Week 6 HW: Genetic Circuits Part1

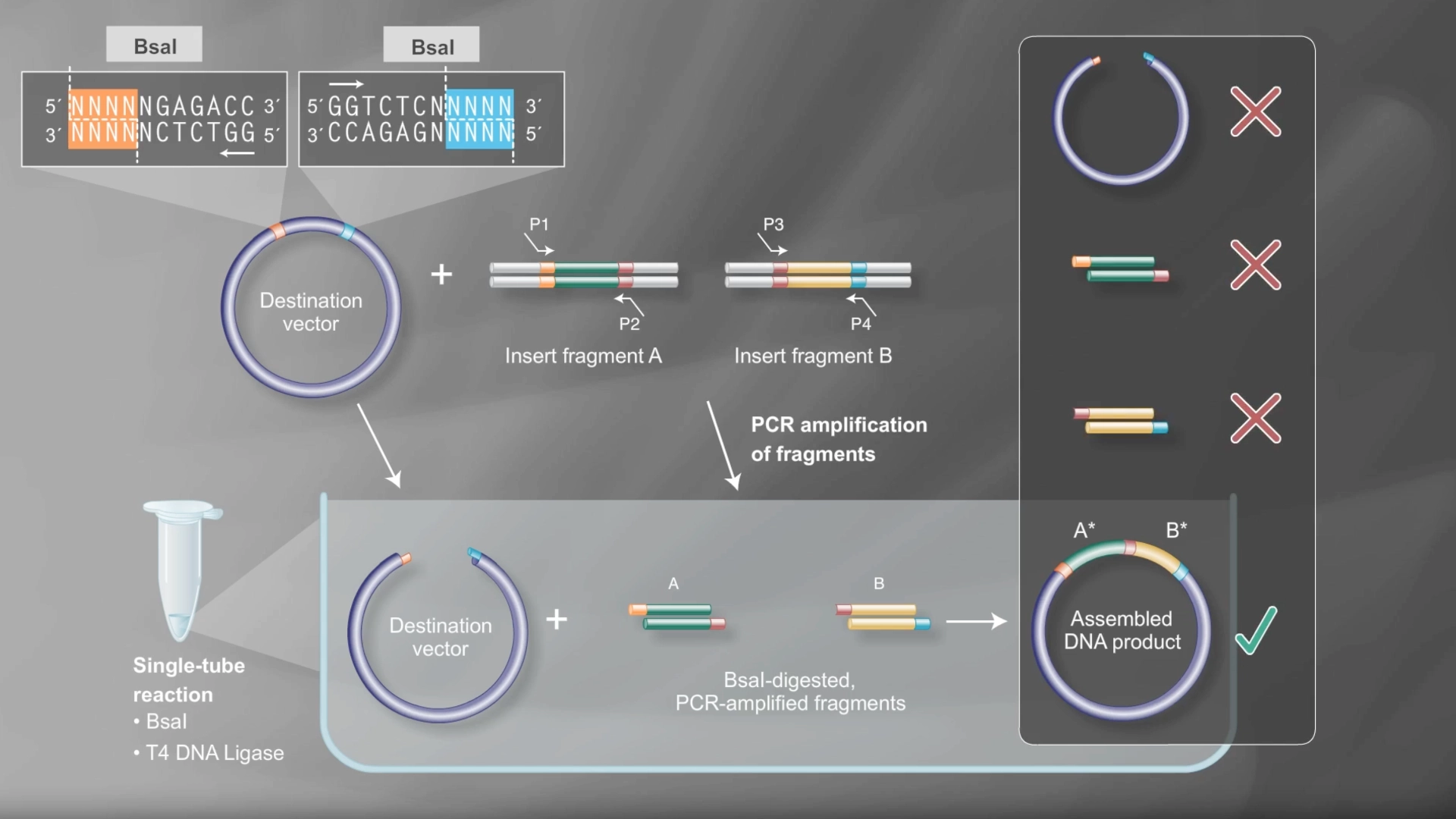
Assignment: DNA Assembly
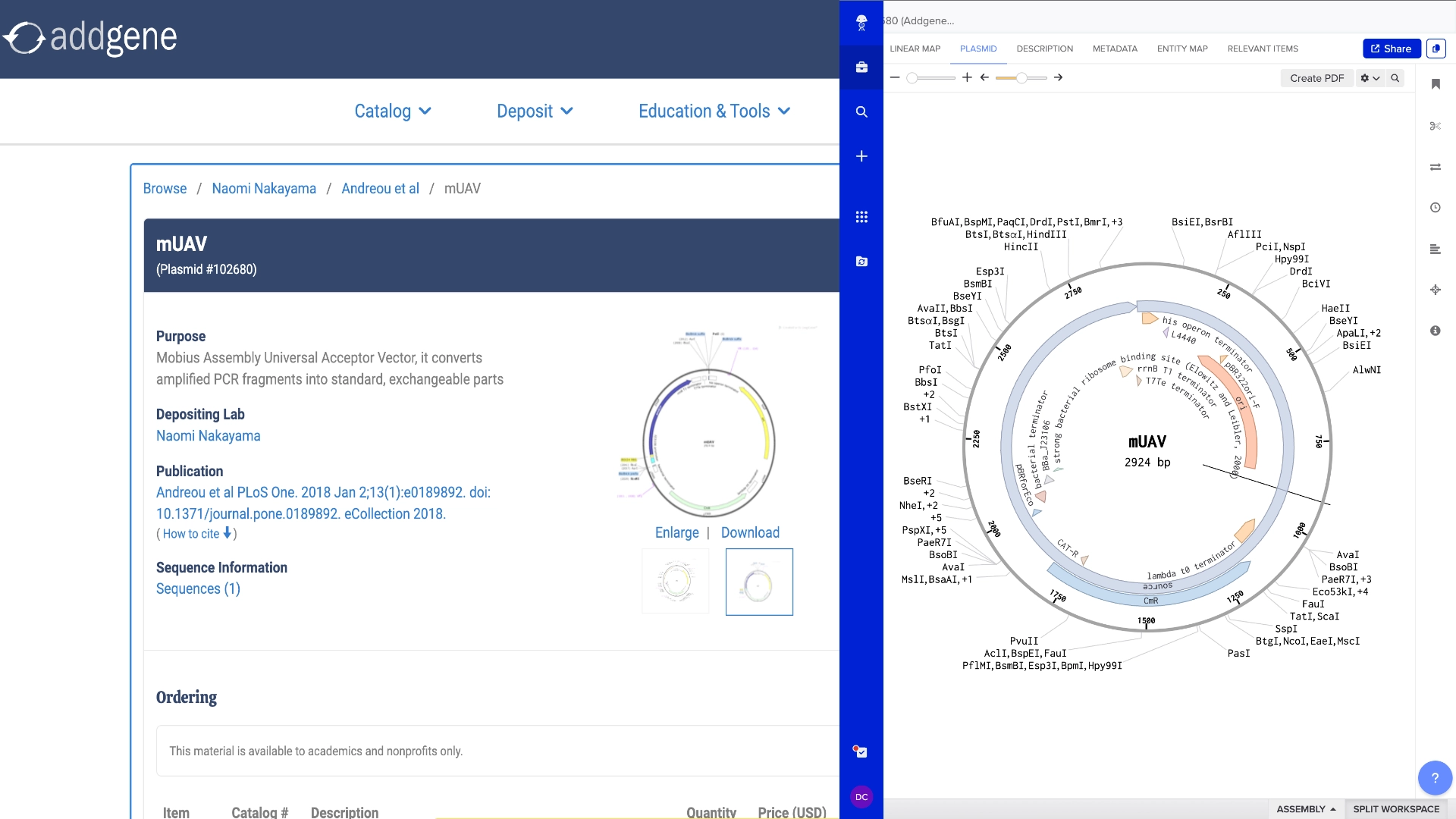
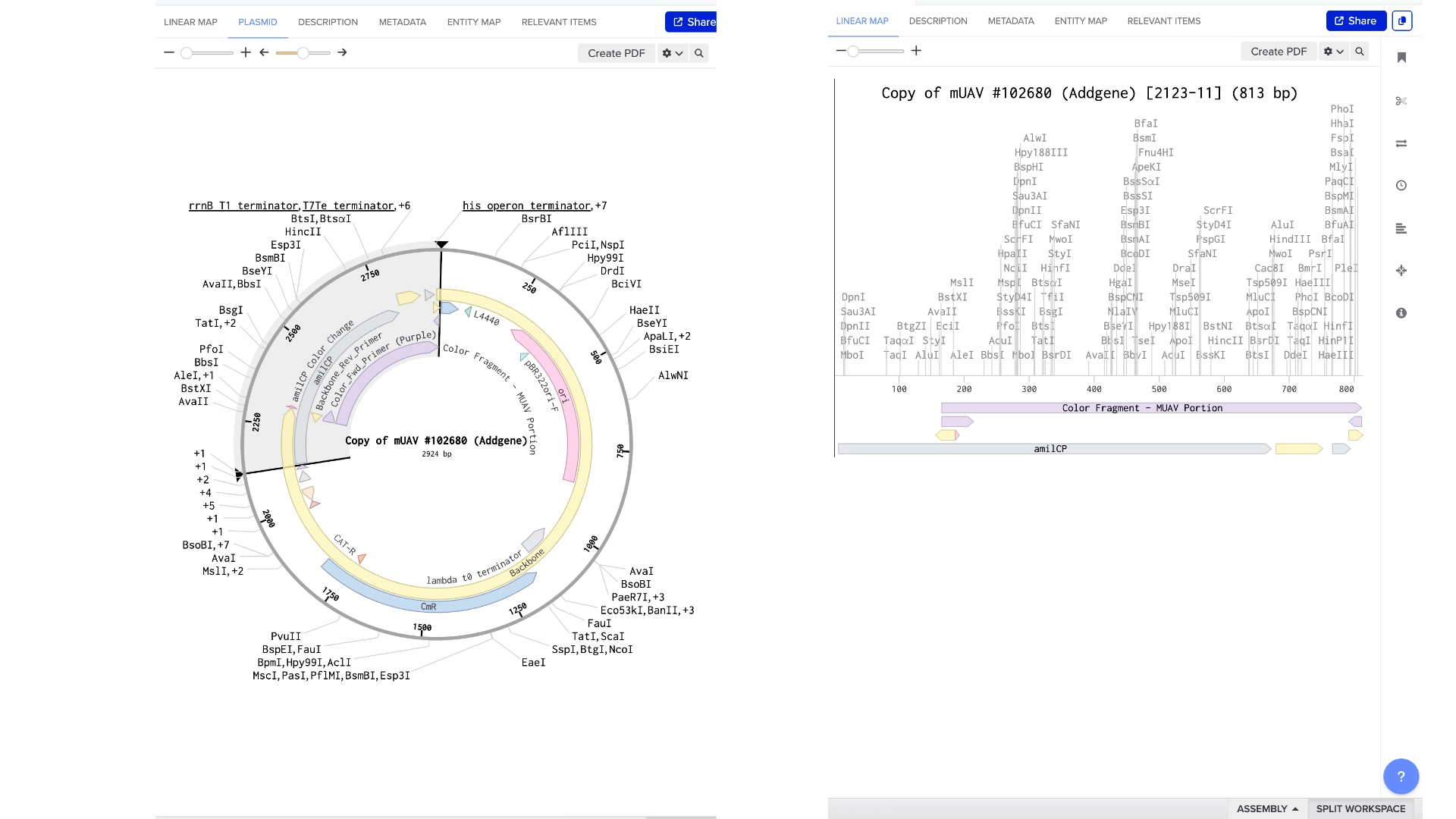
