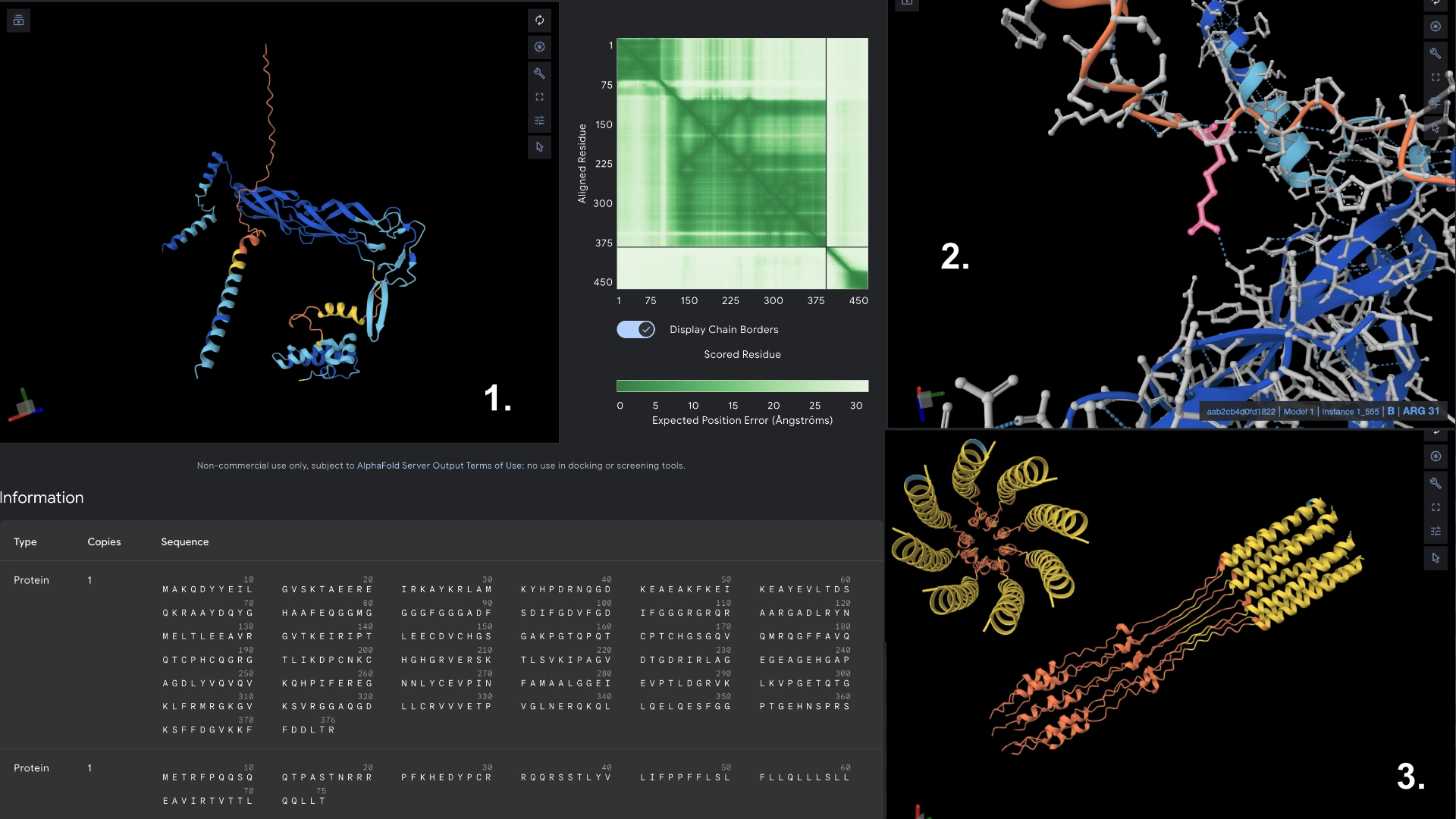
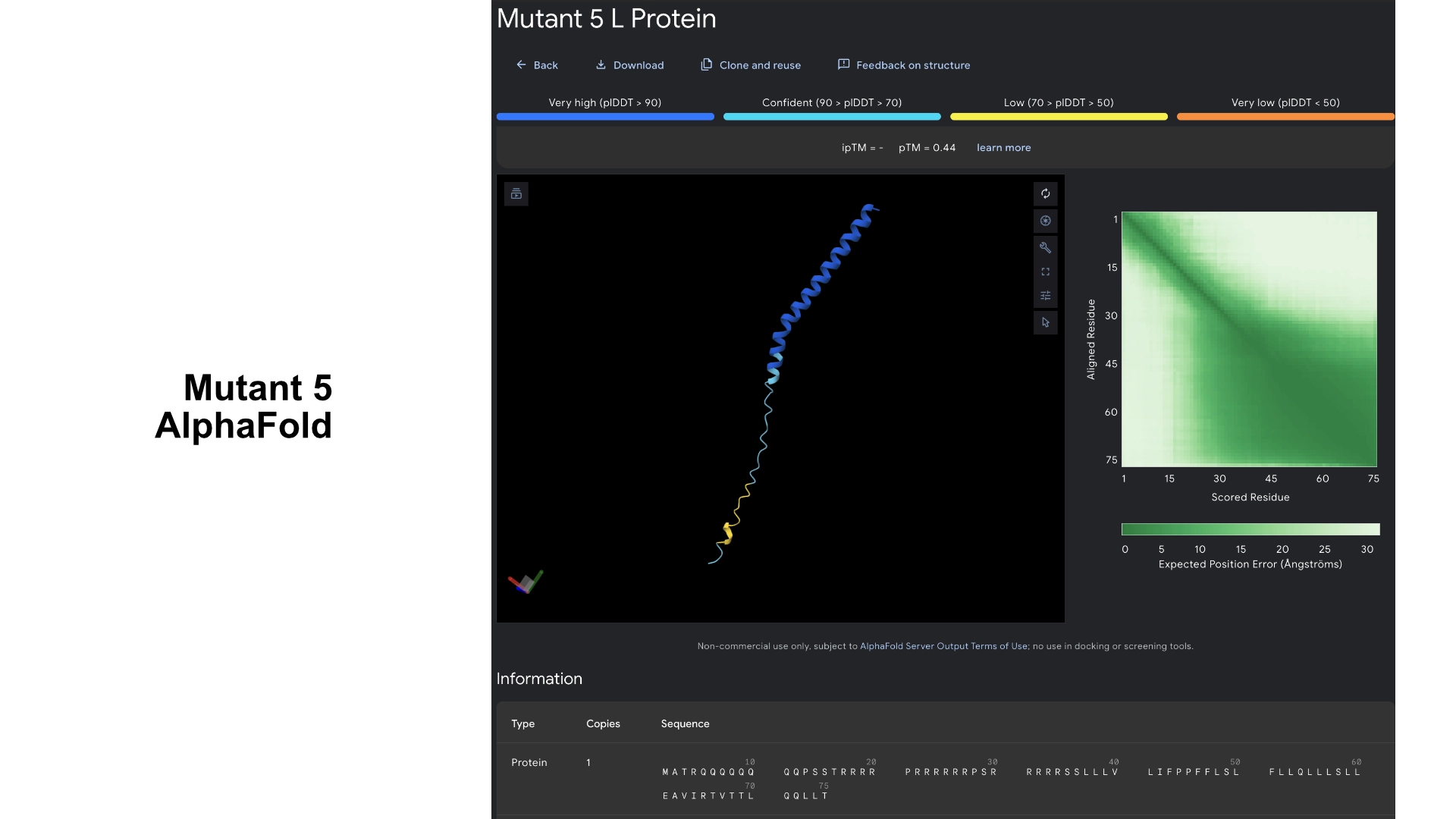
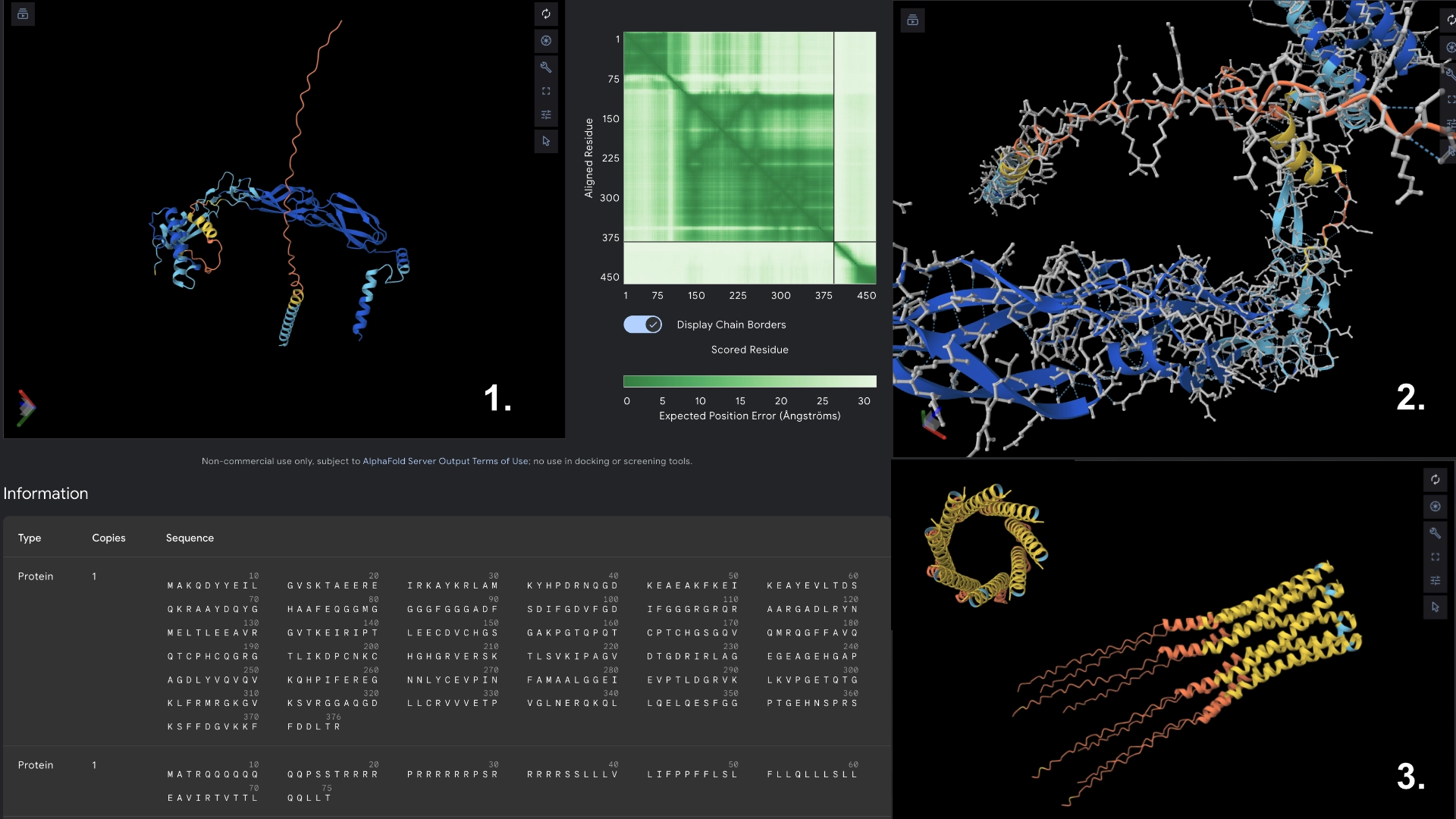
Part C: Group Project: L-Protein Mutants
I started by making 2 mutants with several mutations to the hydrophilic domain, would be interesting to make it more stable and not dependent on DnaJ, by making changes in the positions best scored by the LLR and confirming them with the experimental data.