Week 11 HW: Building Genomes
Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork
- Colaborated in the stage where a single picture was being formed throughout the 4 plates
- It was a fun premise to explore could lab automation collaboratively
- Next year there could be the “constraint” of creating a single image/pattern throughout all the plates to really engage in the collective activity of trying to figure out how to expand on what other people started like a Cadavre Exquis logic
Part B: Cell-Free Protein Synthesis | Cell-Free Reagents
Part C: Planning the Global Experiment | Cell-Free Master Mix Design
3. The second phase of this lab will be to define the precise reagent concentrations for your cell-free experiment.
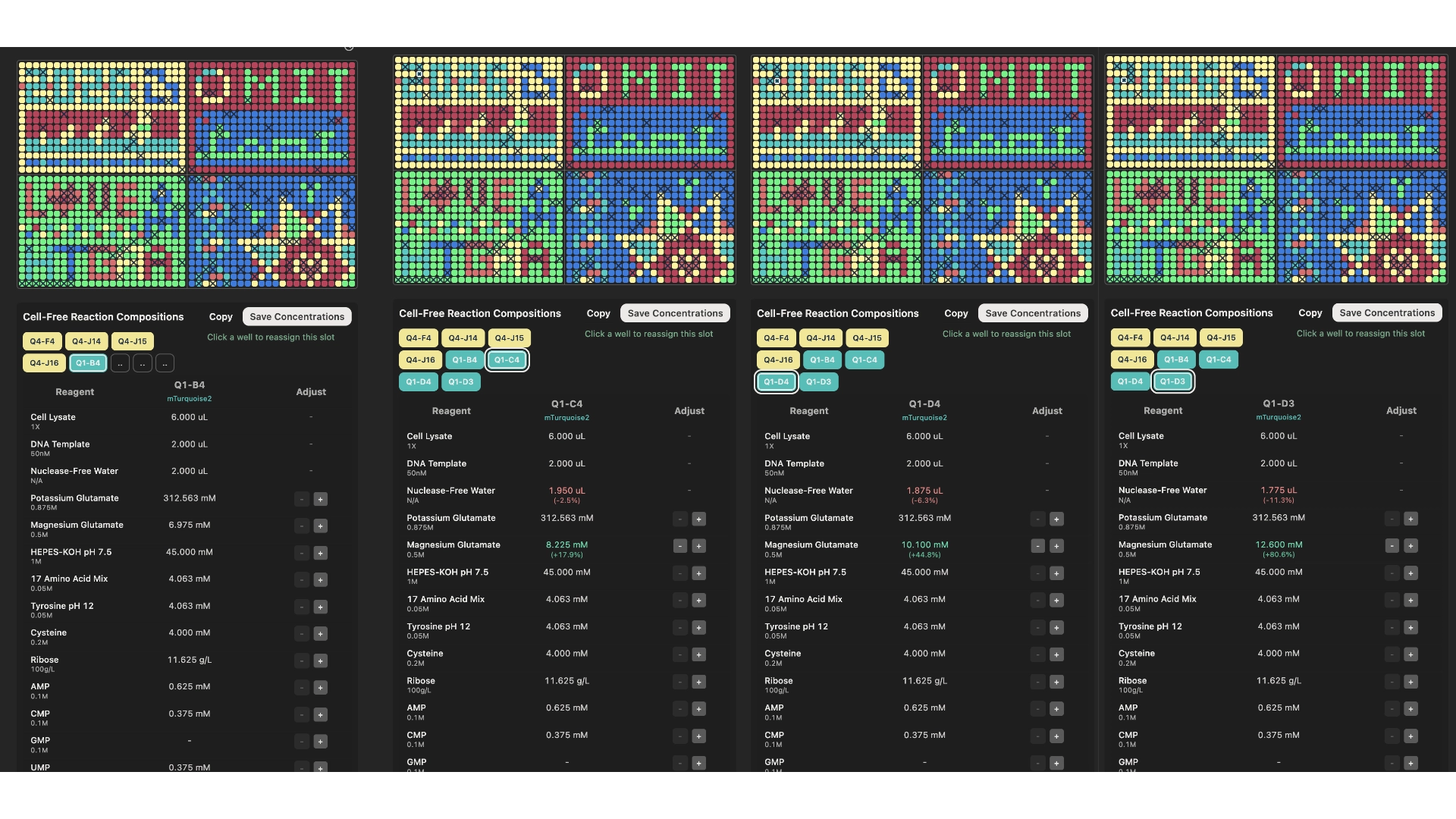
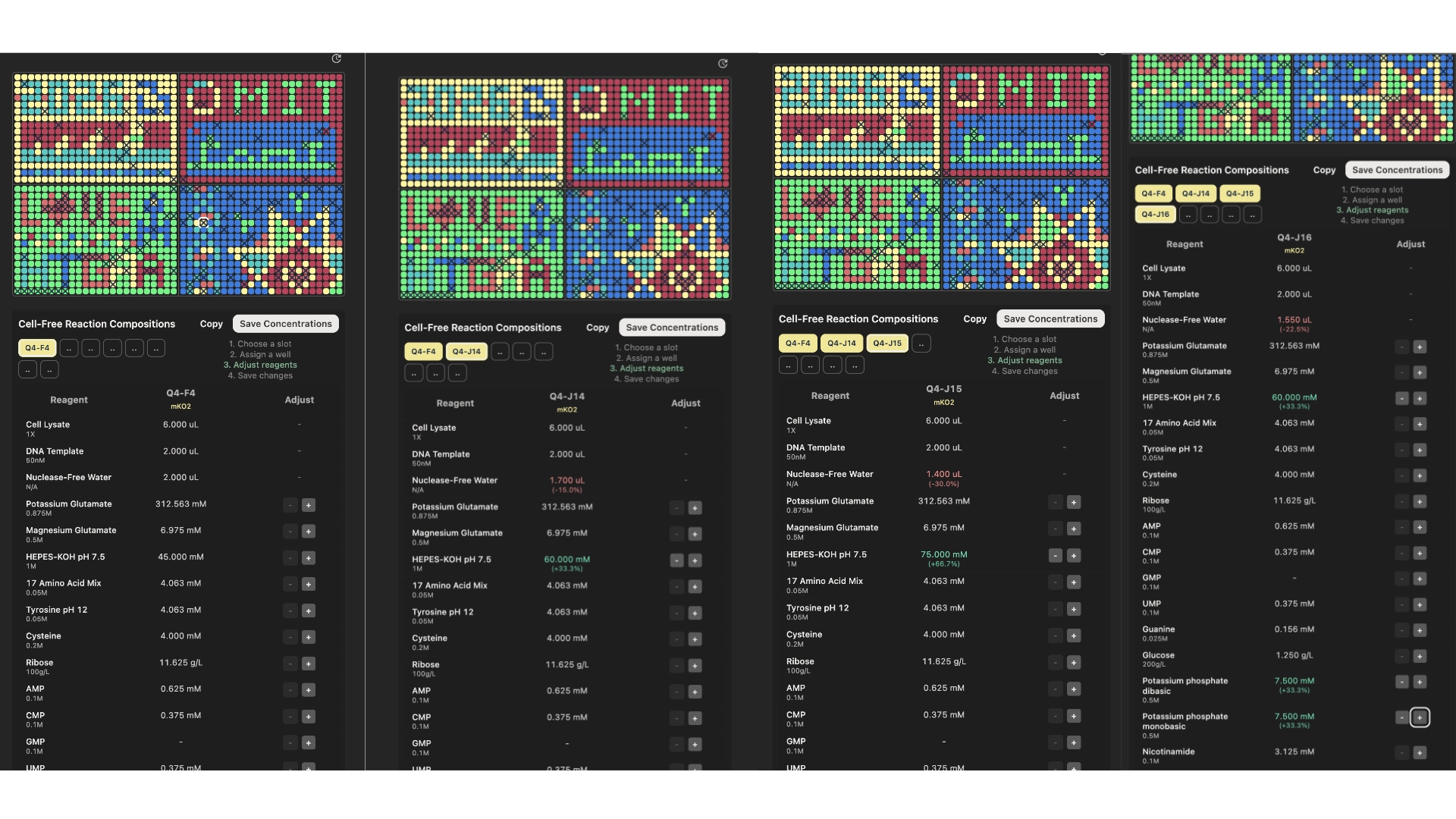
According to the hypothesis presented in the previous step, I created these 8 wells, 4 for the mKO2 and 4 for the mTurquoise2.
mKO2

mTurquoise2