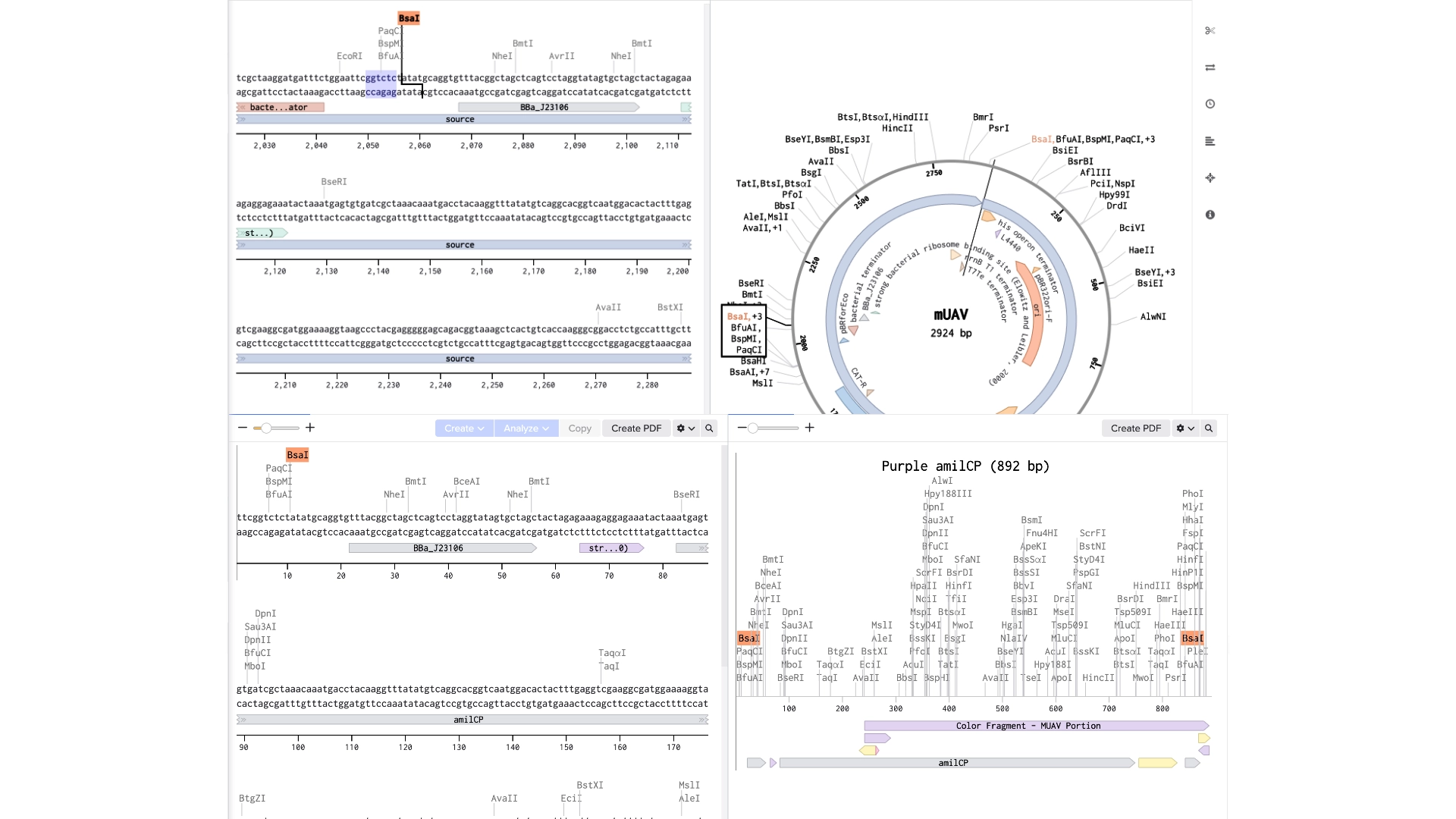
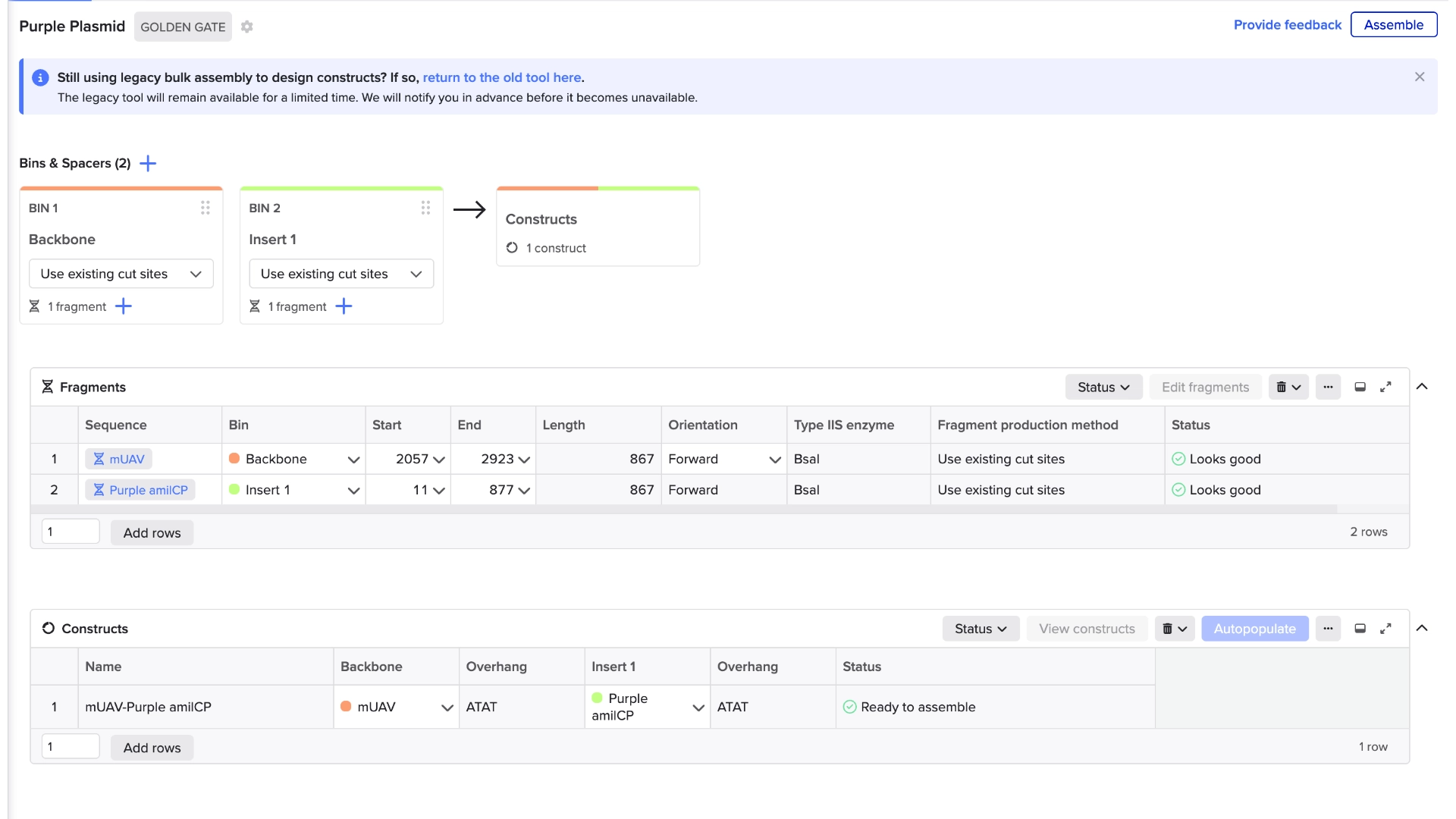
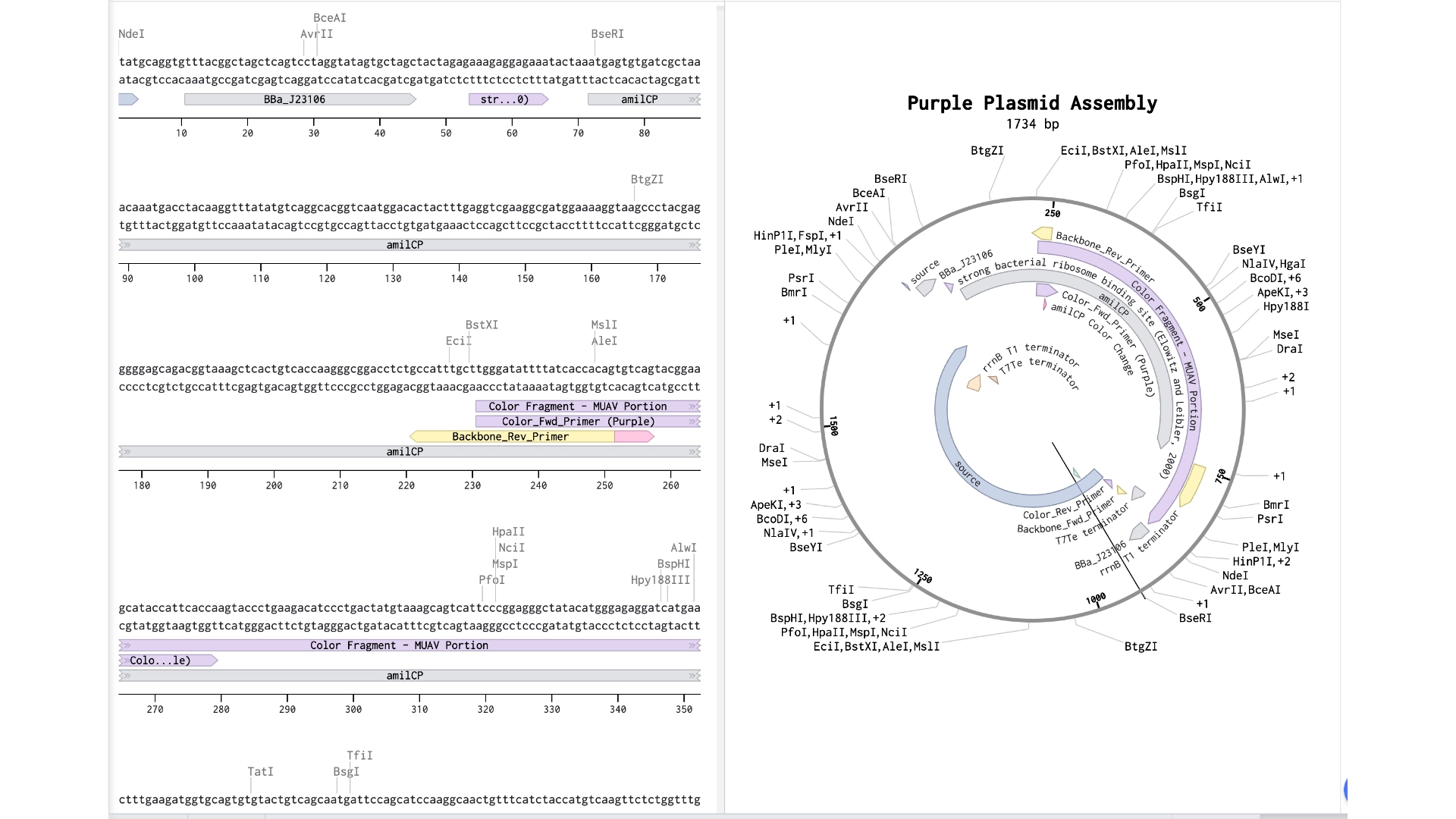
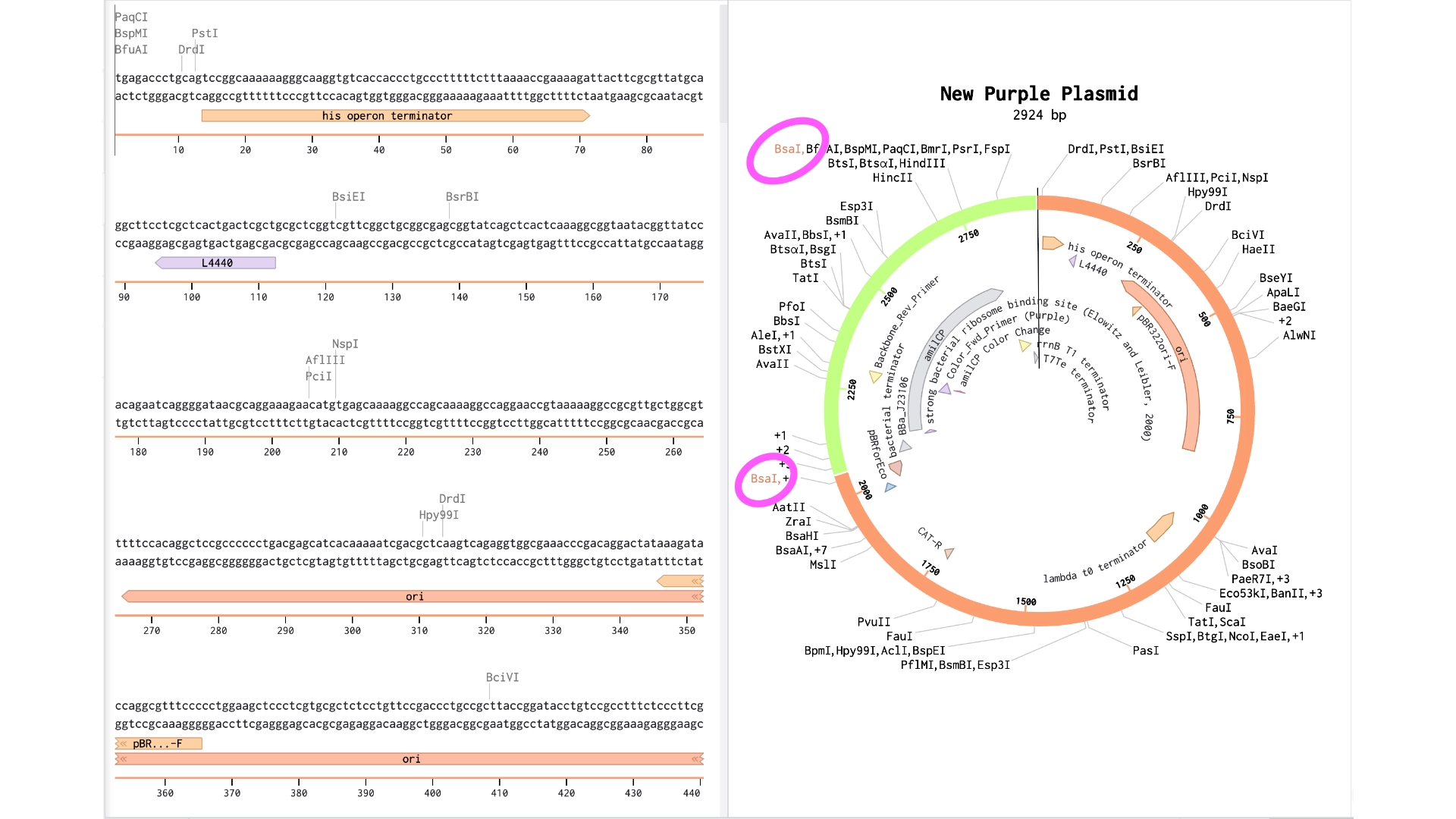
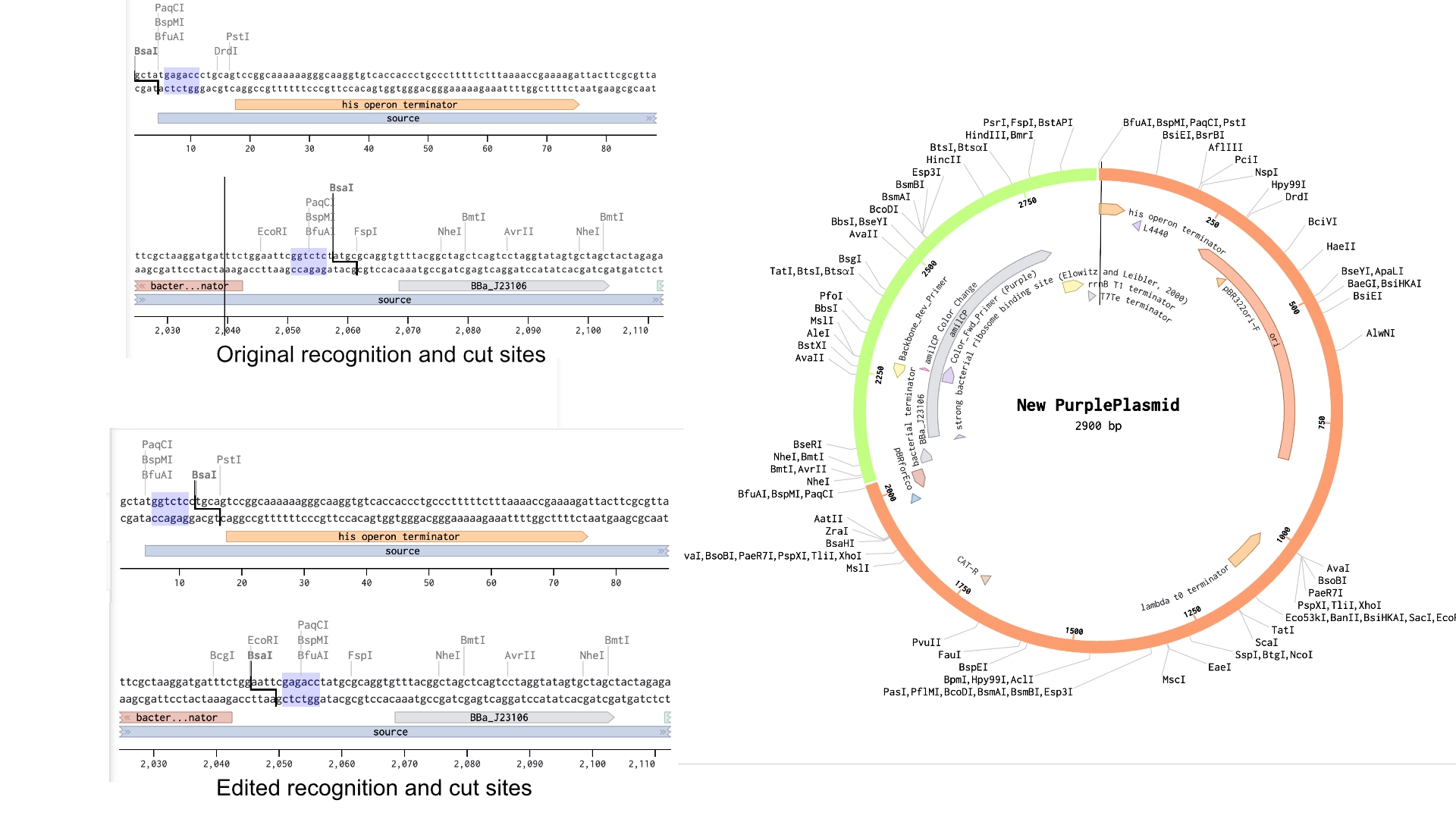
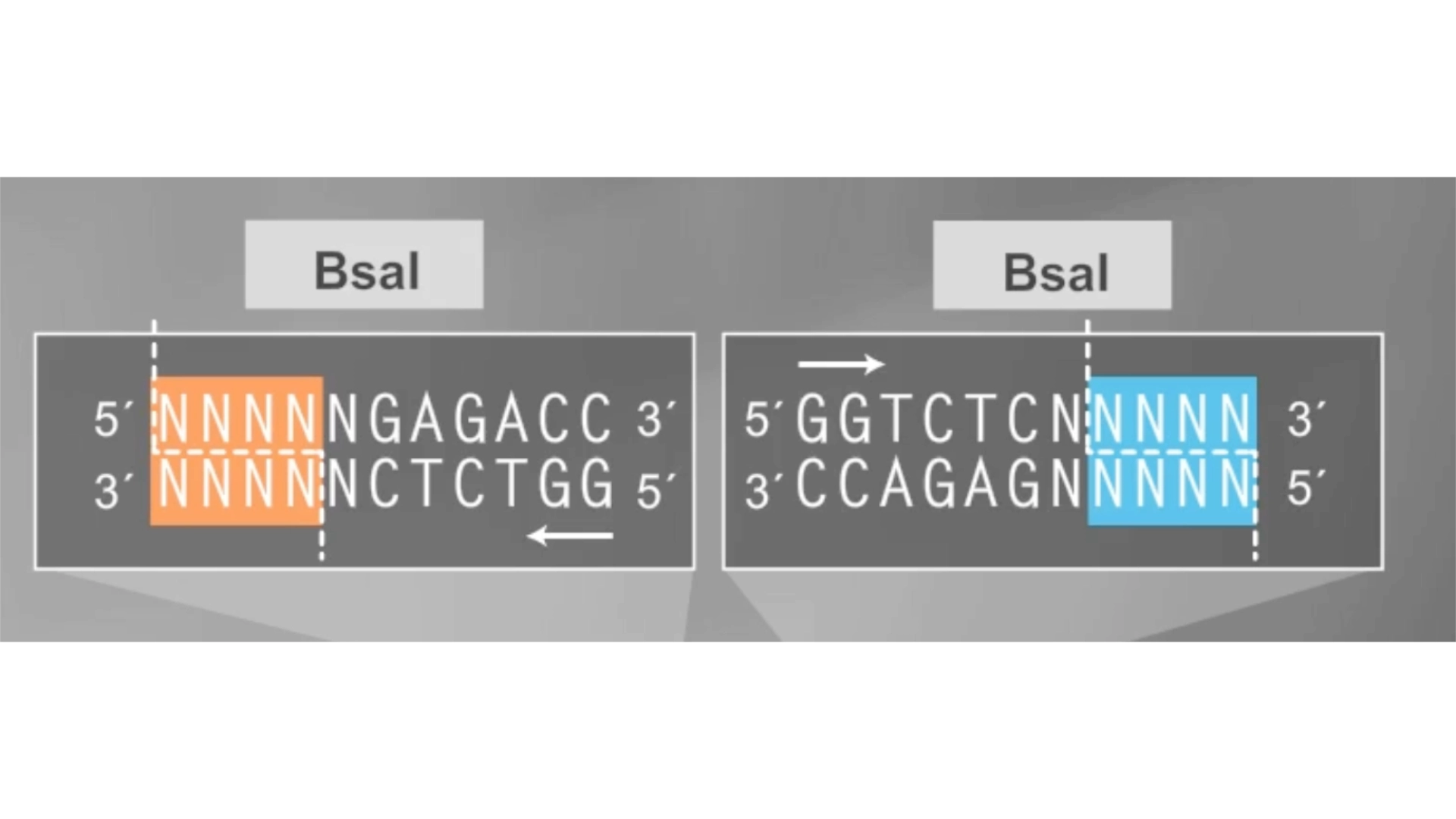
6.2 Model this assembly method with Benchling or Asimov Kernel!
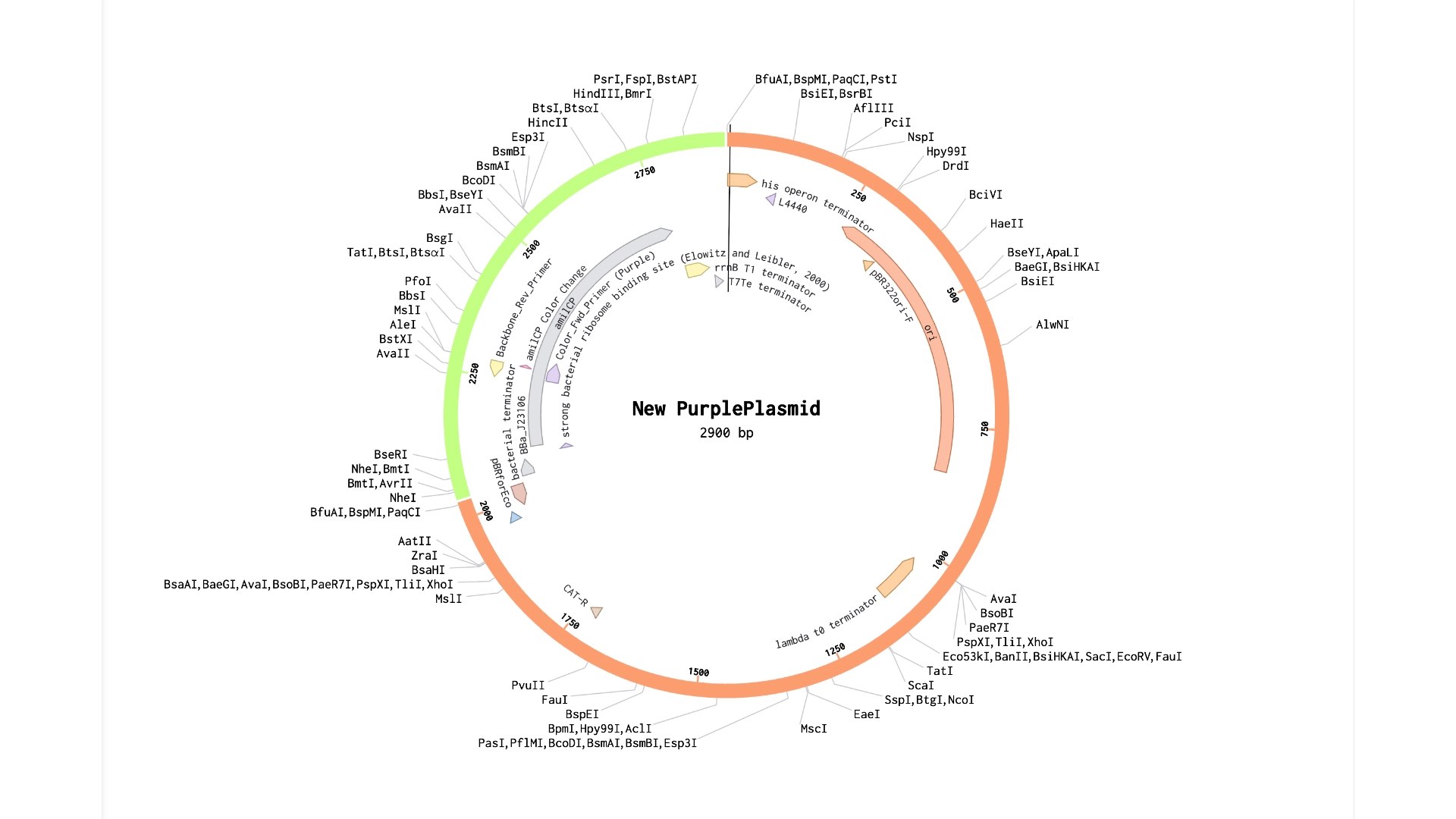

Final result of assembling a mUAV backbone with an insert of purple amilCP
Assignment: Asimov Kernel
Repressilator:
Glyph
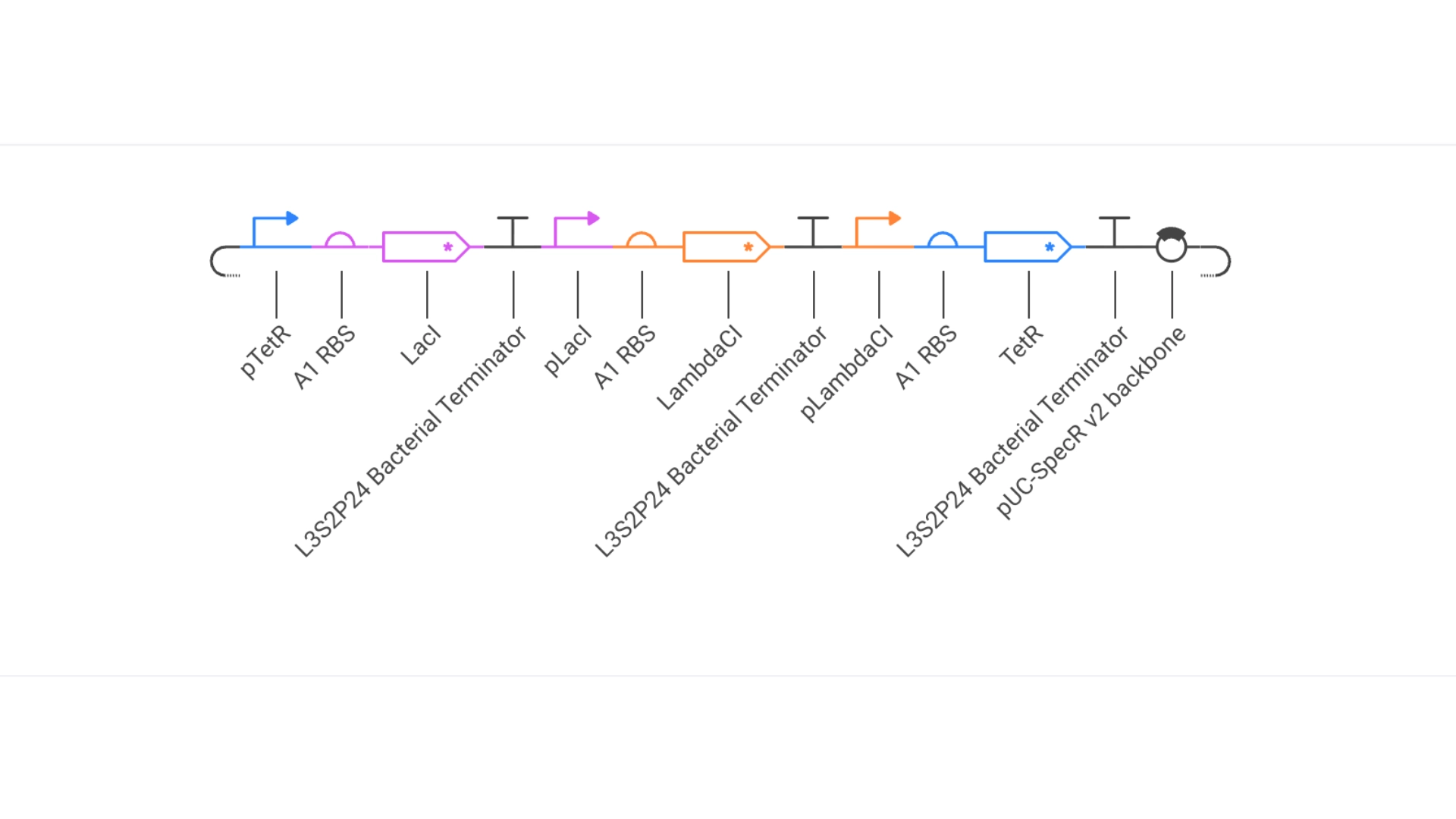
 My Repressilator
My Repressilator
Simulation
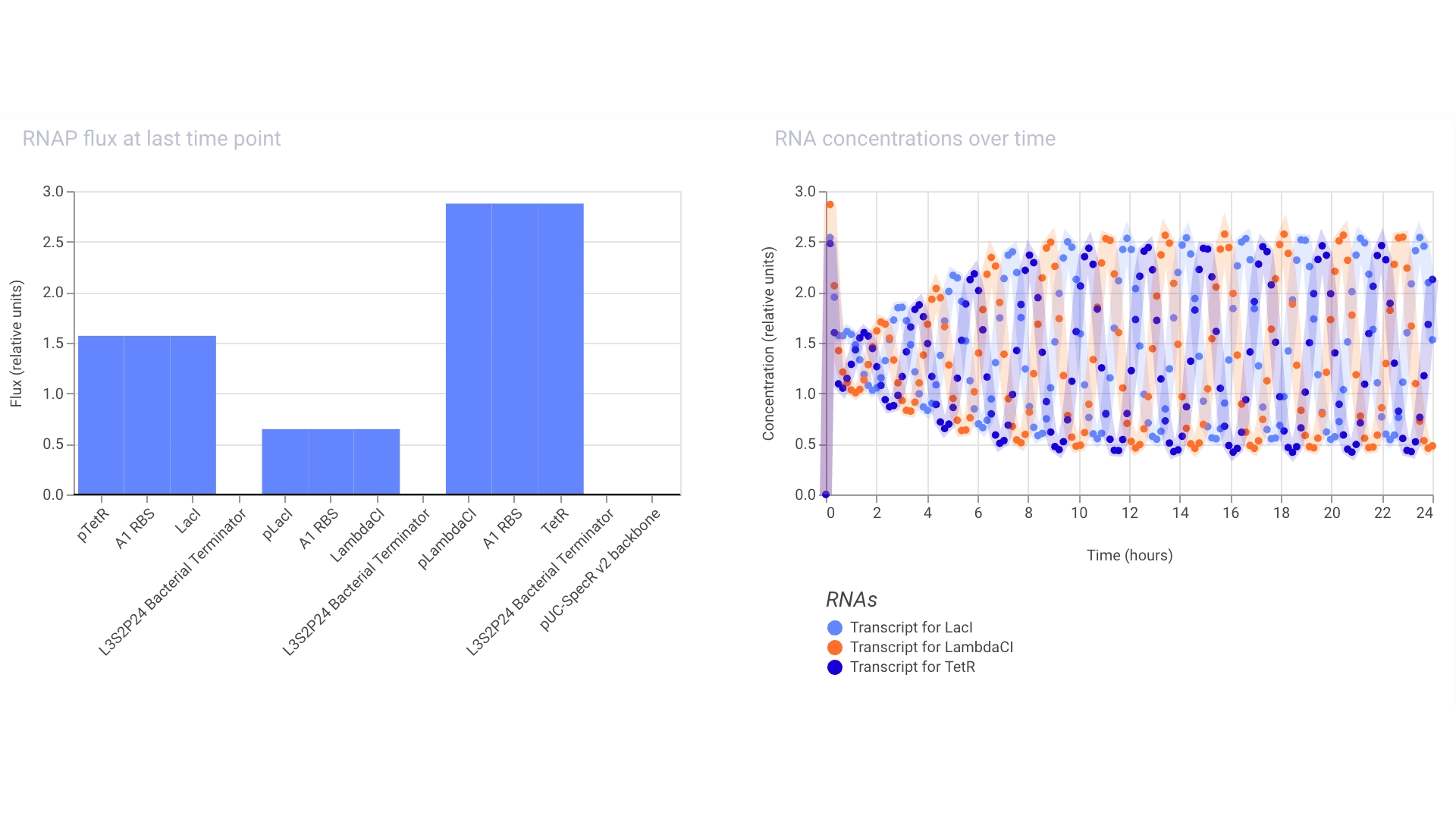
 Proteins expressed by the Repressilator seem to alternate in the expected way!
Proteins expressed by the Repressilator seem to alternate in the expected way!
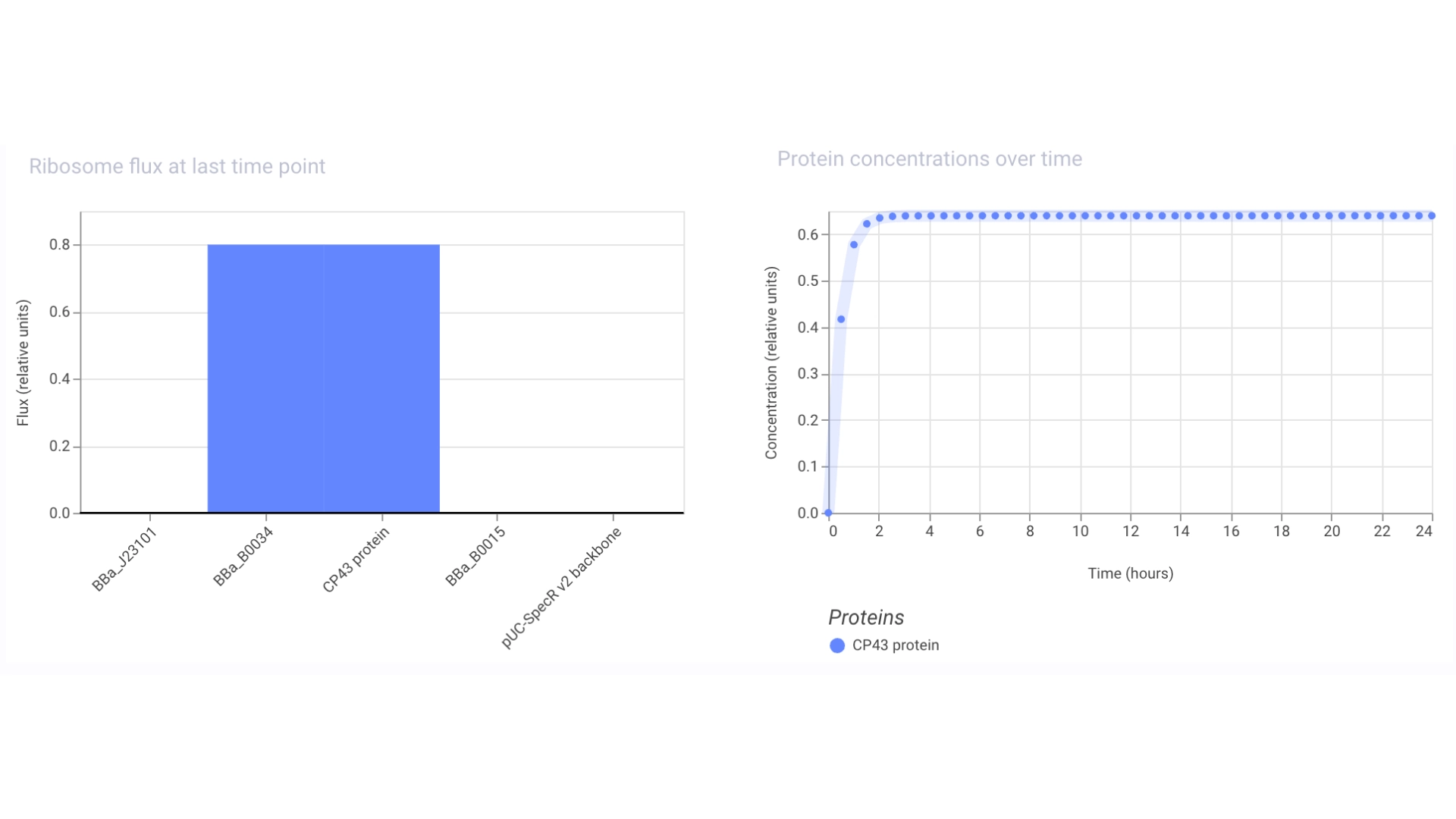
Construct #1:
Objective: Create a simple construct that expresses the chlorophyll binding protein CP43
For this construct I created a new part — a CDS part coding for the CP43 protein
Then assembled the circuit using a strong promoter, RBS, terminator and pUC-SpecR v2 backbone
At first only the RNA for the protein was being expressed but not translated into the actual protein, then I found out that the construct had been created for Chinese hamster instead of E. coli
Glyph
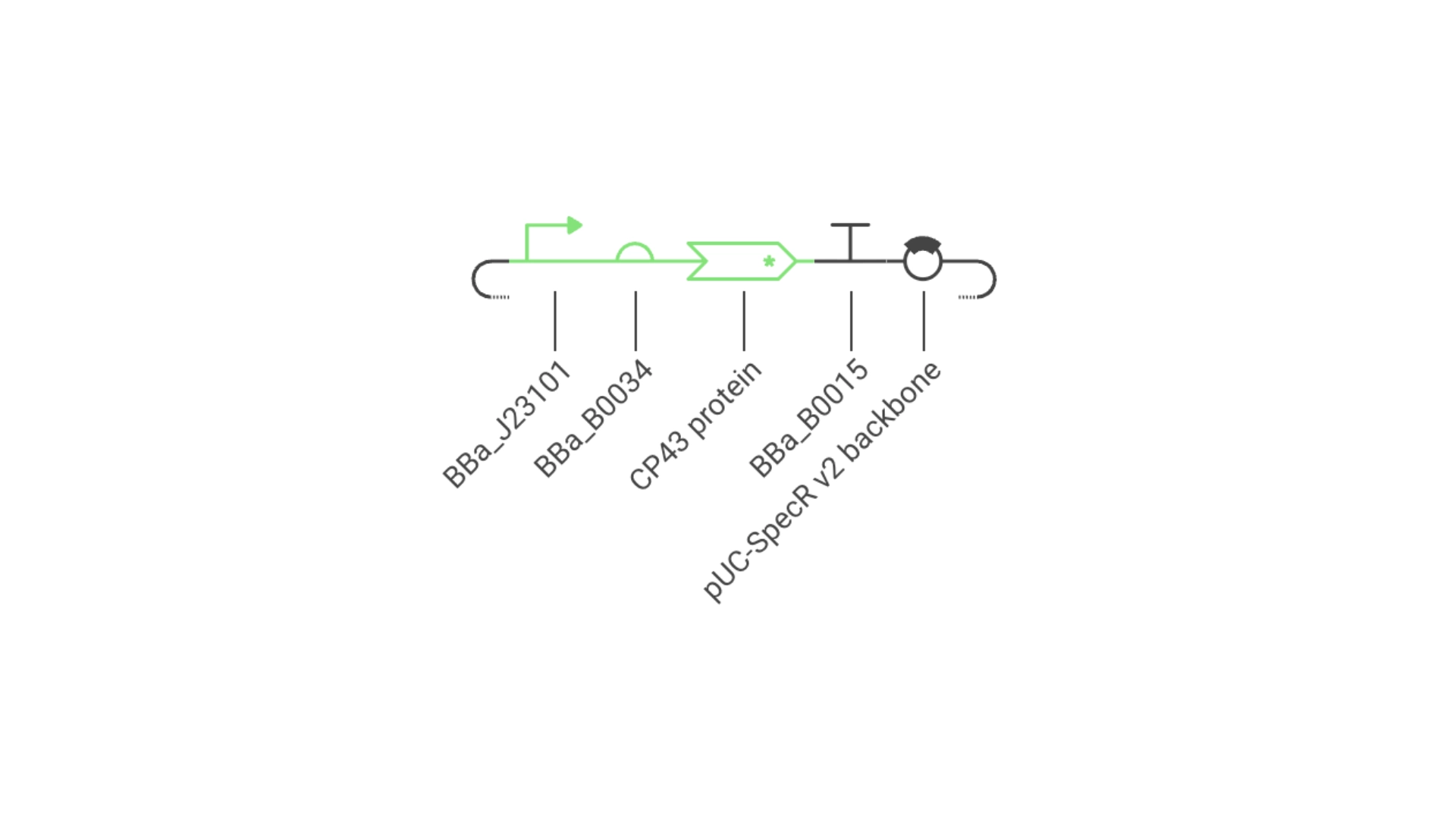
 Construct #1
Construct #1
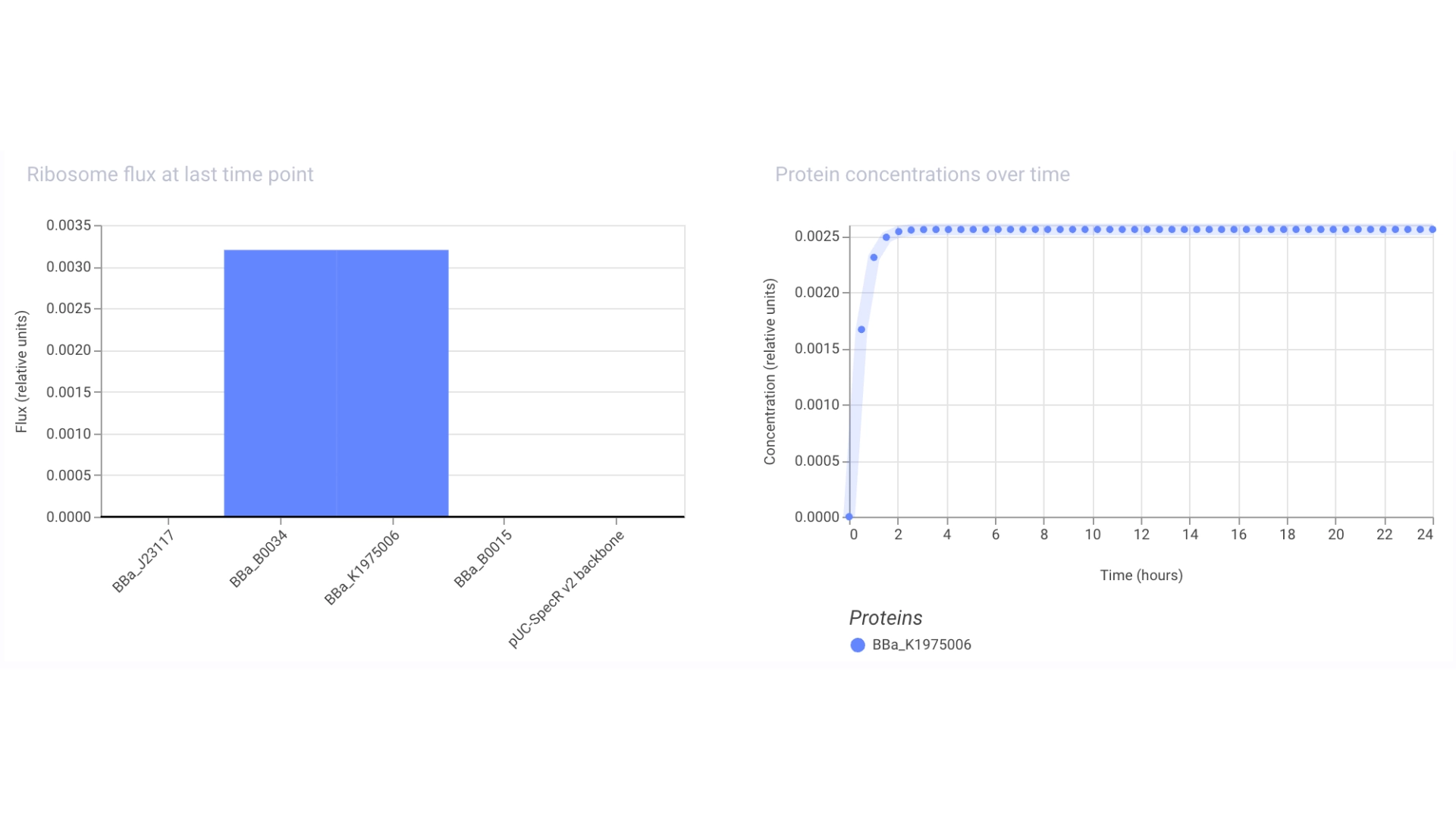
Construct #2:
Objective: Create a simple construct that expresses a Red Fluorescent Protein
For this I had to change the RBS part I was using for the CP43 protein, presumably because the CP43 is a CDS (-Start,+Stop) and the RFP is CDS (+Start,+Stop)
Glyph
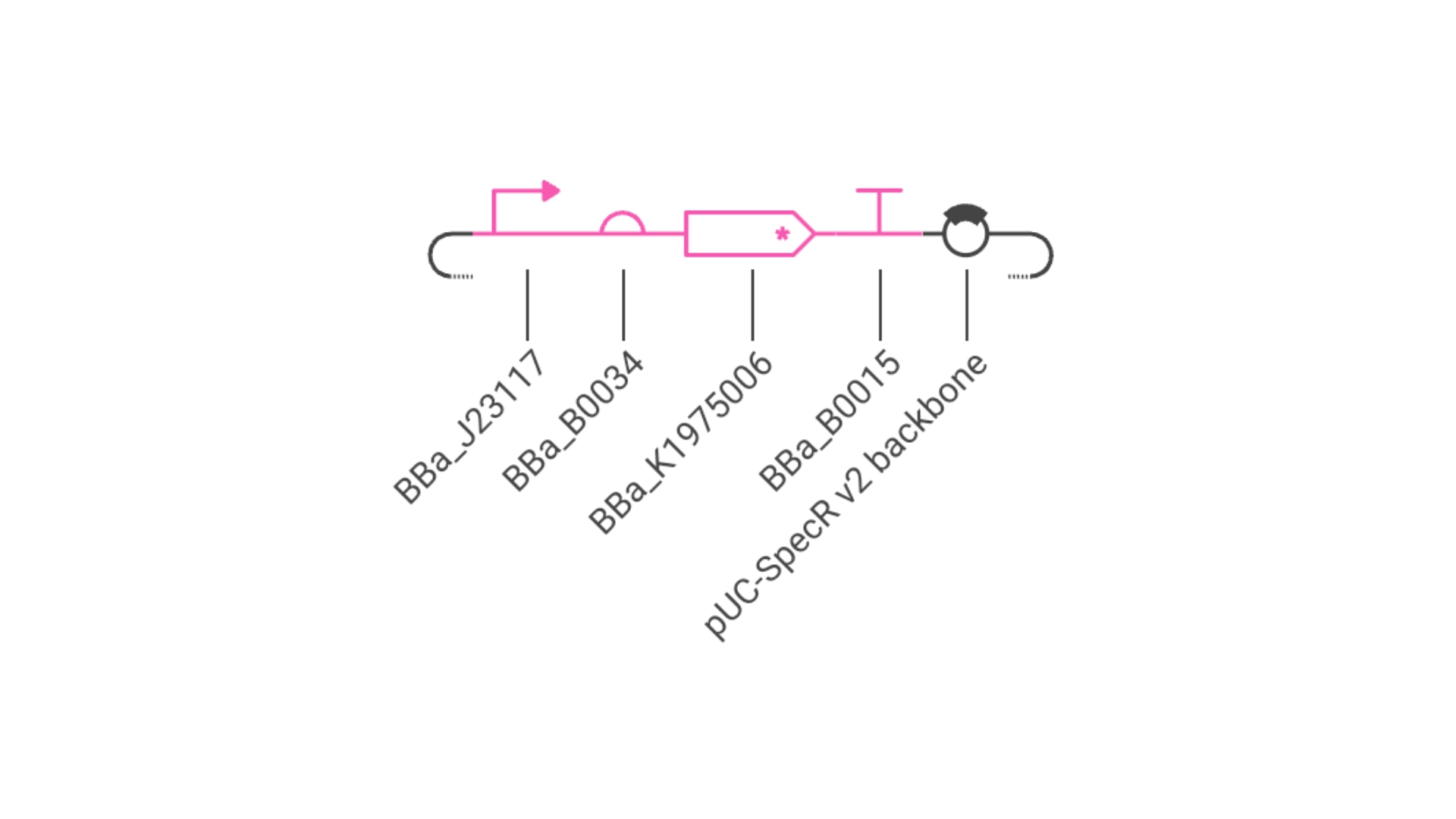
 Construct #2
Construct #2
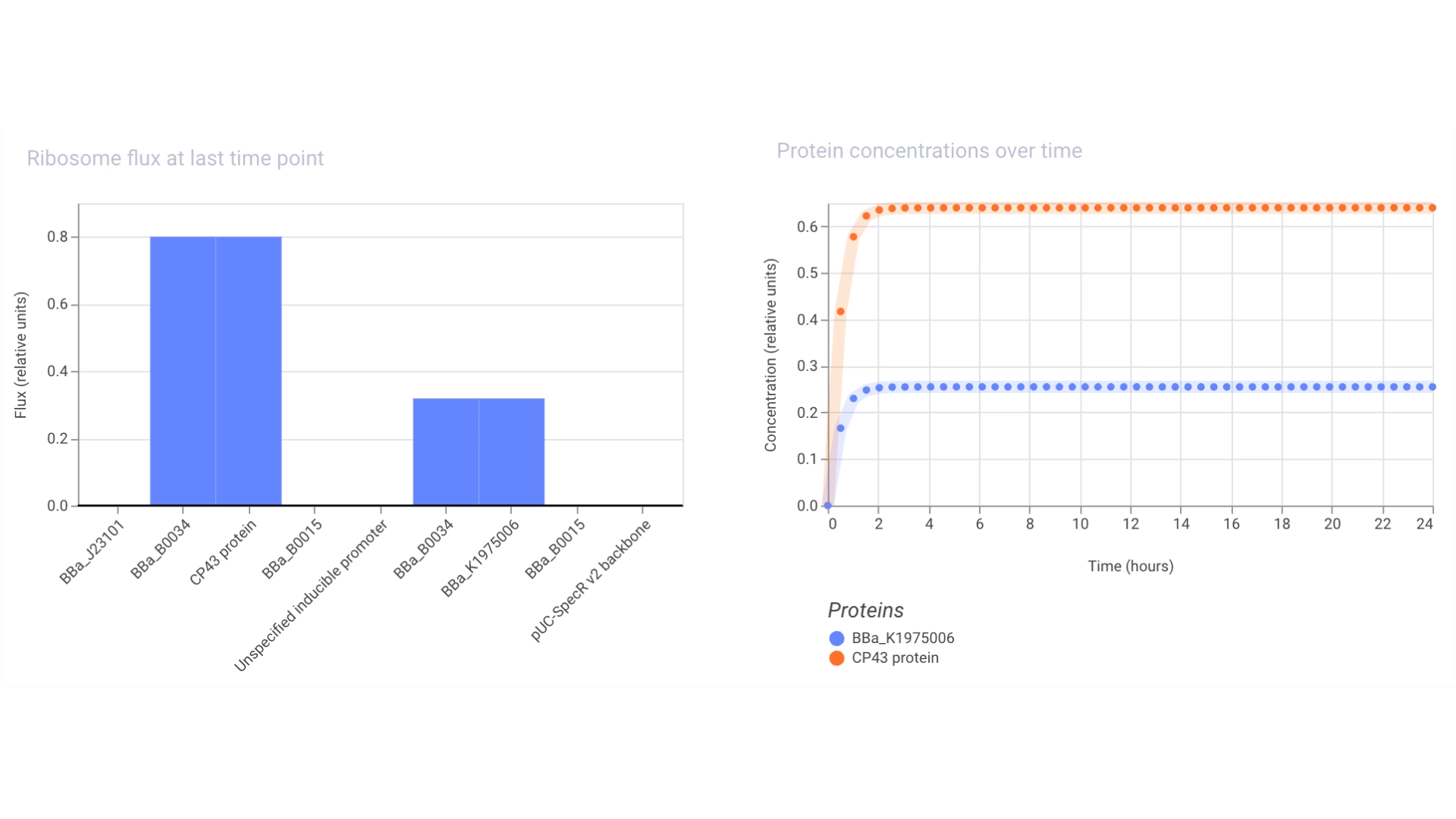
Construct #3:
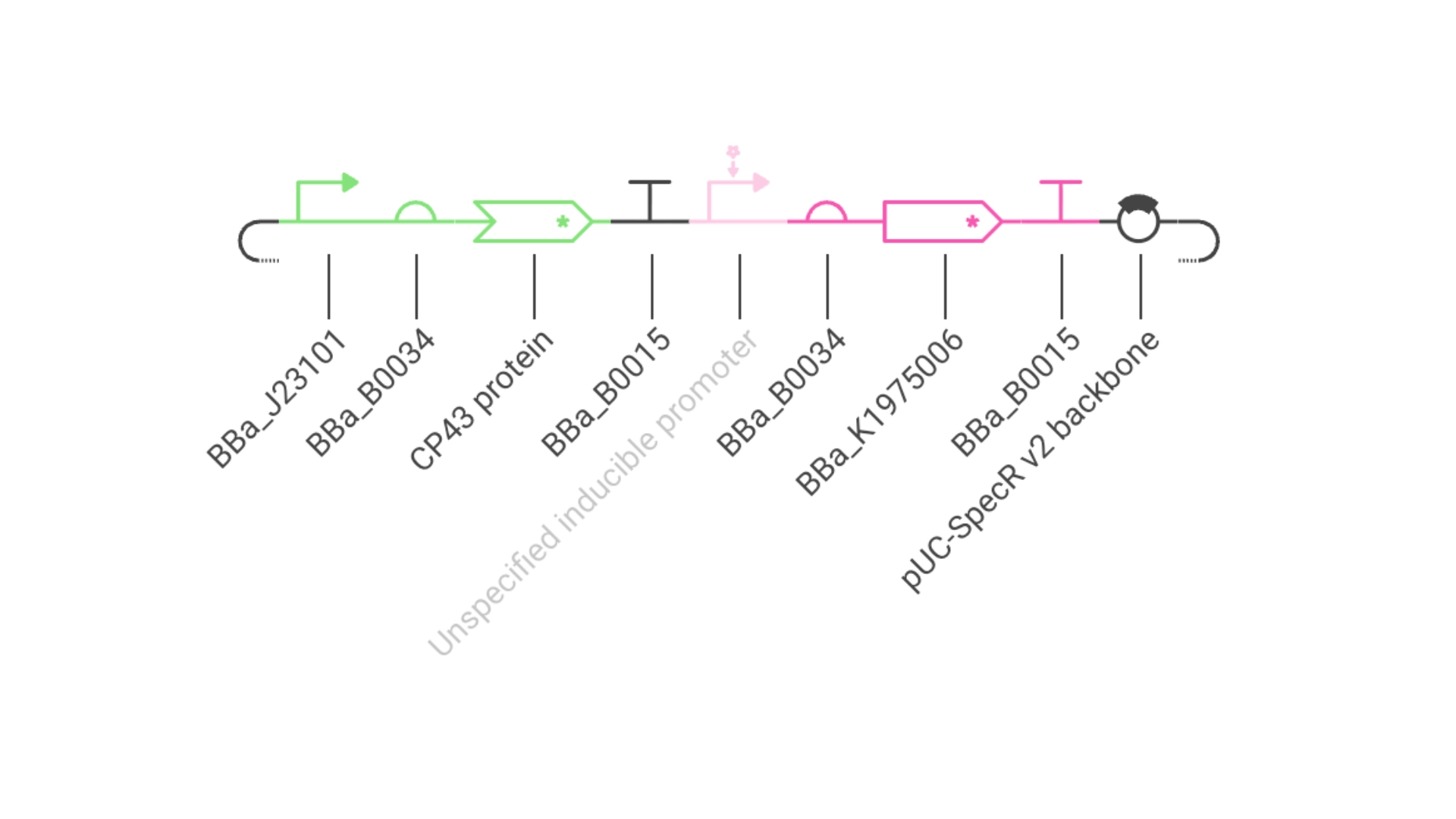
The objective of this construct would be to create a circuit that only expressed a fluorescent protein if the CP43 folded correctly— as it is a lipoprotein it needs an environment with chlorophyll which acts as folding co-factor— and to achieve this an inducible promoter could be designed to be induced by the folded CP43 protein. This way we would have a construct that both expresses the needed protein and gives feedback on the success of the reaction.
For this exercise I just used an Unspecified Inducible Promoter part
Glyph

 Construct #3
Construct #3